Clear Sky Science · en
PlantCLR: contrastive self-supervised pretraining for generalizable plant disease detection
Why smarter plant disease spotting matters
Plant diseases quietly steal food from the world’s dinner table, cutting crop yields and hurting farmers’ incomes. In many regions, only a few trained experts are available to identify problems in the field, and getting their help can be slow or impossible. This study introduces PlantCLR, a computer system that learns to recognize diseases from photos of leaves with far fewer human-provided labels than usual. By making automated diagnosis more accurate, more reliable, and easier to deploy on modest hardware, the work points toward smartphone or low-cost camera tools that could help farmers find trouble early and protect their harvests.
From leaf photos to early warnings
Today, many plant diseases are diagnosed the old-fashioned way: a person looks at a leaf and decides whether spots, yellowing, or curling are signs of infection. That judgment can vary from expert to expert and is easily thrown off by shadows, cluttered backgrounds, or different growth stages. Computer vision systems based on deep learning have begun to help, but they usually demand many thousands of carefully labeled photos. In agriculture, such labeled images are scarce and expensive to collect, while huge piles of unlabeled pictures from mobile phones and field cameras often go unused. PlantCLR is designed to tap into this unlabeled data, learning what diseased and healthy leaves tend to look like before it ever sees a label.
Teaching a model to learn by comparison
PlantCLR relies on a recent approach called contrastive self-supervised learning, in which a model teaches itself by comparing images rather than by reading labels. First, the system takes an unlabeled leaf image and creates two slightly different versions through random crops, flips, color shifts, or blur. These two versions should clearly represent the same leaf, so the model is trained to treat them as a matching pair and give them similar internal representations, while pushing apart representations of different leaves in the same training batch. This pretraining stage uses a compact but modern image-processing backbone called ConvNeXt-Tiny, paired with a small extra module that is used only during this learning-by-comparison step. 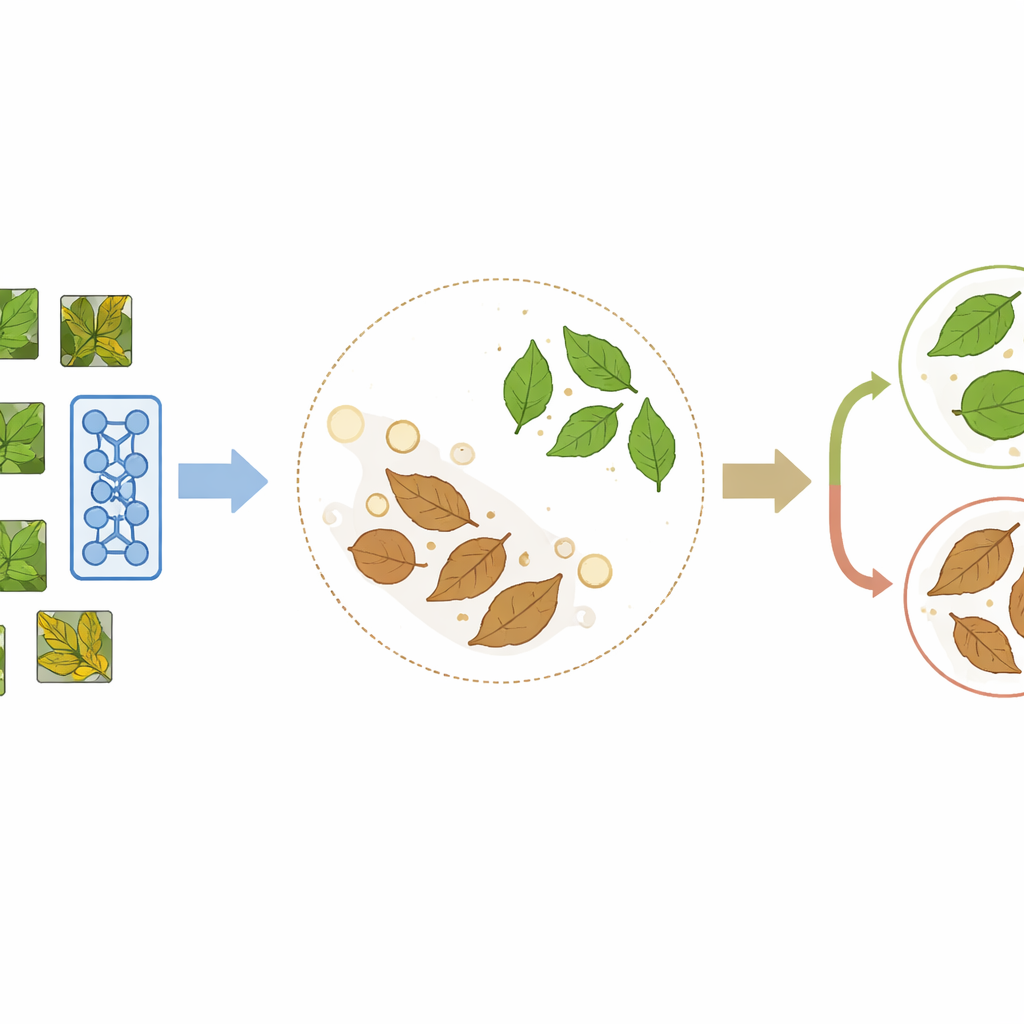
Putting the system to the test
To test how well this strategy works in practice, the authors turned to two popular leaf datasets that mimic very different real-world settings. The PlantVillage dataset contains more than 54,000 images of leaves photographed under neat, controlled conditions, usually with clean backgrounds and clear symptoms across 38 disease and crop categories. In contrast, the Cassava Leaf Disease dataset contains roughly 21,000 images of cassava leaves taken directly in the field, with messy backgrounds, uneven light, and leaves that overlap or twist in many directions across five classes, including several serious viral and bacterial infections. The study uses PlantVillage mainly as a rich source of unlabeled images for pretraining and then evaluates performance both on that dataset and, more critically, on the harder, field-style cassava photos.
Rugged performance across changing conditions
PlantCLR reached 99.10% accuracy on the PlantVillage test set and 96.83% accuracy on the Cassava test set, with similarly high F1-scores that show the model does well even on less common diseases. These numbers beat a range of well-known deep networks, including DenseNet, ResNet, VGG, and a vision transformer model, all trained in a purely supervised way under carefully matched conditions. 
Why this approach is a step forward
For non-specialists, the key message is that PlantCLR shows how a machine can become a capable plant doctor by first learning from large collections of unlabeled pictures and then refining its skills with a smaller, labeled set. This strategy not only reaches very high accuracy but also holds up well when the camera moves from the lab to the field, where conditions are far less tidy. Because the underlying model is relatively lightweight, it could eventually be deployed on affordable hardware, making advanced disease detection more accessible to farmers and extension workers worldwide. In short, the study demonstrates a practical path toward scalable, robust, and label-efficient plant health monitoring tools that could help safeguard food supplies.
Citation: Shah, S.S.A., Saeed, F., Raza, M.U. et al. PlantCLR: contrastive self-supervised pretraining for generalizable plant disease detection. Sci Rep 16, 10550 (2026). https://doi.org/10.1038/s41598-026-45684-x
Keywords: plant disease detection, self-supervised learning, contrastive learning, agricultural AI, crop health monitoring