Clear Sky Science · en
Domain-agnostic weakly supervised surgical instrument segmentation
Why smarter views of surgical tools matter
Modern surgeons increasingly operate with the help of cameras, microscopes, and advanced scanners. To guide robots, align 3D views, or hide tools from certain images, computers must reliably find every surgical instrument in each frame, a task called segmentation. Today, that usually demands thousands of painstaking, pixel-perfect annotations from medical experts—and even then, systems often break when the imaging setup or procedure changes. This study introduces a way to let powerful vision models find instruments across very different kinds of medical images, without needing detailed drawings of every tool beforehand.
The challenge of finding tools in many kinds of images
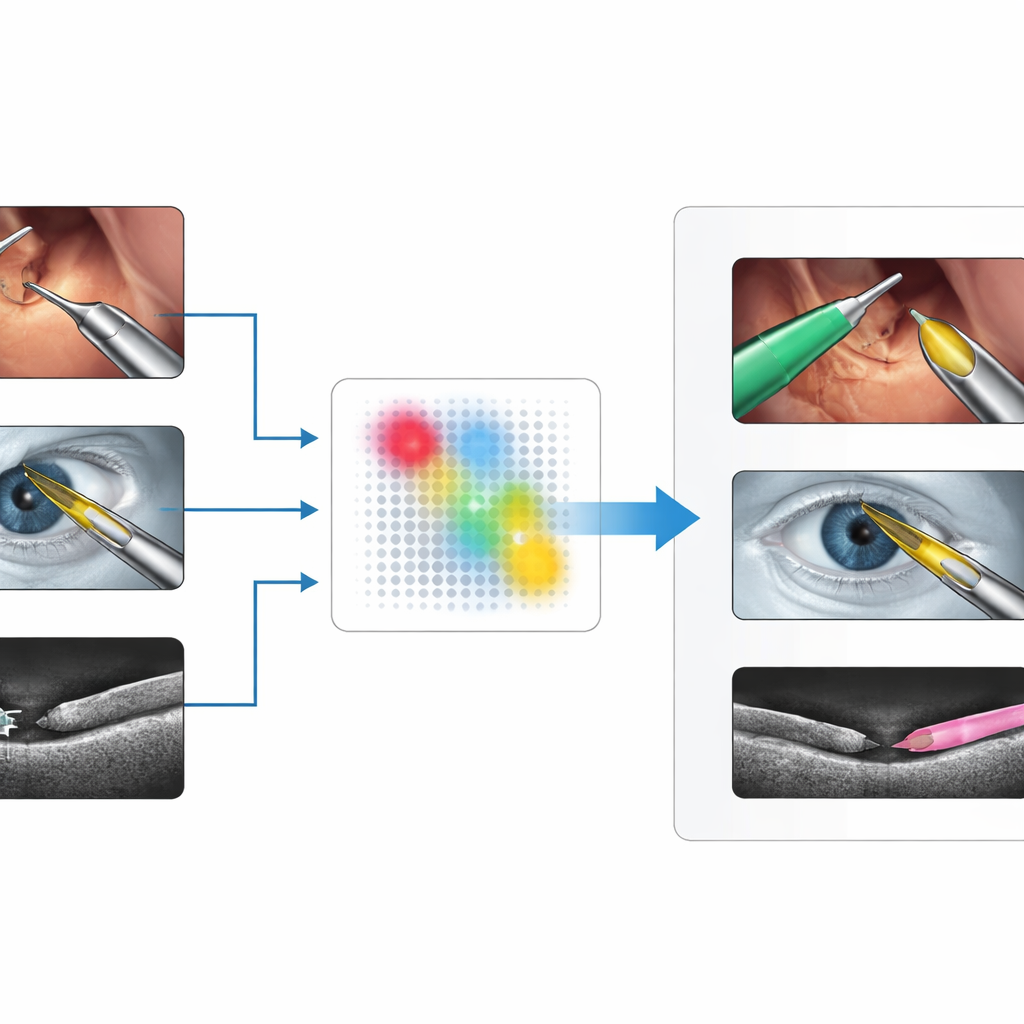
Surgeons use a wide range of imaging systems: color videos from laparoscopic cameras inside the abdomen, microscope views of the eye during cataract surgery, and cross-sectional scans such as optical coherence tomography (OCT) or ultrasound. In each of these, surgical instruments look quite different—shiny metal rods in color images, thin bright lines or crescents in OCT, or speckled blobs in ultrasound. Existing deep-learning methods can work very well, but only after being trained on large, carefully labeled datasets from a specific setting. When the imaging device, anatomy, or instrument type changes, performance often drops sharply, and collecting new annotations is slow, costly, and bound by privacy and expertise constraints.
