Clear Sky Science · en
Translating spatial transcriptomic signatures in adenosquamous carcinoma into bulk prognostic biomarkers in lung adenocarcinoma: a bottom-up approach
Hidden warning signs inside lung tumors
Lung cancer remains one of the deadliest cancers, in part because tumors are mixtures of many different kinds of cells. Some of these cells may quietly drive the disease to become more aggressive or resist treatment, yet they are too rare or too blended in to be seen with standard lab tests. This study shows how ultra-detailed "maps" of gene activity inside tumors can uncover these hidden troublemakers and turn them into simple lab markers that help predict how patients with lung cancer will fare.
Looking closer at a rare, mixed tumor
The researchers focused on a rare lung cancer called adenosquamous carcinoma, which combines two main types of non–small cell lung cancer in a single mass: gland-like adenocarcinoma and flatter, skin-like squamous carcinoma. Because both components grow side by side in the same tumor, this cancer is an ideal model for studying how tumor cells might shift from one identity to another as they become more malignant. Using an imaging method called spatial transcriptomics, the team measured which genes were turned on in thousands of individual cells while preserving their exact positions in thin slices of tumor tissue. At the same time, they used standard staining tests for two common diagnostic markers, TTF‑1 (for adenocarcinoma) and p40 (for squamous cells), to anchor their high-tech maps in routine pathology practice. 
Color-coding tumor cells to find the odd ones out
Simply grouping cells by their gene patterns was not enough, because normal lung cells and tumor cells can sometimes look similar in gene readouts. To sharpen their view, the team used a visualization trick they call RGB‑UMAP. In this approach, every cell’s overall gene activity pattern is compressed into three numbers and translated into a red–green–blue color. Cells that behave similarly appear in similar colors when projected back into the tissue image. This allowed the scientists to spot pockets of normal cells hiding among tumor cells, and to remove them from their analysis, ensuring that they were studying true cancer cells rather than healthy bystanders.
Finding a hybrid cell state and a key sugar gate
