Clear Sky Science · en
Label-free multiphoton microscopy and machine learning for recognition of hepatocellular carcinoma
Why this matters for people with liver cancer
Liver cancer, particularly hepatocellular carcinoma, is both common and deadly, and surgery is often the only chance for a cure. During an operation, surgeons must remove the entire tumor while sparing as much healthy liver as possible, a difficult balancing act when diseased liver tissue already looks abnormal. This study explores a new way to "see" cancer in real time during surgery, without dyes or lengthy lab work, by combining advanced laser microscopy with artificial intelligence. The goal is to help surgeons find the true edge of the tumor more quickly and accurately, potentially improving outcomes for patients.
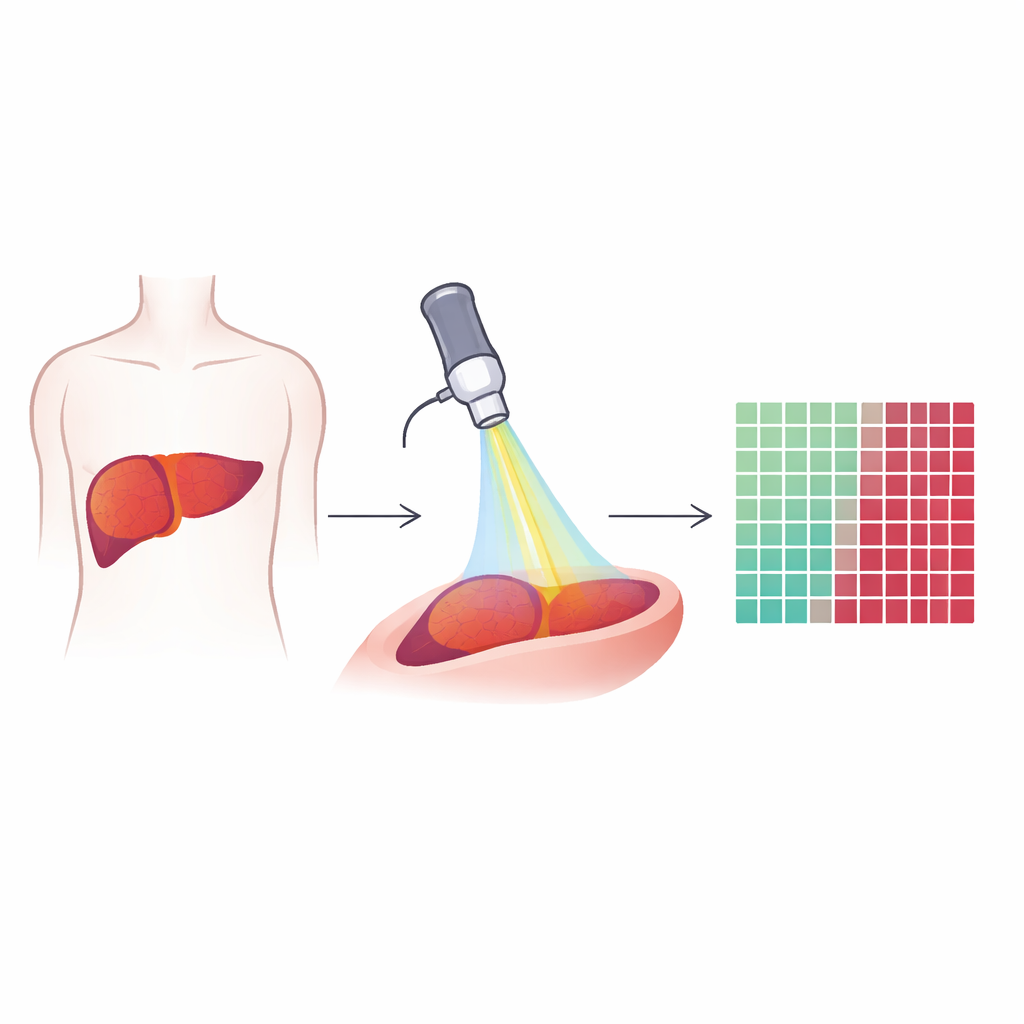
Looking inside tissue with special light
The researchers used a technique called multiphoton microscopy, which relies on very short laser pulses to make natural molecules in tissue light up or generate signals. Instead of adding stains or contrast agents, the method reads out what is already there: fats, collagen fibers, and naturally fluorescent substances inside cells. Three types of signals were recorded from each tiny field of view in liver samples: one that highlights lipids, one that shows autofluorescence from molecules like NADH and vitamin A, and one that reveals stiff collagen fibers. Together, these signals create colorful, high-resolution images that show both individual cells and the overall architecture of liver tissue and tumors.
Capturing the many faces of liver cancer
The team examined tissue from 76 patients who had surgery for liver cancer, looking at both the tumor and the surrounding liver. They showed that this label-free imaging can depict classic growth patterns of hepatocellular carcinoma, such as thin or thick plates of tumor cells, solid sheets, and gland-like structures. It also reveals changes in the surrounding liver, including fat build-up and scarring, which are common in patients with chronic liver disease. Importantly, these patterns were visible even when the images were acquired at relatively low resolution, similar to what future endoscopic devices inside the body could provide. This suggests the approach could be practical in the operating room, not just in a lab.
