Clear Sky Science · en
Cullin 3 substrate-adaptor protein 1 (MtCSP1) modulates nodulation through interaction with the GTPase ARFA1
How Beans Make Their Own Fertilizer
Modern agriculture leans heavily on nitrogen fertilizers, which boost yields but come with environmental costs. Legume plants such as clover, peas, and alfalfa offer a natural workaround: they host friendly soil bacteria inside special root structures called nodules, where air nitrogen is converted into plant food. This study uncovers how a previously uncharacterized plant protein, MtCSP1, helps control the formation of these nodules by fine-tuning the internal traffic and recycling of key molecular switches inside root cells.
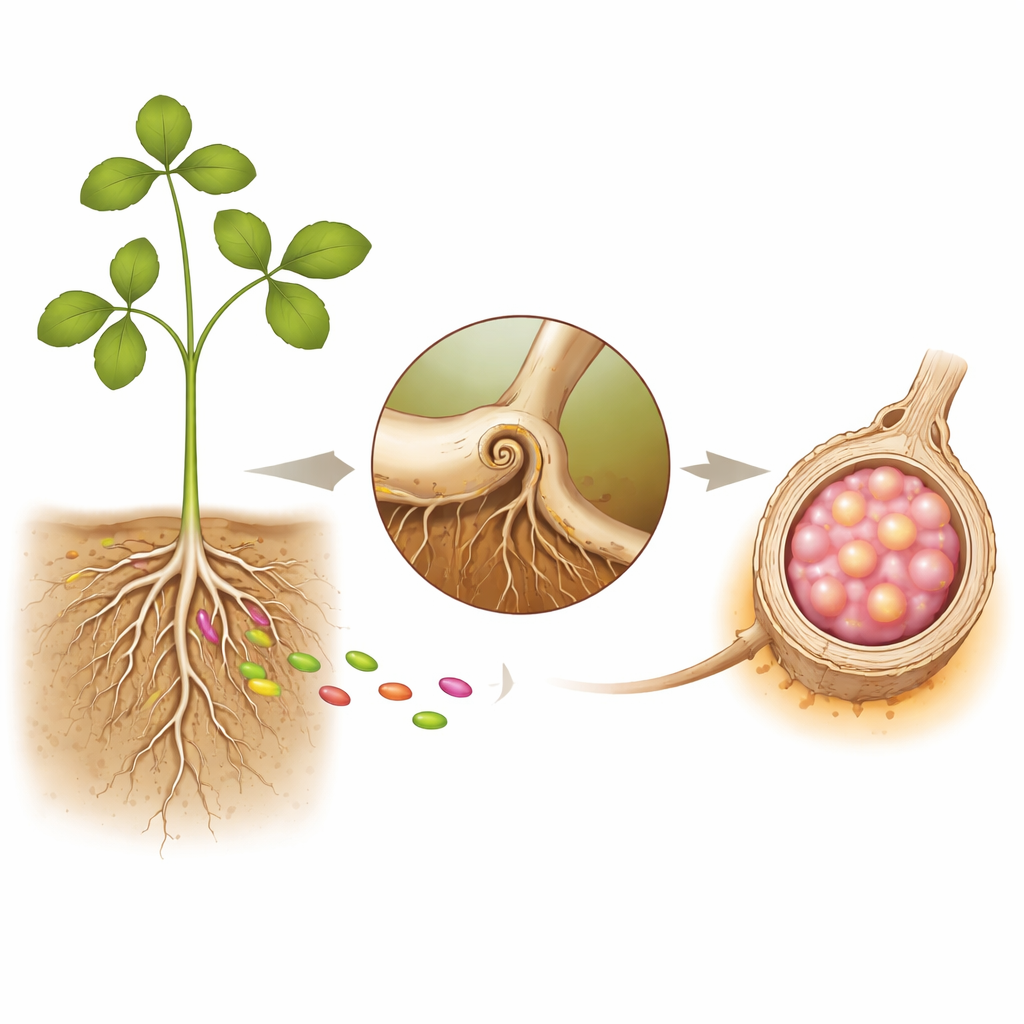
Plant Roots and Their Bacterial Houseguests
Legumes partner with rhizobia, soil bacteria that can turn nitrogen gas from the atmosphere into usable forms like ammonium. To house these bacteria safely, roots build nodules—tiny factories on the root surface. The process begins when bacteria attach to root hairs, which curl and form a narrow tunnel called an infection thread. Through this tunnel the bacteria travel into deeper root tissues, where a new organ, the nodule, starts to grow. Inside the mature nodule, bacteria are wrapped in plant-made membranes and become specialized “bacteroids” that fix nitrogen for the plant, in exchange for sugars and energy.
Cellular Traffic Cops Inside Root Cells
Building infection threads and nodules requires intense remodeling of plant cell membranes. This remodeling depends on vesicles—small membrane bubbles that ferry materials inside cells. Their movement and timing are controlled by small molecular switches known as GTPases. One such switch, ARFA1, helps shape and direct vesicles in many organisms. But in the context of nitrogen-fixing nodules, it was unclear how ARFA1 is regulated or how long it remains active before being removed.
A New Protein Link Between Traffic and Recycling
The researchers searched for plant proteins that physically interact with ARFA1 in the model legume Medicago truncatula. Using yeast two-hybrid screens and protein interaction tests in plant leaves, they identified MtCSP1, a protein that belongs to a family of adaptors that link specific targets to a cellular “shredder” called the ubiquitin system. MtCSP1 carries a BTB/POZ domain, a signature motif of proteins that recruit chosen partners to Cullin 3–based complexes, which tag proteins for destruction. Fluorescence imaging showed that MtCSP1 and ARFA1 meet on vesicles located in late endosomes—cellular waystations that often direct cargo toward degradation.
