Clear Sky Science · en
An interpretable deep learning model for predicting endometrial cancer molecular subtypes from H&E-stained slides
Why this matters for women’s health
Endometrial cancer, which starts in the lining of the uterus, is one of the most common cancers in women, and its death rate is rising worldwide. Doctors now know that this cancer actually comes in several molecular “flavors” that respond differently to surgery, radiation, chemotherapy, and newer immune therapies. Today, finding those molecular subtypes usually requires costly and time‑consuming gene tests that many hospitals cannot easily offer. This study explores whether a carefully designed artificial intelligence (AI) system can read routine pathology slides—the pink‑and‑purple tissue images already taken for every patient—and accurately infer these molecular subtypes, potentially making precision care more widely available.
A closer look at tumor diversity
Not all endometrial cancers behave the same. Some grow slowly and stay confined to the uterus; others spread early and are harder to treat. Modern guidelines divide these tumors into four molecular subtypes based on DNA changes and how the cells repair genetic damage. These categories help predict outcome and guide decisions such as how extensive surgery should be and whether a patient might benefit from immunotherapy. However, the required genetic tests and special stains are expensive, depend on expert interpretation, and are often unavailable in smaller or resource‑limited hospitals. Pathologists have long suspected that many of these molecular differences leave visual clues in how cells and supporting tissue look under the microscope—but those clues can be too subtle and complex for the human eye to judge consistently.
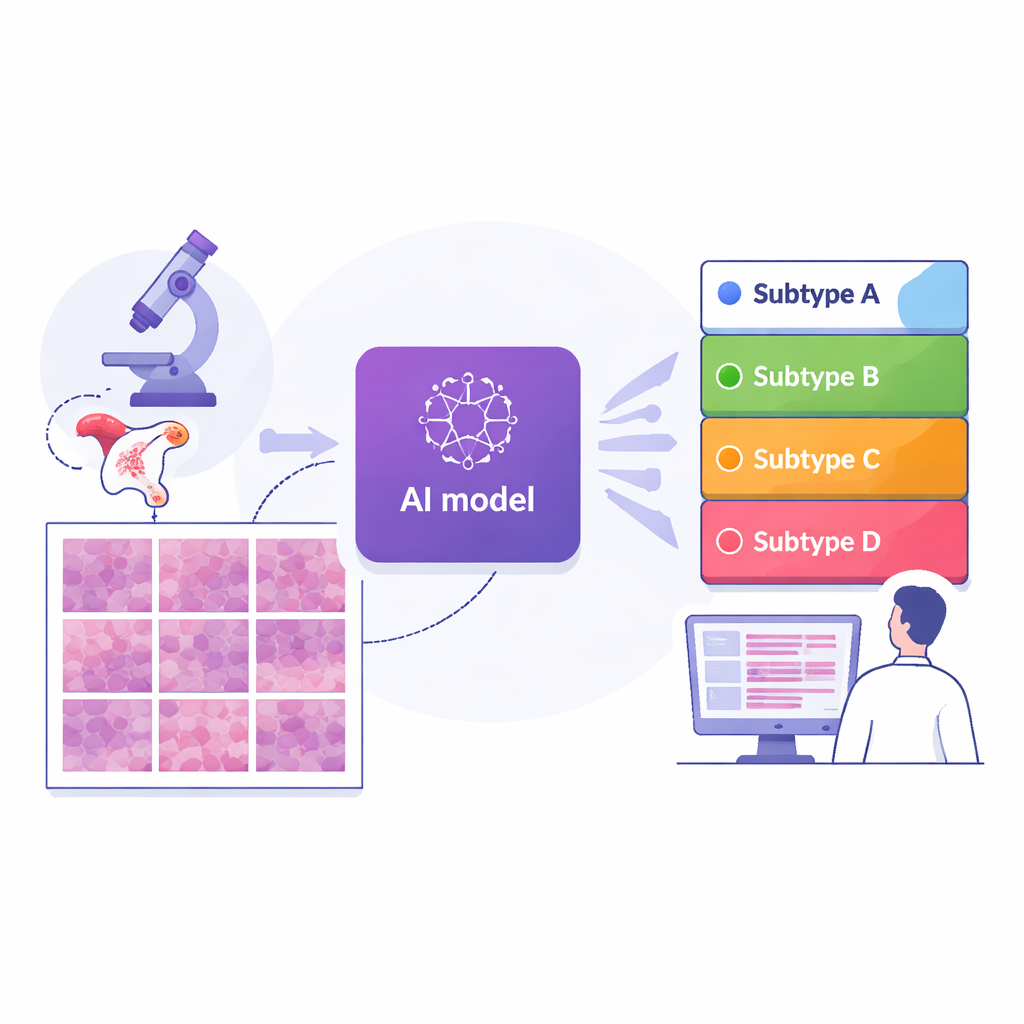
Teaching computers to read pathology slides
The researchers assembled digital images of 364 tissue slides from 324 women treated at a major cancer center in Shanghai, along with two independent comparison groups: 296 slides from an international public dataset and 36 from another hospital in Suzhou. Every case had already been assigned to one of the four molecular subtypes using advanced gene testing. The team then cut each whole‑slide image into thousands of small tiles and trained a deep learning model—a type of AI used in image recognition—to view each tile and estimate how likely it was to belong to each subtype. By averaging tile predictions across an entire slide, the system produced a single subtype prediction for each patient, closely mirroring how doctors think about the tumor as a whole.
