Clear Sky Science · en
Combining tissue-derived microRNAs with clinical risk models for prediction of HCC recurrence after liver transplantation: A proof-of-concept study
Why this research matters to patients and families
Liver cancer is one of the deadliest cancers worldwide, and for some patients the only real chance of cure is to receive a new liver. Even then, the cancer can come back in the transplanted organ, cutting lives short and using up scarce donor livers. This study explores whether tiny molecules inside the removed tumor, called microRNAs, can help doctors better predict whose cancer is most likely to return after liver transplantation so that follow-up care can be more closely tailored to each person.
A closer look at liver cancer after transplant
Hepatocellular carcinoma, the most common form of primary liver cancer, often arises in people with long-standing liver damage from causes like alcohol use, viral hepatitis, or fatty liver disease. For small tumors that cannot be removed safely, liver transplantation offers a powerful two-in-one treatment: it replaces the scarred liver and removes the cancer at the same time. Because donor organs are limited, however, transplant centers rely on strict rules based mainly on tumor size and number—the well-known Milan and UCSF criteria—to decide who is likely to benefit most. Yet even when these rules are followed, 8–20% of patients still see their cancer return, showing that measurements of size alone do not capture how aggressive each tumor really is.
Tiny RNA molecules with big potential
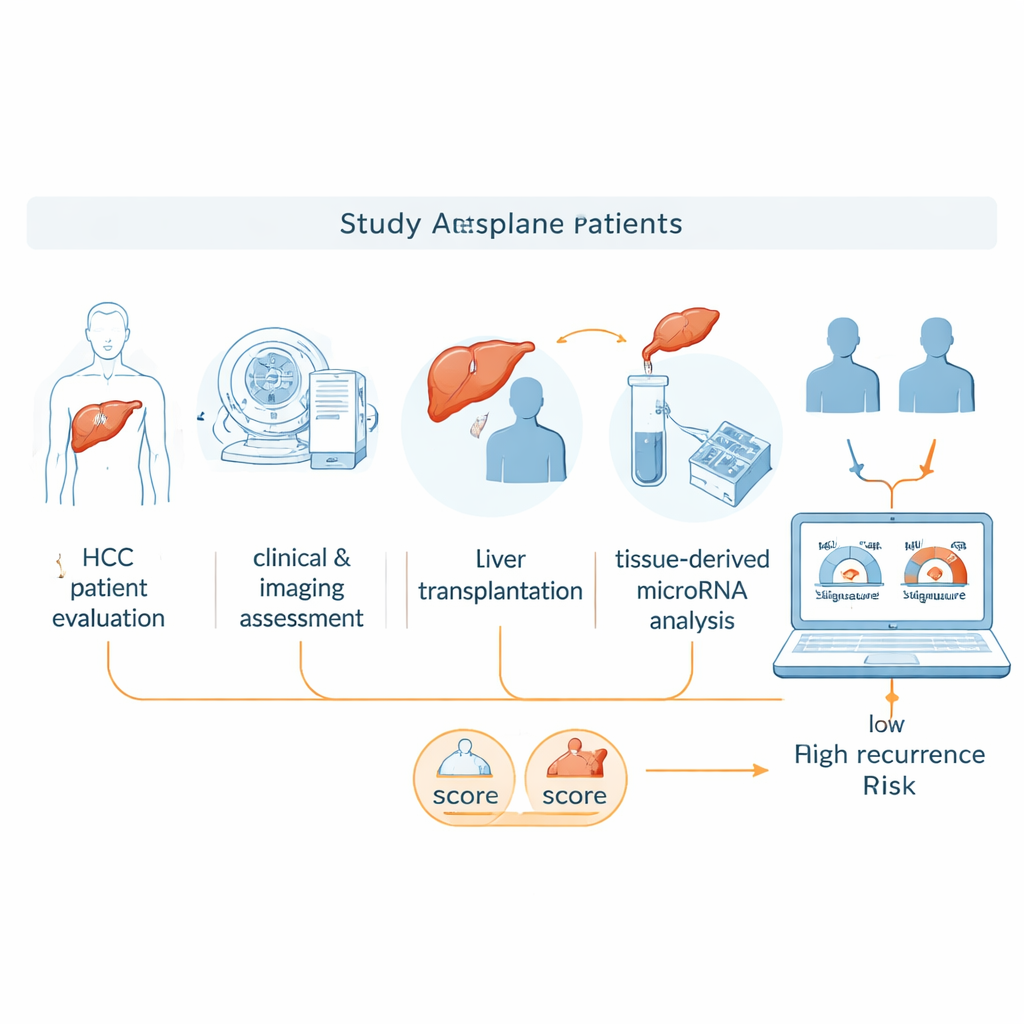
MicroRNAs are short pieces of genetic material that do not make proteins themselves but help control how other genes are switched on and off. Different cancers can have their own microRNA “fingerprints,” and these patterns may reveal how fast a tumor grows or how likely it is to spread. In this proof-of-concept study, the researchers asked whether microRNAs taken from the liver cancer tissue removed at the time of transplantation could be combined with existing clinical rules to sharpen predictions about cancer recurrence. They focused on tissue samples that had been preserved in wax blocks, a routine method used in hospitals, to test whether such molecular testing would be practical in real-world settings.
How the study was done
The team reviewed 20 patients who had liver transplantation for liver cancer between 2007 and 2021. Half of them experienced cancer recurrence after the transplant, while the other half remained free of recurrence for at least five years. All had undergone standard clinical and CT scan assessments before surgery. From each patient’s removed tumor, the scientists extracted RNA and measured the activity of about 3,600 microRNAs using a microarray chip. They first looked for microRNAs whose levels differed strongly between the recurrence and non-recurrence groups, then narrowed the list based on statistical strength and previous evidence linking particular microRNAs to liver cancer behavior.