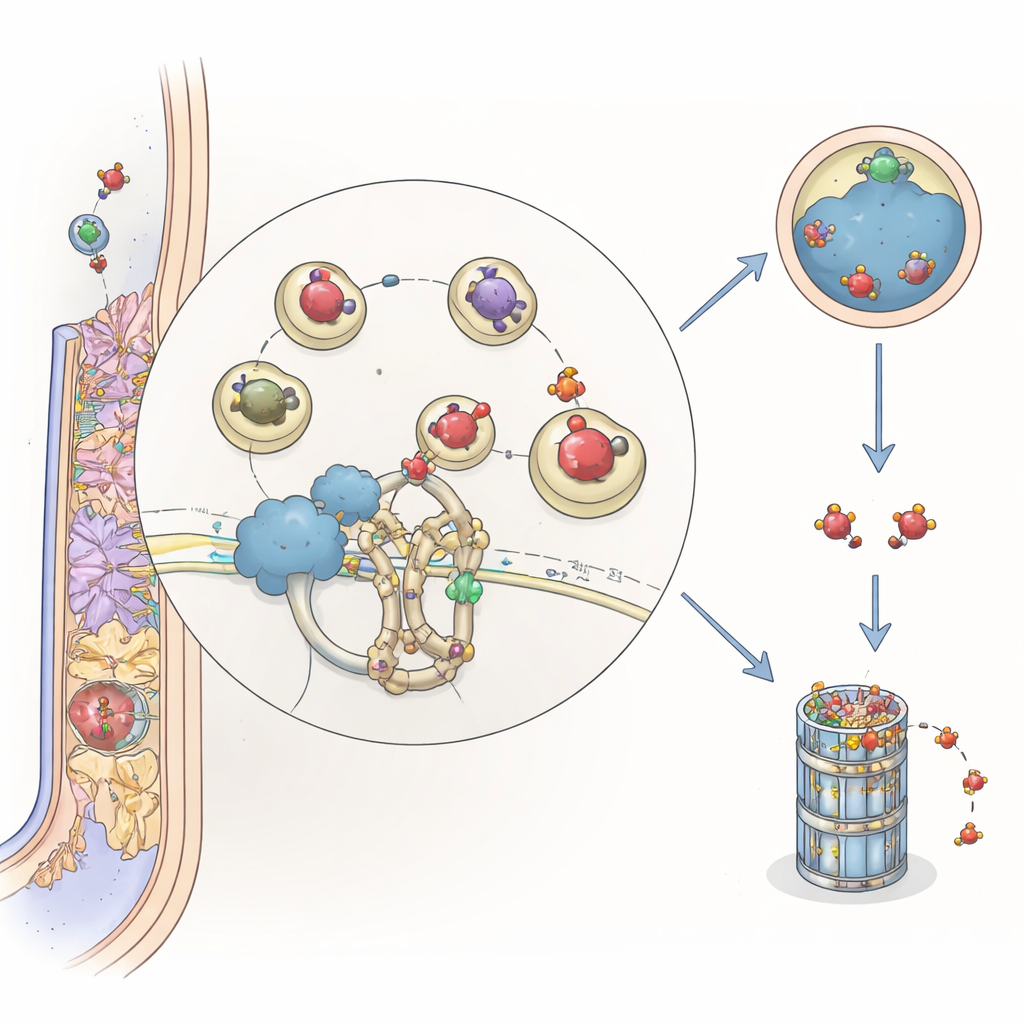
When and Where the New Player Acts
To understand when MtCSP1 is used, the team tracked the activity of its promoter, the DNA switch that controls its production. They found that MtCSP1 is switched on in root tips, in emerging lateral roots, and in key regions of developing and mature nodules, particularly the meristem (the growth zone) and infection regions. Public gene-expression datasets revealed that MtCSP1 and ARFA1 are often activated together, hinting that the plant coordinates the presence of the switch (ARFA1) and the adaptor (MtCSP1) during organ formation in roots.
Tuning Nodule Number and Infection Progress
The scientists then altered MtCSP1 levels in roots. When MtCSP1 was silenced by RNA interference, plants formed fewer nodules over time, and many infection threads stalled in the root hairs instead of reaching inner root layers. However, the nodules that did form were of normal size, indicating that MtCSP1 mainly affects the initiation and progression of infection, rather than later growth. Conversely, when MtCSP1 was overproduced, roots developed more nodules and showed changes in how infection events advanced, again without major changes in nodule size or shape. These results indicate that MtCSP1 is not needed to start infection threads, but is crucial for their proper progression and for the successful onset of new nodules.
Recycling Switches to Control Symbiosis
Putting the pieces together, the authors propose that MtCSP1 acts as a guide that brings ARFA1, in its active or inactive forms, to the Cullin 3 ubiquitin machinery on late endosome vesicles. There, ARFA1 can be tagged and sent either to the vacuole for destruction or to the proteasome, helping shut down or adjust vesicle traffic once it has served its purpose. By regulating how long ARFA1 remains available, MtCSP1 allows the plant to fine-tune the delicate choreography of infection threads and nodule formation. For a layperson, the message is that legumes use an internal recycling system to control tiny molecular traffic lights inside their roots—ensuring that their natural fertilizer factories are built efficiently and at the right time.
Citation: Rípodas, C., Cretton, M., Eylenstein, A. et al. Cullin 3 substrate-adaptor protein 1 (MtCSP1) modulates nodulation through interaction with the GTPase ARFA1. Sci Rep 16, 8938 (2026). https://doi.org/10.1038/s41598-026-41112-2
Keywords: nitrogen fixation, legume nodules, protein degradation, vesicle trafficking, plant–microbe symbiosis