Clear Sky Science · en
Proof of concept for aqueous two-phase system-based extraction of cell-free DNA from plasma for liquid biopsy applications
Why a Blood Test Matters for Cancer and Other Diseases
Many modern medical tests look for tiny fragments of DNA that drift freely in our blood. These bits, known as cell-free DNA, can reveal whether a tumor is shedding genetic material, whether a transplanted organ is being rejected, or how a pregnancy is progressing. But because these DNA fragments are scarce, short, and mixed with a soup of proteins and other molecules, pulling them out cleanly and quickly is surprisingly hard. This study introduces a new, simpler way to extract these fragments from blood plasma using a gentle water-based separation method, potentially making advanced “liquid biopsy” tests more accessible and affordable.
The Challenge of Fishing Out Tiny DNA Pieces
Doctors and researchers use liquid biopsies to read genetic clues from a simple blood draw instead of cutting out tissue. The catch is that cell-free DNA in plasma usually exists at only a few dozen nanograms per milliliter and is chopped into short pieces. A small fraction of this material may come from a tumor or from a transplanted organ, so every fragment counts. Standard extraction kits capture DNA by making it stick to silica surfaces in the presence of strong salts, followed by several washing and spinning steps. These kits work well but take time, require special equipment, and can be expensive. They can also struggle to recover the shortest fragments and may bring along unwanted long stretches of normal genomic DNA, which can mask rare disease-related signals.
A Two-Layer Water Trick to Catch DNA
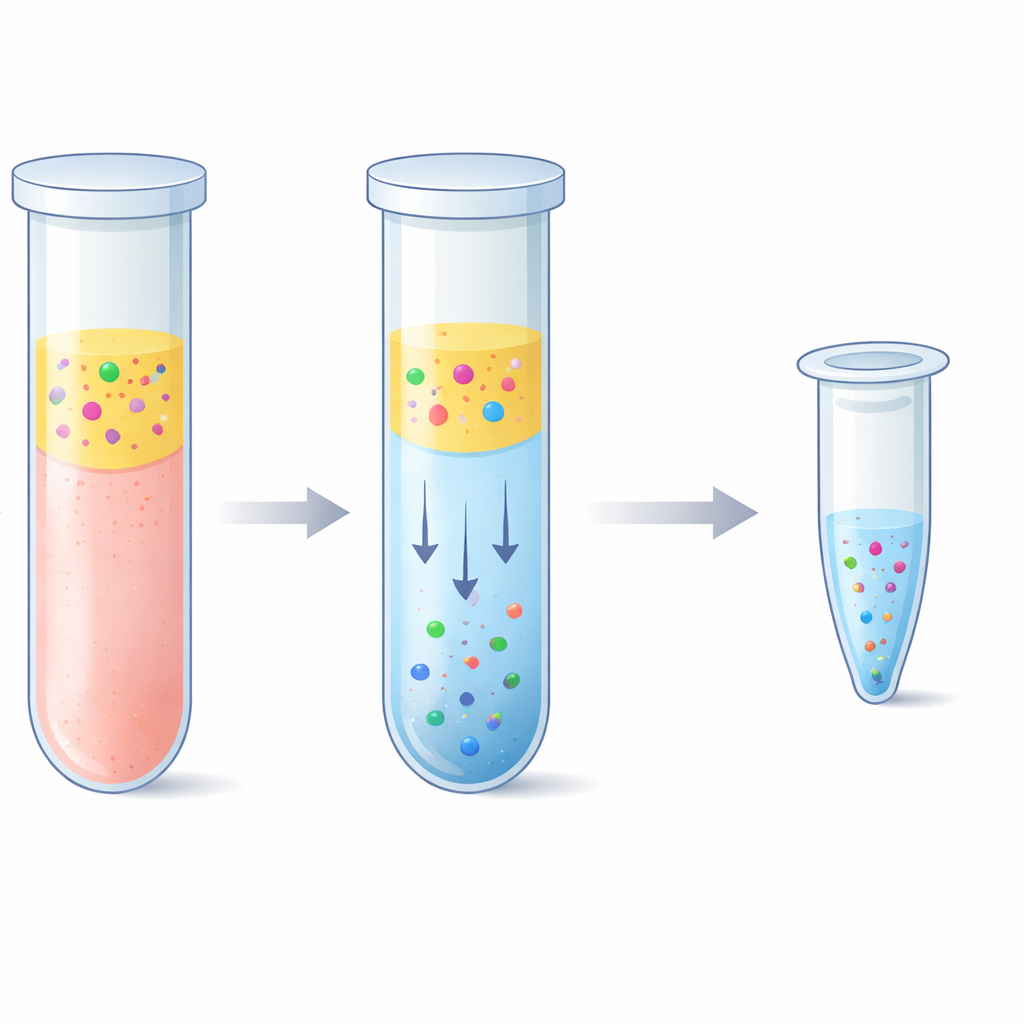
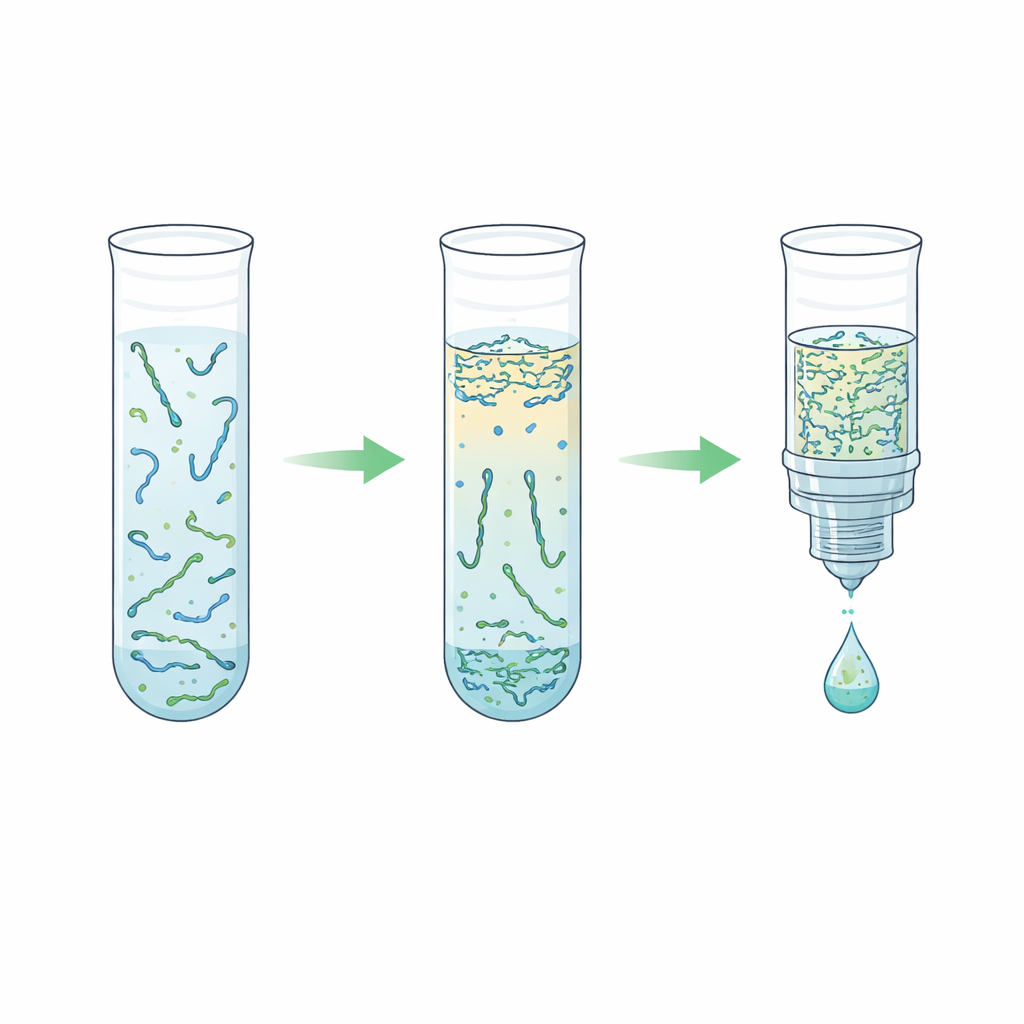
The team explored an alternative based on an "aqueous two-phase system"—in plain terms, two water-rich liquids that do not fully mix, like oil and vinegar but both made mostly of water. By combining plasma with a polymer called polyethylene glycol and simple phosphate salts, the mixture naturally separated into a top and bottom layer. Remarkably, the short DNA fragments strongly preferred the salt-rich bottom layer, while most blood proteins and long DNA strands stayed in the top layer or at the interface. The researchers carefully tuned the recipe so that this bottom layer stayed small enough to concentrate the DNA but gentle enough to avoid damaging it. They then paired this step with a “reverse elution” purification, which pushes the salty bottom layer through a porous matrix to remove salt and shrink the final volume, ending with a cleaner and more concentrated DNA solution.
Refining the Process for Real-World Use
