Clear Sky Science · en
Early detection of chronic kidney disease based on a SURD-enhanced machine learning model
Why catching kidney trouble early matters
Chronic kidney disease often creeps up silently, showing few warning signs until the kidneys are badly damaged. Yet simple blood and urine tests can reveal problems years earlier, when treatment can slow or even prevent serious decline. This study explores a new way to sift through those routine test results using advanced, but interpretable, computer models so that people at high risk can be flagged sooner and doctors can understand why.
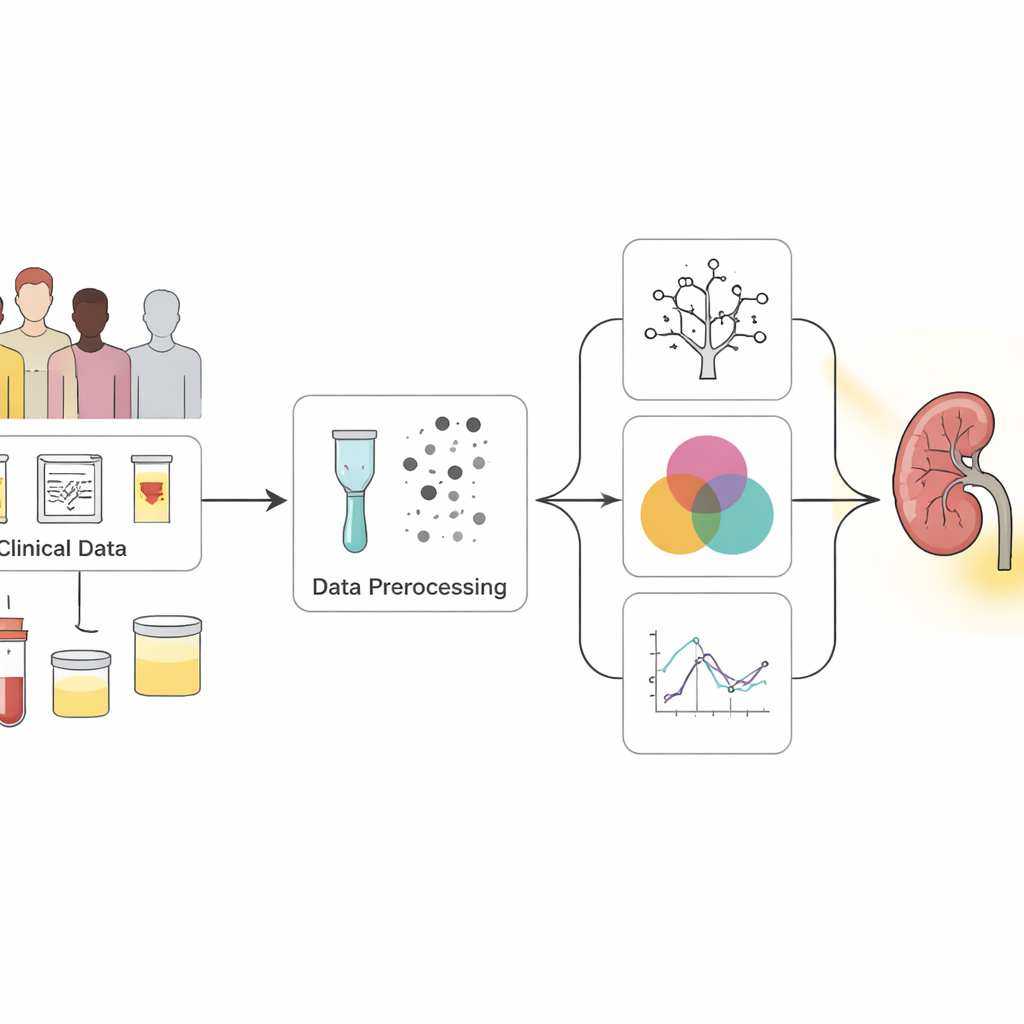
Turning messy checkup data into clear signals
The researchers started with a widely used public dataset of 400 people, most of whom had already been diagnosed with chronic kidney disease. Each person had 25 measurements, ranging from blood pressure and blood counts to urine findings and medical history such as diabetes and high blood pressure. Many entries were incomplete, so the team used careful statistical techniques to fill in missing values rather than simply discarding patients. They also balanced the data so that healthy and diseased cases were more evenly represented, helping the computer models learn to recognize both groups fairly.
Looking beyond simple correlations
Most medical prediction tools treat each test result separately: they look at how strongly one measurement, such as blood sugar, is linked to disease. But in the body, risk factors rarely act alone. Some tests convey nearly the same information, while others only become informative in combination. To capture this, the authors used a framework called SURD that breaks down each feature’s contribution into three parts: information that is shared with other tests, information that is unique, and information that only appears when features work together. This allowed them to group lab values and clinical findings into “unique,” “redundant,” and “synergistic” sets before feeding them to the prediction models.
Teaching many models and choosing the most reliable
With these SURD-based feature groups in hand, the team trained ten different machine learning models, from simple decision trees to more complex approaches like random forests and neural networks. They compared performance when models used all available features versus only a combined set of unique and synergistic ones. Across almost all model types, this trimmed, SURD-guided feature set performed as well as or better than the full collection of 25 variables, often improving the balance between correctly identifying sick patients and avoiding false alarms. In particular, tree-based models such as random forests and boosted trees achieved nearly perfect scores on the original dataset.