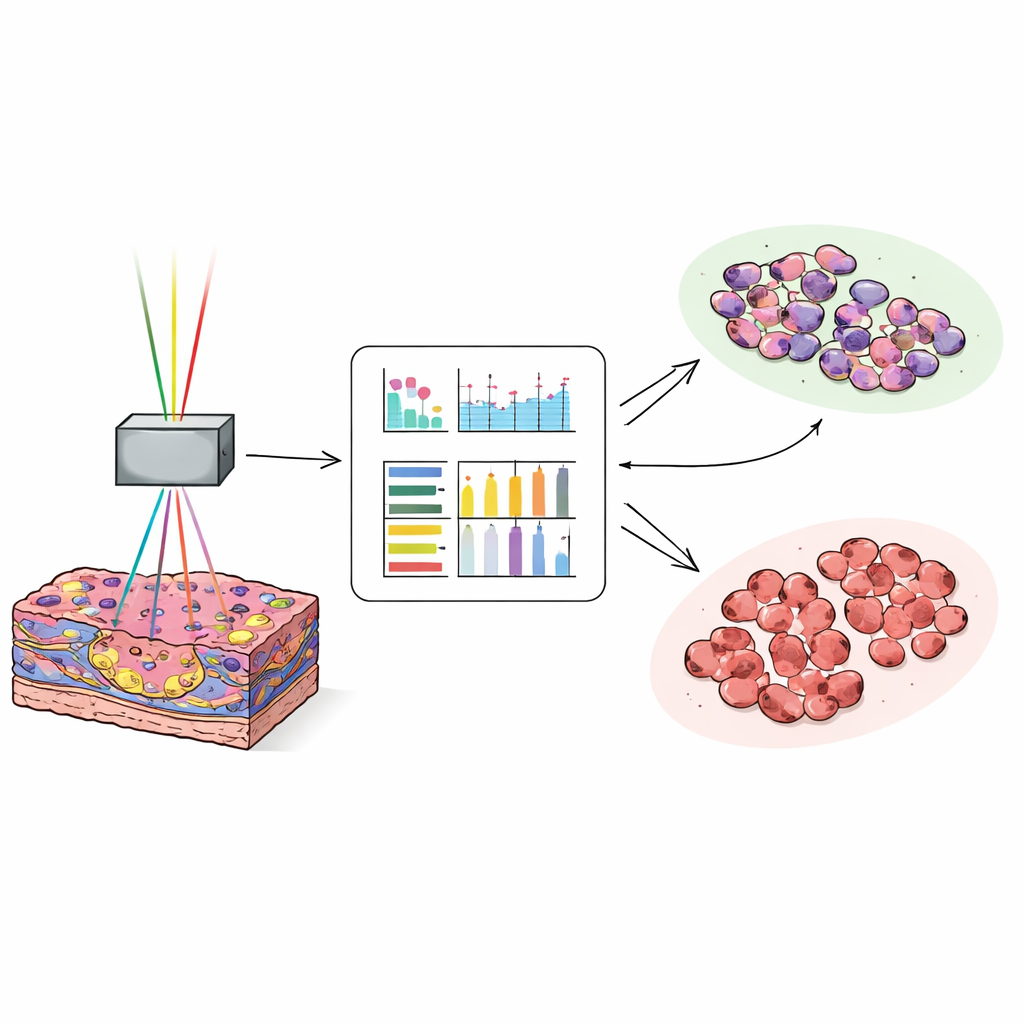
Teaching a computer to spot the tumor edge
To turn rich but complex pictures into quick decisions, the scientists transformed each image into numbers that describe texture, such as how smooth, coarse, or varied the pixel patterns are. For each image channel, they computed 17 such measurements and then trained a neural network on about 25,000 images from 35 patients to distinguish tumor tissue from non-cancerous liver. They then tested the model on around 27,000 new images from another 38 patients. When the computer used a combination of the autofluorescence and collagen-sensitive signals, it correctly labeled images as tumor or normal in about 97 out of 100 cases. Even the most subtle, well-differentiated tumors, which can be hard for pathologists to recognize on quick frozen sections, were classified with high accuracy.
What the light patterns reveal about cancer
By comparing different signal combinations, the study found that autofluorescence carried the most useful information for telling cancer from non-cancer. This glow arises from many natural molecules inside cells and from structures like elastin and collagen, so it encodes cell density, nuclear size and shape (seen as darker areas), fat droplets, and the makeup of the surrounding tissue. Adding the collagen-specific signal further sharpened the contrast between tumor and background liver, reflecting known differences in how fibers and blood vessels are arranged. Surprisingly, the lipid-focused signal contributed less to automatic classification, a practical insight because it may allow simpler device designs that rely on one laser beam instead of more complex two-beam setups.
From lab bench to operating room
To mimic real surgical use, the team also applied their best model to tissue samples that included the actual border between tumor and liver, creating probability maps that highlighted where cancer stopped and normal tissue began. These maps generally agreed well with conventional histology, though highly fatty or collagen-rich regions sometimes confused the classifier. The authors note remaining hurdles before this technology can guide live surgery, including the limited depth of optical imaging, the need for robust, sterile endoscopes, motion and blood in the surgical field, and the challenge of validating artificial intelligence on large, real-world datasets. Still, the work shows that label-free multiphoton imaging, combined with machine learning, can act like a rapid, digital microscope that may one day help surgeons cut more precisely and safely, increasing the chances that all of the cancer is removed while preserving as much healthy liver as possible.
Citation: Galli, R., Korn, S., Aust, D. et al. Label-free multiphoton microscopy and machine learning for recognition of hepatocellular carcinoma. Sci Rep 16, 8734 (2026). https://doi.org/10.1038/s41598-026-43831-y
Keywords: liver cancer imaging, multiphoton microscopy, autofluorescence, surgical margins, machine learning