Clear Sky Science · en
Spatial transcriptomics reveals molecular heterogeneity and subtype-specific therapeutic targets in small cell lung cancer
Why this lung cancer study matters
Small cell lung cancer is one of the deadliest forms of cancer, striking mostly people with a history of smoking and often returning quickly after chemotherapy. Doctors know that not all small cell lung cancers behave the same, yet they have lacked tools to see how different cancer cell types and immune cells are arranged inside each tumor. This study used a new technology that reads gene activity in thousands of tiny spots across a tumor slide, creating a “molecular map” of each sample. By pairing this with custom computer analysis, the researchers uncovered hidden patterns that could point to more precise, subtype‑specific treatments.
Mapping tumors in place
The team analyzed tumor samples from 21 patients with limited‑stage small cell lung cancer who had surgery before receiving any drugs. Instead of grinding the tissue and losing all spatial information, they applied spatial transcriptomics: a technique that measures which genes are turned on at many defined spots across a tissue section. Each spot sits in its original position, allowing the scientists to link gene activity to where cancer cells, normal cells, and immune cells physically reside. They added two key tools on top of this: a new “Edgeindex” score to quantify how deeply tumor cells infiltrate neighboring tissue, and an artificial neural network—a form of machine learning—to automatically mark which spots are tumor and which are not.
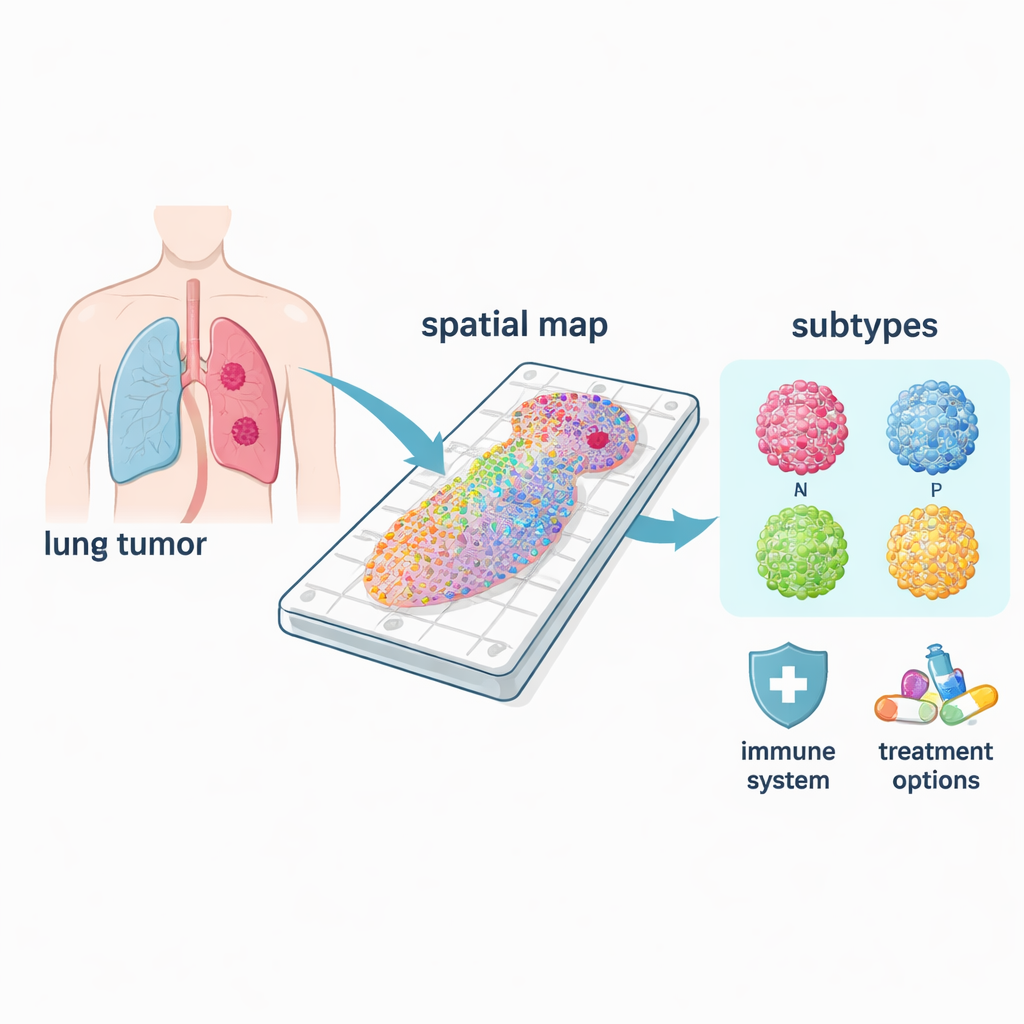
Different cancer types in one disease
Small cell lung cancer is not a single disease but includes at least four major molecular subtypes, often called ASCL1, NEUROD1, POU2F3, and YAP1, after key control genes that define them. Using their spatial maps, the researchers could see where each subtype dominated within tumors and how their biology differed. The common ASCL1 and NEUROD1 forms were strongly tied to