A three-microRNA warning signal
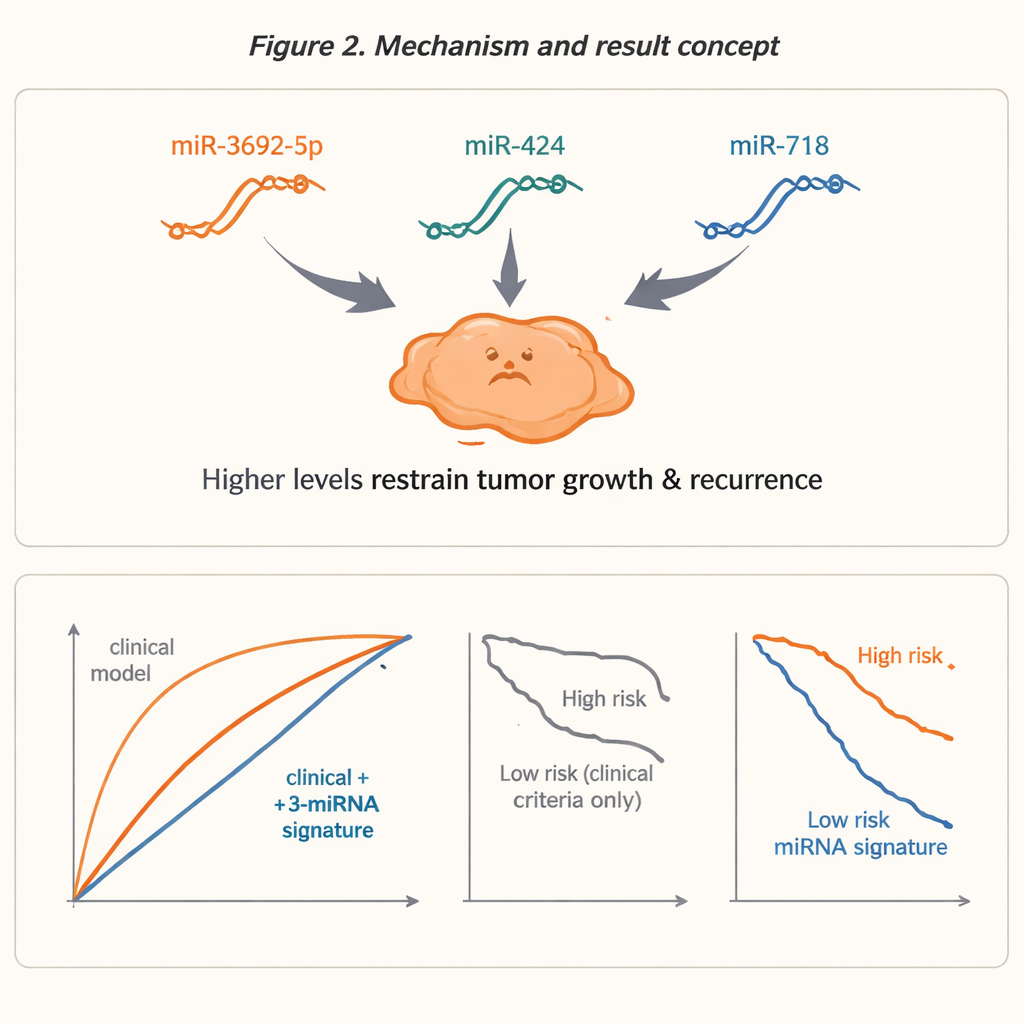
This process highlighted one strongly informative microRNA, called miR-3692-5p, which was noticeably lower in patients whose cancer later returned. Two additional microRNAs—miR-424 and miR-718—were added because earlier studies had tied them to liver cancer outcomes and because they showed similar downward shifts in the recurrence group. Together, this trio formed a “3-miRNA signature” that, on its own, could already distinguish recurrence from non-recurrence in this small group with high accuracy. When the researchers layered this microRNA signature on top of four commonly used clinical models, including the Milan and UCSF criteria and a blood test–based AFP score, the combined models performed far better than the clinical rules alone in predicting who would relapse.
What better prediction looks like in practice
To judge performance, the team used standard statistical tools that measure how well a test separates high- and low-risk patients. The clinical models by themselves only modestly distinguished between these groups. After adding the three microRNAs, the predictive power rose sharply, with accuracy measures approaching 0.94–0.96 on a scale where 1.0 represents a perfect test. The improved models reassigned around half of the patients to a different risk category, suggesting that many people currently labeled as “safe” or “high risk” might be misclassified by size-based rules alone. Importantly, patients labeled high risk by the combined models had much shorter recurrence-free survival—about 17 months versus 38.5 months in one example—showing that these molecular signals tracked meaningfully with real outcomes.
What this means for future care
For non-specialists, the takeaway is that examining the tumor’s internal “control signals,” captured here by three specific microRNAs, may reveal more about its true nature than size or count alone. While this was a small, exploratory study that needs confirmation in larger, multi-center groups and in easier-to-access samples like blood, it demonstrates that such testing is technically feasible on routine hospital tissue. If validated, this approach could help transplant teams choose follow-up scans and treatments more wisely, concentrating the most intensive monitoring and preventive strategies on those whose tumors carry a molecular warning of likely return, and potentially sparing lower-risk patients from unnecessary tests.
Citation: Lederer, T., Lehr, K., Bobe, S. et al. Combining tissue-derived microRNAs with clinical risk models for prediction of HCC recurrence after liver transplantation: A proof-of-concept study. Sci Rep 16, 7742 (2026). https://doi.org/10.1038/s41598-026-41688-9
Keywords: liver cancer, liver transplantation, microRNA biomarkers, cancer recurrence risk, precision oncology