Testing the method in real-world hospital data
Excellent performance on a small benchmark dataset can be misleading if a model fails when exposed to more varied patients. To guard against this, the authors validated their approach using a much larger hospital database of over 27,000 intensive-care patients. Here, the random forest model built on the SURD-selected features still distinguished patients with and without kidney disease with extremely high accuracy. Its performance clearly exceeded that of a simpler decision tree, indicating that the method can generalize beyond a carefully curated research dataset to messier real-world records.
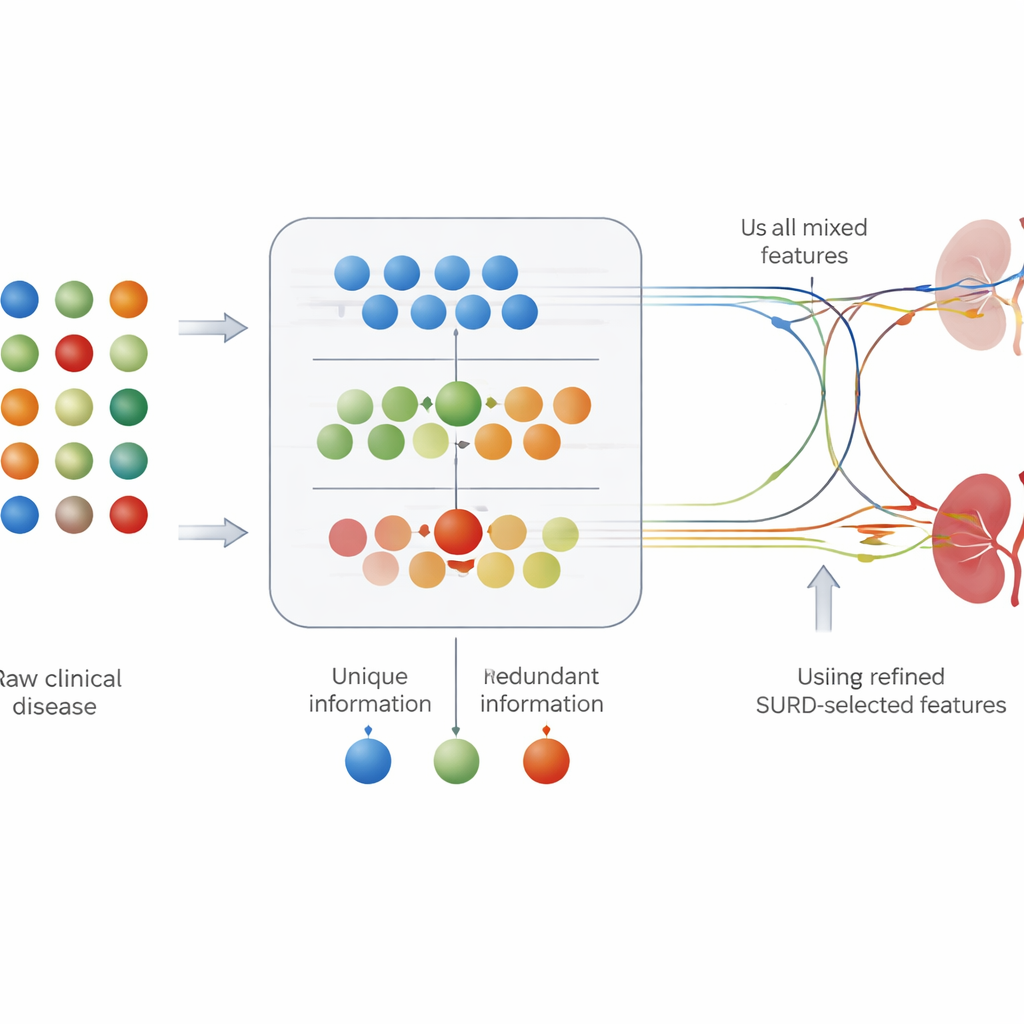
Seeing which tests matter and how
Accuracy alone is not enough for clinical use; doctors also need to know which test results are driving a prediction. The study combined SURD with modern explanation tools that assign each feature a contribution to the model’s decision for a given patient. This analysis highlighted familiar risk markers, such as serum creatinine (a direct indicator of kidney function), hemoglobin levels, urine concentration, and the presence of diabetes or high blood pressure. Interestingly, SURD showed that some of these factors mostly act in concert with others, while creatinine stands out as a strongly informative signal on its own. Together, these techniques offer both a global view of which tests the model relies on and patient-level breakdowns of why a particular person is predicted to be at high risk.
What this means for everyday care
In plain terms, the study shows that it is possible to build a kidney disease risk calculator that is both highly accurate and reasonably transparent. By separating overlapping from truly unique information in routine lab and history data, the SURD-guided models make sharper predictions without becoming a mysterious black box. Although further work is needed in broader and more diverse patient groups, this approach could eventually help clinicians spot kidney problems earlier, focus attention on the most informative tests, and explain to patients in straightforward terms which aspects of their health are putting their kidneys at risk.
Citation: Xue, N., Bai, T., Jia, X. et al. Early detection of chronic kidney disease based on a SURD-enhanced machine learning model. Sci Rep 16, 10444 (2026). https://doi.org/10.1038/s41598-026-41050-z
Keywords: chronic kidney disease, kidney risk prediction, medical machine learning, explainable AI, electronic health records