After filtering out normal cells, the researchers searched for genes that were strongly and consistently active only in tumor cells. Among these, they discovered a group of tumor cells that were negative for both TTF‑1 and p40 yet showed a mixed gene pattern, sharing traits of both adenocarcinoma and squamous carcinoma. One gene stood out in this hybrid group: SLC2A1, which makes a protein known as GLUT1. GLUT1 acts as a gate that lets glucose, the cell’s fuel, enter from the bloodstream. In several tumors, SLC2A1 was abundant in squamous areas and in the TTF‑1/p40‑negative zones, but almost absent from normal lung tissue. The team confirmed this pattern in multiple patient samples, suggesting that high SLC2A1 marks a squamous-like, metabolically active subpopulation of cancer cells.
From microscopic maps to patient outcomes
To test whether this microscopic finding matters for real patients, the scientists turned to large existing datasets where bulk gene activity (averaged across whole tumor samples), DNA changes, and clinical outcomes were available for hundreds of people with lung adenocarcinoma. When they grouped these patients by SLC2A1 levels in their tumors, those with higher SLC2A1 had more gene changes typically seen in squamous cancers, a stronger signature of sugar-hungry metabolism, and significantly worse survival. This link held true even after accounting for factors like age, stage, smoking history, and sex, and it was confirmed in an independent public dataset. In contrast, in pure squamous tumors, SLC2A1 levels did not clearly separate patients by outcome, underscoring that the marker is especially informative within adenocarcinoma. 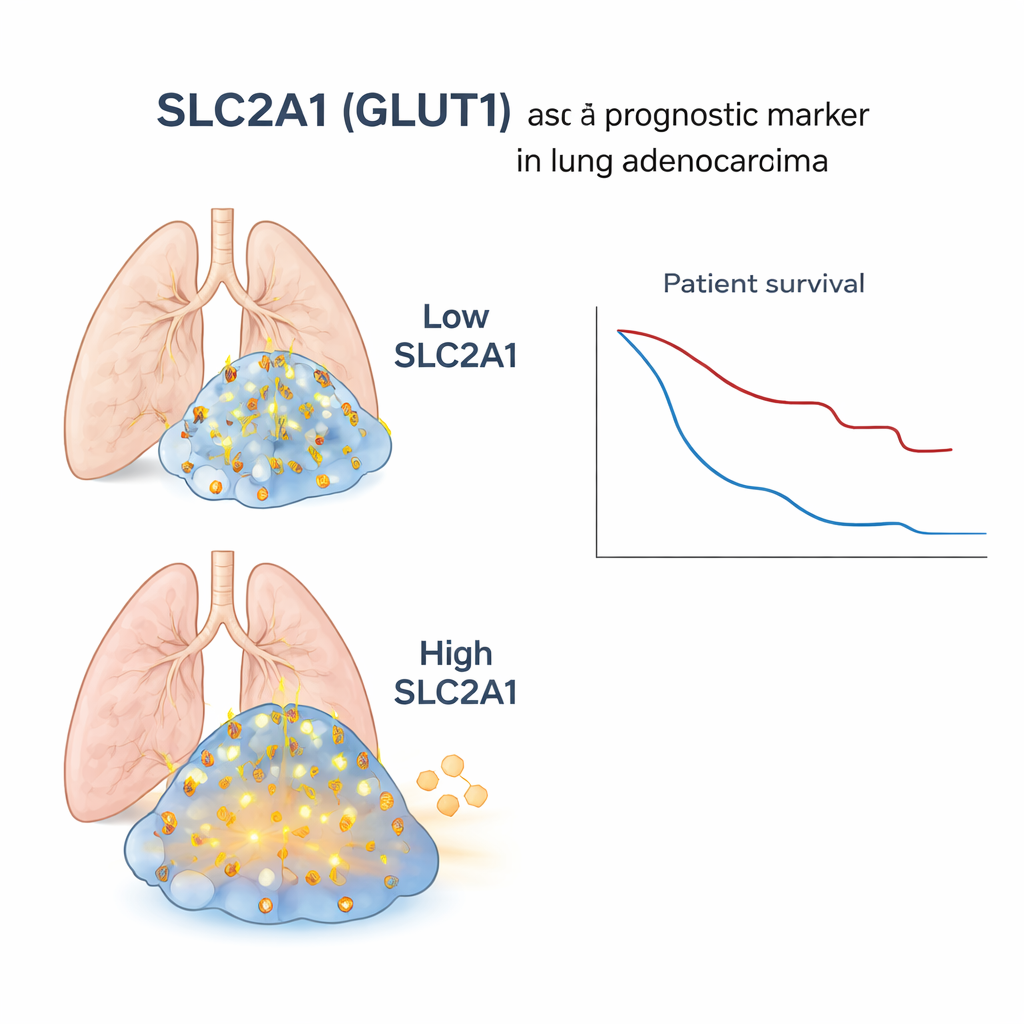
What this means for patients with lung adenocarcinoma
For a non-specialist, the key message is that within what looks like a single type of lung cancer under the microscope, there can be hidden pockets of cells that behave more like another, more aggressive type. By using high-resolution gene mapping, the researchers identified SLC2A1 as a simple, measurable sign of these squamous-like, fuel‑hungry cells lurking inside adenocarcinomas. High levels of this gene in standard tissue samples flag patients whose tumors are more likely to behave aggressively and who may need closer monitoring or different treatment strategies. More broadly, the study showcases a "bottom‑up" path—from single cells in a tissue slice to whole-tumor tests—that could be applied to other cancers to reveal dangerous cell states that routine methods currently miss.
Citation: Hatakeyama, K., Kawata, T., Muramatsu, K. et al. Translating spatial transcriptomic signatures in adenosquamous carcinoma into bulk prognostic biomarkers in lung adenocarcinoma: a bottom-up approach. npj Precis. Onc. 10, 77 (2026). https://doi.org/10.1038/s41698-026-01297-1
Keywords: lung adenocarcinoma, spatial transcriptomics, adenosquamous carcinoma, SLC2A1 GLUT1, cancer biomarkers