cell‑division programs, matching their reputation as highly proliferative. In contrast, POU2F3 regions were linked to immune‑related pathways, while YAP1 areas showed signals related to metabolism and tissue remodeling, features that may connect to spread and treatment resistance. Because these patterns are resolved in space, the method can identify the true “driver” subtype within a mixed tumor, something bulk tests often miss.
The invading edge and the immune shift
One of the study’s central innovations, the Edgeindex, captures how much tumor cells intermingle with surrounding non‑tumor tissue. Low scores reflect compact, well‑bounded tumors, while high scores indicate scattered islands and ragged, invasive borders. Tumors with high Edgeindex were enriched for gene programs related to the extracellular matrix, metastasis, and cell death. They also showed a strikingly altered immune landscape: fewer helper T cells and other adaptive immune players, but more innate cells such as certain natural killer cells and monocytes. The authors interpret this as an “immunological niche remodeling,” where the tumor pushes away precise, targeted immune responses and leaves behind a less effective, largely innate defense that cannot fully control the cancer.
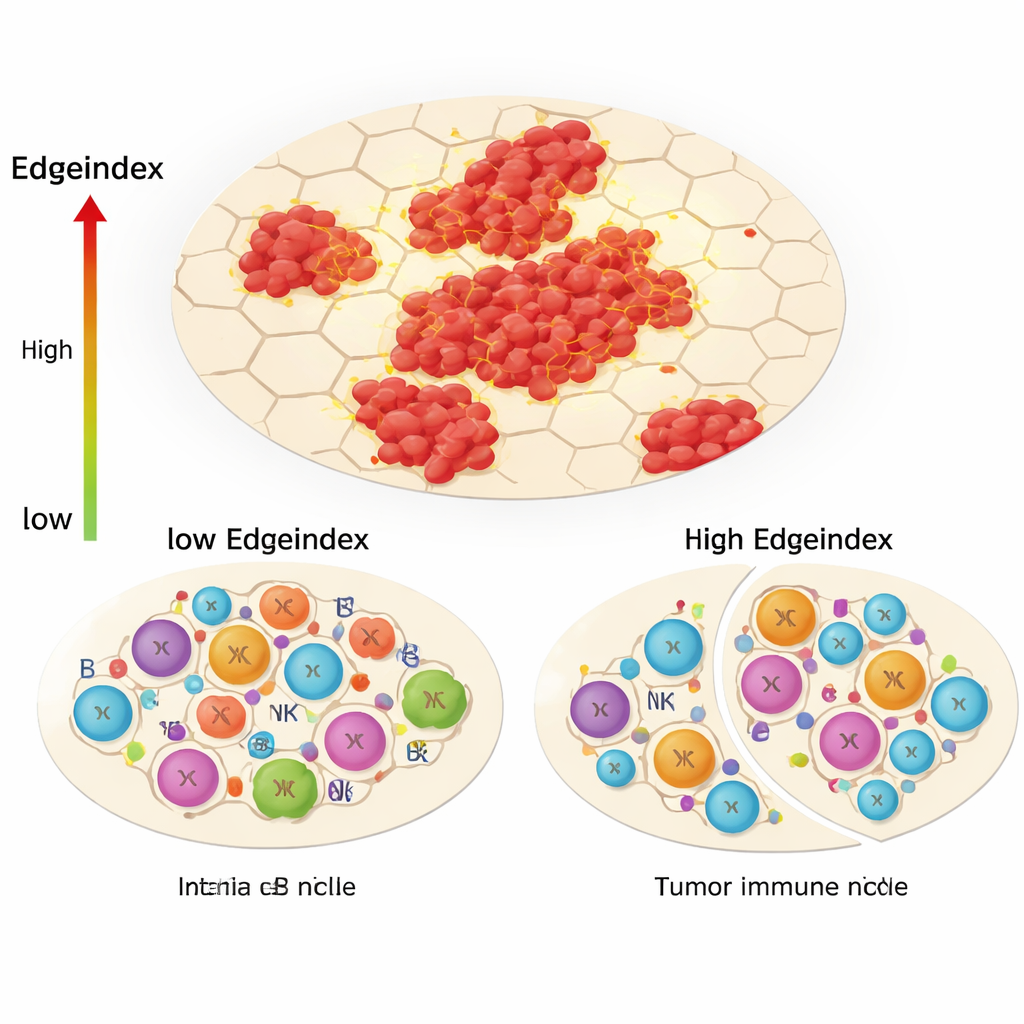
Hidden diversity in and around tumors
Beyond subtype and invasion patterns, the spatial data revealed that both the cancer cell populations and their surrounding microenvironments were internally diverse. By clustering spots within each tumor, the team counted how many distinct gene‑expression neighborhoods existed in the cancer regions and in the nearby non‑tumor tissue. Higher diversity inside the tumor was tied to immune‑related pathways, while certain patterns in the surrounding tissue linked to cell‑cycle activity and genetic regulation. The researchers also examined how strongly different regions “communicated” using known signaling molecules. Tumors with more intense communication between regions tended to show stronger immune‑related signals, reinforcing the idea that immune activity and tumor growth are tightly interwoven in space.
