Clear Sky Science · en
Crystallographic data for Pyrococcus furiosus dolichylphosphate mannose synthase suggest that the enzyme could flip its glycolipid product
How heat-loving microbes keep their cell coats in shape
Many microbes that thrive in boiling-hot environments survive thanks to a protective sugar coat on their cell surface. Building this coat requires special fat–sugar molecules that must be made on one side of the cell membrane and then moved to the other. This study dissects one such molecule-making enzyme from a heat-loving microbe and suggests that, remarkably, the same protein may also help flip its product across the membrane—doing two jobs at once.
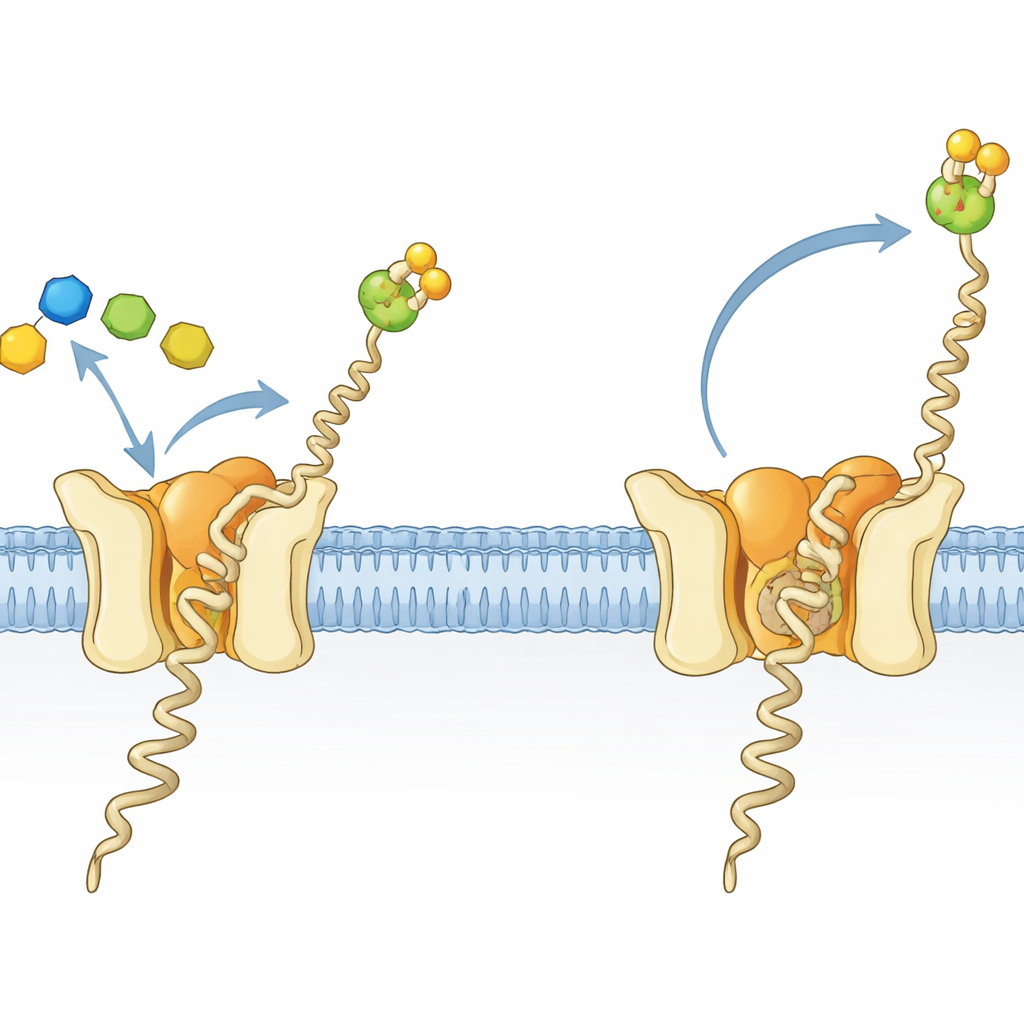
A tiny factory for a crucial sugar–fat molecule
Cells in all domains of life decorate many proteins with chains of sugars, a process essential for proper folding, stability, and cell–cell communication. In archaea and eukaryotes, a key building block for these decorations is a sugar–fat molecule called dolichylphosphate mannose (Dol-P-Man). It consists of a long greasy tail anchored in the membrane and a sugar–phosphate head that sticks out into the watery surroundings. Dol-P-Man is made by the enzyme dolichylphosphate mannose synthase (DPMS), which transfers a mannose sugar from a soluble donor onto a membrane-bound lipid. In the hyperthermophilic archaeon Pyrococcus furiosus, DPMS is a single-chain protein, dubbed type-III DPMS, with a soluble catalytic region attached to a four-helix membrane-spanning section whose role had been unclear.
Capturing the enzyme in the act
The researchers crystallized P. furiosus DPMS together with its substrates and followed the reaction by X-ray crystallography. Their new structure, combined with three earlier ones, captures a series of snapshots around the moment when the sugar is transferred. In one state, the donor molecule (GDP-mannose) and a Dol-P–like acceptor are bound in a precise geometry that places the reactive carbon of mannose directly in front of the phosphate group on Dol-P—an ideal “pre-transfer” arrangement. Loops on the enzyme act like front and back doors: they close to hold donor and metal ion in place, then rearrange to open a path for Dol-P to enter and for the spent GDP to leave once the sugar has been passed on. These details clarify how the enzyme ensures efficient and accurate sugar transfer at the membrane surface.