Through many rounds of testing, the authors adjusted the balance of polymer and salt, spinning speeds, volumes, and optional lysis steps that break up protein–DNA complexes. They found that increasing polymer content along certain composition lines doubled the DNA concentration without harming recovery, and that stronger conditioning in the reverse elution step roughly quadrupled concentration compared with their earlier work. Surprisingly, they discovered that skipping harsh lysis before the two-layer separation often worked best; lysis tended to disturb the phases and reduce yield. A streamlined, largely lysis-free workflow removed about 99.7% of plasma proteins while recovering up to roughly two-thirds of the short DNA input in a fourfold smaller volume, all within about ten minutes of total processing time and only a couple of minutes of hands-on work.
How the New Method Stacks Up
The researchers compared their approach to a widely used commercial silica kit. The standard kit produced somewhat higher total DNA yields and stronger concentration because it elutes into very small volumes. However, the two-phase method showed consistent recoveries above 60% across a range of DNA inputs, even when starting with very small amounts, and required far fewer manual steps and less equipment. Importantly, the new process acted like a built-in filter for fragment length: it enriched for short, cell-free–like DNA while largely excluding long strands that can come from broken blood cells. Tests using quantitative PCR showed that the cleaned extracts did not inhibit downstream amplification. When the team used a commercial reference sample containing known cancer-related mutations, the DNA recovered with their method could be turned into sequencing libraries, run on a high-throughput sequencer, and used to detect all expected variants at the right frequencies.
What This Could Mean for Future Blood Tests
In plain terms, this work shows that a relatively simple, water-based separation can replace more complex solid-surface chemistry for preparing cell-free DNA from plasma. Although the new method currently yields slightly less DNA than top-of-the-line commercial kits, it offers rapid processing, strong removal of unwanted proteins and long DNA, and ready compatibility with PCR and next-generation sequencing. If further validated on real patient samples and refined for automation, this approach could lower costs and simplify laboratory workflows, helping bring precise, genetics-based blood tests for cancer, transplantation, and other conditions into wider everyday use.
Citation: Meutelet, R., Buerfent, B.C., Hess, T. et al. Proof of concept for aqueous two-phase system-based extraction of cell-free DNA from plasma for liquid biopsy applications. Sci Rep 16, 11232 (2026). https://doi.org/10.1038/s41598-026-45585-z
Keywords: liquid biopsy, cell-free DNA, cancer blood test, DNA extraction, next-generation sequencing