A new idea: treat instruments as out-of-place objects
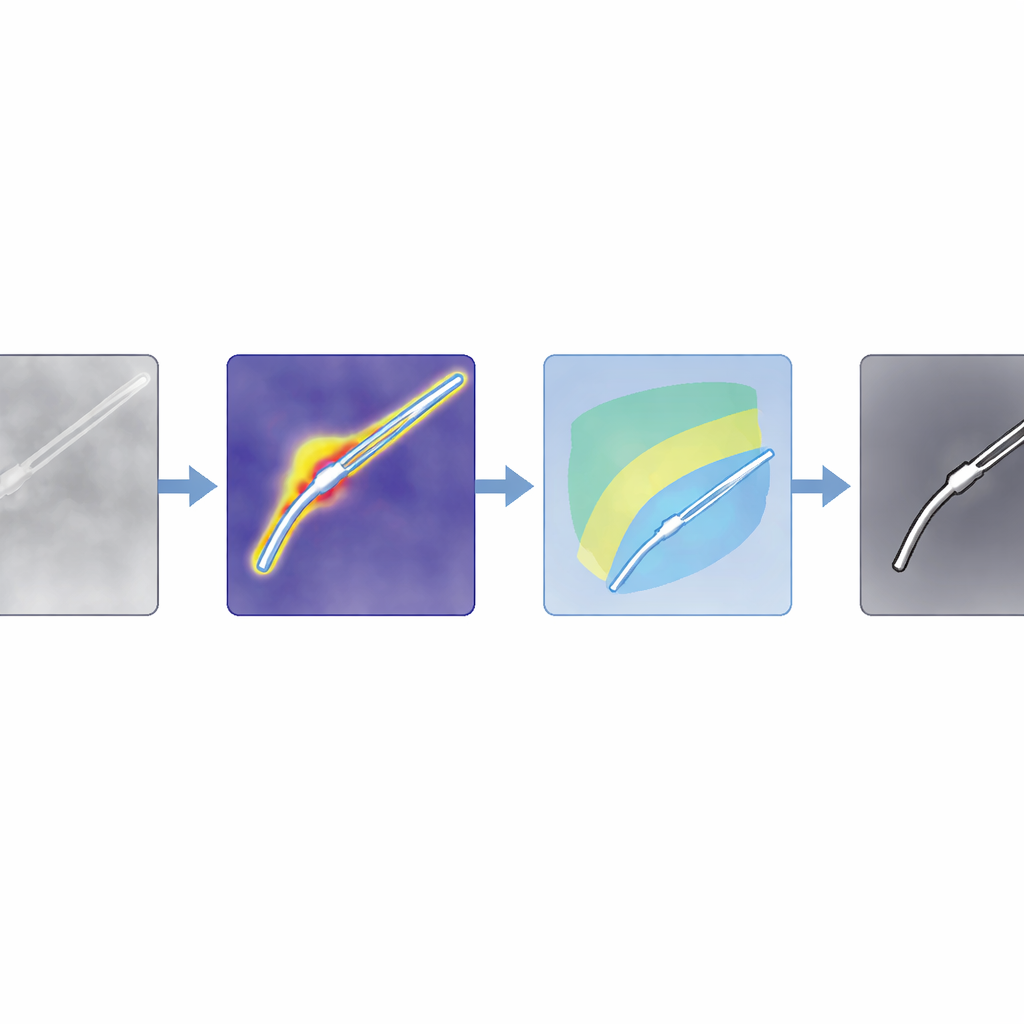
The authors propose a method they call SAM4SIS that turns the problem on its head. Instead of teaching the system exactly what every instrument looks like, they first show it images without any tools, letting it learn what “normal” tissue looks like. They use an anomaly detector called PatchCore to build a memory of these normal patterns. When a new image arrives, PatchCore highlights regions whose appearance does not fit this memory—areas that are likely to contain surgical instruments. This step needs only simple image-level information about whether a tool is present, not pixel-wise drawings of its outline, making setup far easier.
From rough hints to precise outlines
Anomaly maps are coarse, so the team combines them with a powerful foundation model, Segment Anything Model 2 (SAM2), which can draw sharp outlines if given a point inside the object of interest. The key trick is to automatically choose those points from the anomaly map, rather than asking a human to click. The authors design filters tailored for ordinary color images and for intensity-based scans like OCT, boosting regions that are likely to contain instruments while suppressing shadows and bright artifacts. They then score potential tool regions and pick the strongest spots as prompts for SAM2. Because SAM2 returns several candidate outlines, the authors introduce a new scoring rule, SAM4SIS, which measures how well each candidate matches the anomaly-based map and selects the best-fitting mask.
Works across many surgeries and scanners
The researchers test their approach on three demanding datasets: abdominal robotic surgery videos (EndoVis2017), microscope images of cataract operations (CaDIS), and cross-sectional OCT scans of porcine eyes with tiny tools (PASO-SIS). These cover very different views, colors, and noise patterns. Without ever retraining the large segmentation model or drawing new masks, SAM4SIS reaches boundary-accuracy scores between about 53% and 73%, rivaling or surpassing text-based prompting methods and approaching some supervised systems. It performs especially well where traditional methods struggle, such as in OCT and ultrasound data, and needs less than a minute of setup time. The team also shows that the same idea can highlight other foreign objects, like cotton balls in brain ultrasound scans, suggesting that the concept is not limited to instruments.
What this means for future smart surgery
For readers, the core message is that computers can now learn to “segment anything new” in surgical scenes by first understanding what normal tissue looks like and then flagging unfamiliar shapes as likely tools, which are then refined by a general-purpose vision model. This approach avoids heavy annotation work, adapts to different imaging technologies, and can be slotted into surgical workflows with minimal preparation. While carefully trained, specialized models still win when ample labeled data are available, SAM4SIS offers a practical alternative for new procedures, rare imaging setups, or early-stage research, bringing robust, automated instrument detection closer to everyday clinical reality.
Citation: Peter, R., Pham, D.X.V., Matten, P. et al. Domain-agnostic weakly supervised surgical instrument segmentation. Sci Rep 16, 9337 (2026). https://doi.org/10.1038/s41598-026-43054-1
Keywords: surgical instrument segmentation, medical imaging AI, anomaly detection, foundation vision models, robotic surgery