An upside-down product hidden in the membrane
More surprising was a second, distinct state seen in the same crystals. Here, the finished Dol-P-Man molecule is not at the active site but lodged deeper inside the membrane-spanning part of the protein, in an “upside-down” orientation. Its sugar–phosphate head sits in a polar pocket between two pairs of helices (TMD1 and TMD2), while the greasy tail runs along a groove formed by the helices. A key amino acid, a phenylalanine, appears to act as a gate that can either block or open access to this pocket. The quality of the electron density and the way the protein packs in the crystal argue that this flipped Dol-P-Man is not an artifact of crystal contacts, but a genuine, well-populated state the enzyme can adopt.
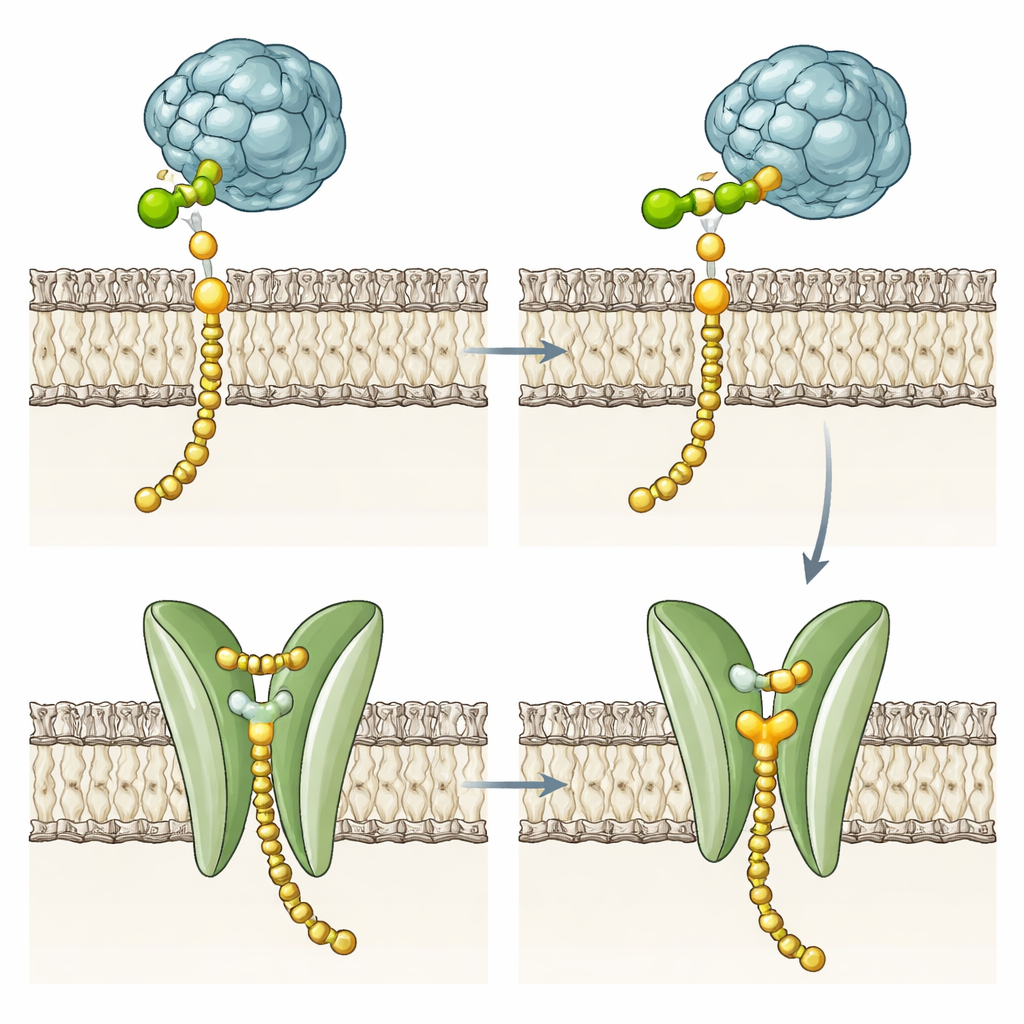
Mutant enzymes hint at a second job
To test whether the membrane-spanning domain is required for the sugar-transfer chemistry, the team engineered multiple variants. They deleted most or all of the helices, swapped them for a different membrane segment from yeast DPMS, or altered key pocket residues and the phenylalanine gate. Remarkably, these mutants still made Dol-P-Man at similar rates to the intact enzyme, although some were slightly less stable in the membrane. This shows that the core chemistry resides in the soluble catalytic region and that the four-helix bundle is not needed to attach mannose to Dol-P. Instead, its conserved polar pocket, groove, and gating residue suggest a specialized role in handling the finished product, especially in hyperthermophilic archaea whose rigid membranes make spontaneous lipid flipping energetically costly.
Why a dual-purpose enzyme matters
Putting the structural and mutational evidence together, the authors propose that type-III DPMS is a “moonlighting” protein. First it synthesizes Dol-P-Man on the inner face of the archaeal plasma membrane; then its transmembrane domain may help flip the sugar–fat molecule so the head group emerges on the cell exterior, ready to be consumed by other enzymes that build the cell’s sugar coat. Evolution seems to have favored this chimeric design mainly in extremely heat-loving archaea, where bringing the site of synthesis close to a dedicated flipping groove could minimize waste and thermal damage. Although this work does not yet prove flipping activity directly, it offers a concrete structural model for how an enzyme can both create and reposition a vital lipid intermediate across a harsh membrane barrier.
Citation: Gandini, R., Keskitalo, M.M., Reichenbach, T. et al. Crystallographic data for Pyrococcus furiosus dolichylphosphate mannose synthase suggest that the enzyme could flip its glycolipid product. Sci Rep 16, 9076 (2026). https://doi.org/10.1038/s41598-026-44343-5
Keywords: dolichylphosphate mannose, lipid flippase, archaea, protein glycosylation, membrane enzymes