How well the system performed
Across the main Shanghai group, the AI reached a high level of accuracy: its overall score for distinguishing among the four subtypes (measured by a standard statistic that ranges from 0.5 for guessing to 1.0 for perfect separation) was about 0.87. Performance remained strong—around 0.84—when tested on the two outside groups drawn from different hospitals and slide‑scanning systems, suggesting that the approach is reasonably robust. When compared with several leading AI strategies that use more complex attention or pooling schemes, this end‑to‑end model, built on a modern image‑analysis backbone, generally performed better. Importantly, the authors designed the system to be interpretable: they used visualization tools to highlight exactly which regions of each tile the AI relied on for its decisions.
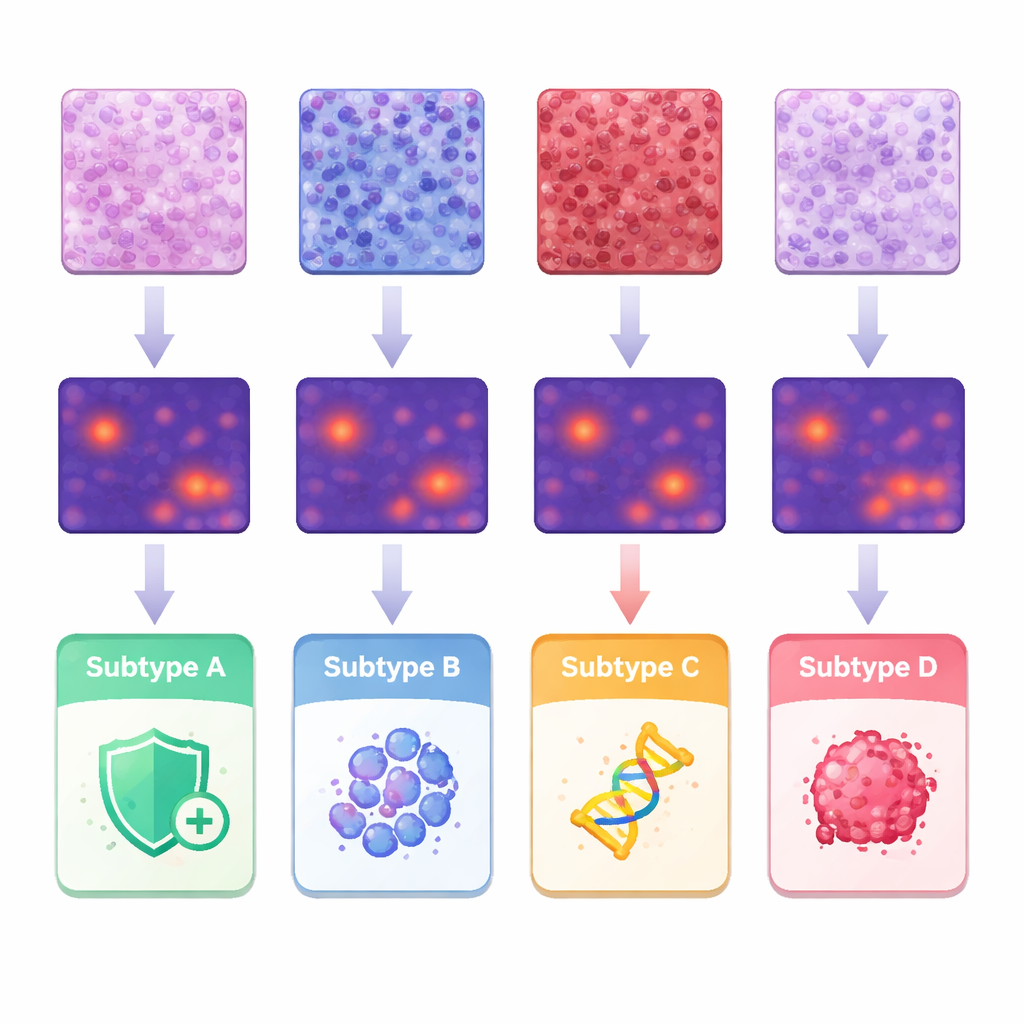
What the AI “saw” in the tumor micro‑environment
To understand what features drove the predictions, the team linked the AI’s heatmaps to classic pathology observations and to detailed measurements of individual cell shapes and arrangements. Tumors in one subtype showed dense infiltration of immune cells in the supporting tissue, while another subtype tended to have tightly packed structural cells. A third group displayed more solid sheets of highly abnormal tumor cells, and the subtype associated with aggressive behavior showed papillary, finger‑like structures and very irregular nuclei. By segmenting and analyzing about 245 million cells, the researchers quantified differences in cell size, variation, and spacing, and showed how certain combinations lined up with specific subtypes. These findings support the idea that molecular differences leave a recognizable imprint on tissue architecture that machines can systematically detect.
From proof of concept to clinical help
This work does not aim to replace genetic testing; instead, it proposes an “H&E‑first” triage tool that uses the standard stain already prepared for every biopsy. In practice, an AI‑generated subtype probability map could help pathologists decide which confirmatory tests to order first, prioritize limited tissue for the most informative assays, and speed up treatment decisions, especially in hospitals where full molecular profiling is hard to obtain. The study also highlights current limits, such as weaker performance for the rarest subtype and the need for larger, more diverse datasets before deployment. Still, it offers a compelling demonstration that routine microscope images carry enough hidden information for AI to approximate complex molecular labels, opening a path toward more equitable, data‑driven care for women with endometrial cancer.
Citation: Guo, Q., Cui, H., Zhang, Y. et al. An interpretable deep learning model for predicting endometrial cancer molecular subtypes from H&E-stained slides. npj Precis. Onc. 10, 71 (2026). https://doi.org/10.1038/s41698-026-01280-w
Keywords: endometrial cancer, digital pathology, deep learning, molecular subtypes, precision oncology