Following tumor development over time
To approximate how tumor cells change as the disease progresses, the team performed a “pseudotime” analysis that orders cells along a developmental path based on their gene activity. Across samples, they repeatedly identified one gene, UCHL1, as central to these trajectories. High UCHL1 levels coincided with strong cell‑cycle activity in tumor spots, while lower levels aligned with immune and cell‑death pathways. Because UCHL1 is a neural‑type protein previously tied to aggressive lung cancers, these findings strengthen the case that it may be a useful drug target, particularly for the neuroendocrine‑like cells that drive small cell lung cancer.
What this means for patients
In plain terms, this study turns flat microscope slides of small cell lung cancer into rich, layered maps that show where different cancer subtypes sit, how aggressively they invade, how they talk to their surroundings, and how the immune system responds or fails. The new Edgeindex score and mapping approach highlight which tumors are especially infiltrative and which immune cells are present or missing at the border. By tying these spatial patterns to subtype‑specific weaknesses—such as drugs that target DLL3, BCL‑2, AURKA, IGF‑1R/PARP, or immune checkpoints—the work moves the field closer to matching each patient with a tailored treatment strategy. While these tools still need to be tested in larger studies, they offer a blueprint for more precise and potentially more effective therapies in a cancer type that urgently needs better options.
Citation: Xie, T., Tang, L., Fan, G. et al. Spatial transcriptomics reveals molecular heterogeneity and subtype-specific therapeutic targets in small cell lung cancer. npj Precis. Onc. 10, 81 (2026). https://doi.org/10.1038/s41698-025-01243-7
Keywords: small cell lung cancer, spatial transcriptomics, tumor heterogeneity, tumor immune microenvironment, precision oncology