Clear Sky Science · en
Evaluating resolution requirements for subtle caenorhabditis elegans strain discrimination using classical descriptors and CNN–transformer models
Why tiny worms and sharp pictures matter
Scientists often use a microscopic worm called Caenorhabditis elegans to study how genes, aging, and drugs affect the nervous system. Many worm strains look and move almost the same to the naked eye, yet those tiny differences can reveal how their brains and muscles are working. This study asks a practical question: how sharp do our images really need to be to spot such subtle changes in movement, and when do modern artificial intelligence tools actually benefit from higher resolution?
Watching worms from far and near
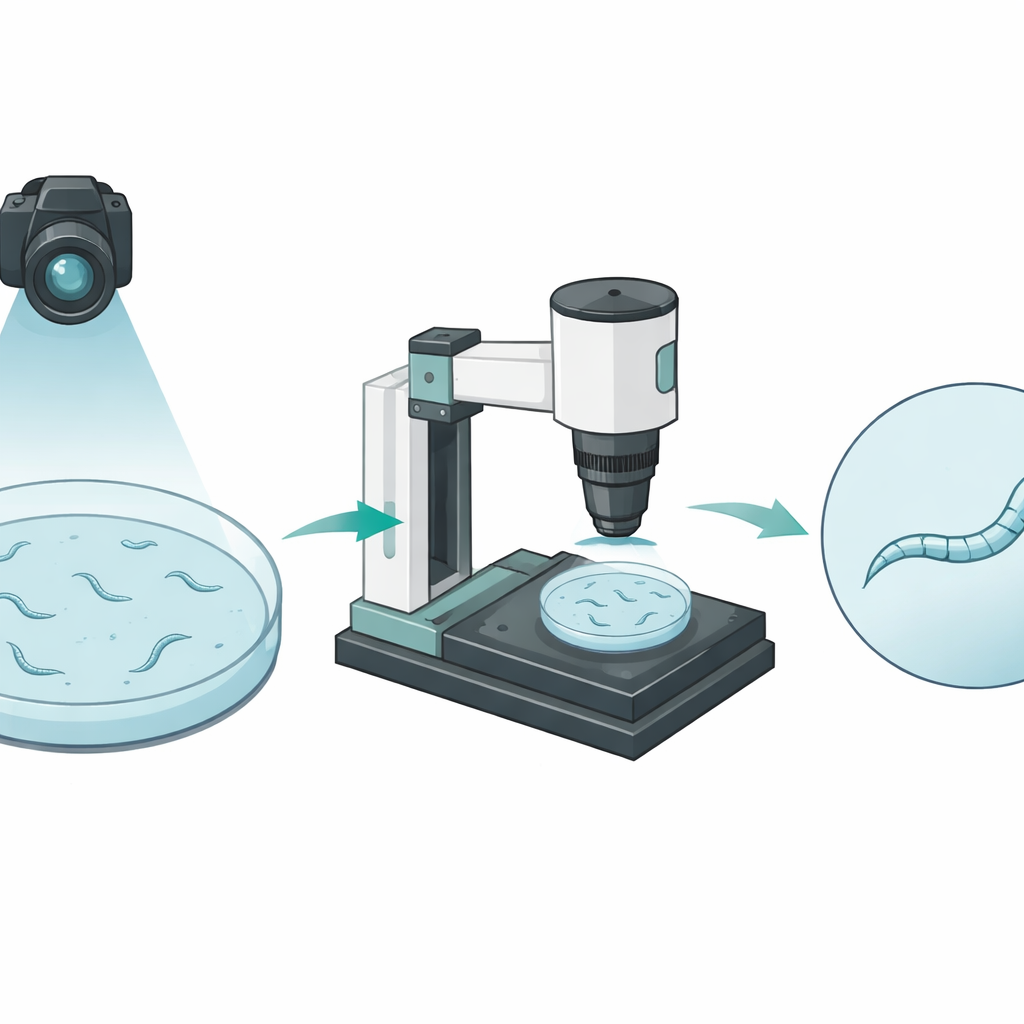
The researchers built an automated imaging platform that views worms at two very different scales. A pair of cameras first looks at an entire petri dish from above, following many worms as they crawl around. This wide view captures how far each animal travels but shows each worm as only a few pixels wide, like a stick figure viewed from across a room. A separate motorized microscope can then zoom in on one chosen worm, keeping it centered and in focus for a full minute. In these close-up movies, the worm’s body spans dozens of pixels in width, revealing fine bends and shape changes as it moves.
Simple measurements hit a wall
To compare what each view could reveal, the team recorded three kinds of worms. One was the standard wild-type strain used as a reference. A second was a mutant with extremely clumsy movement that is easy to spot. The third was a specially engineered strain with only very mild motor problems, known to be hard to distinguish from the reference strain even by eye. From both the wide and close-up recordings, the researchers extracted traditional measures such as how far each worm traveled, how fast it moved, and how its body shape changed over time. As expected, both views clearly separated the very clumsy mutant from the other two strains. However, none of these standard measurements, alone or together, could reliably tell apart the subtly altered worms from the normal ones.
Letting deep learning read the motion
Next, the authors turned to a more flexible approach: a deep learning model that watches the actual image sequence instead of hand-picked measurements. Each frame was first passed through a convolutional neural network that learned to encode the worm’s appearance. These frame-wise features were then fed into a Transformer module, which looked at how posture evolved over the 60-second clip. When this model was trained on the low-detail, dish-wide videos, it performed no better than chance at separating the subtle strain from the reference. But when trained on the high-detail microscope recordings, it consistently classified the two strains with around three-quarters accuracy, uncovering movement patterns too faint for standard descriptors to capture.
How much detail is enough?
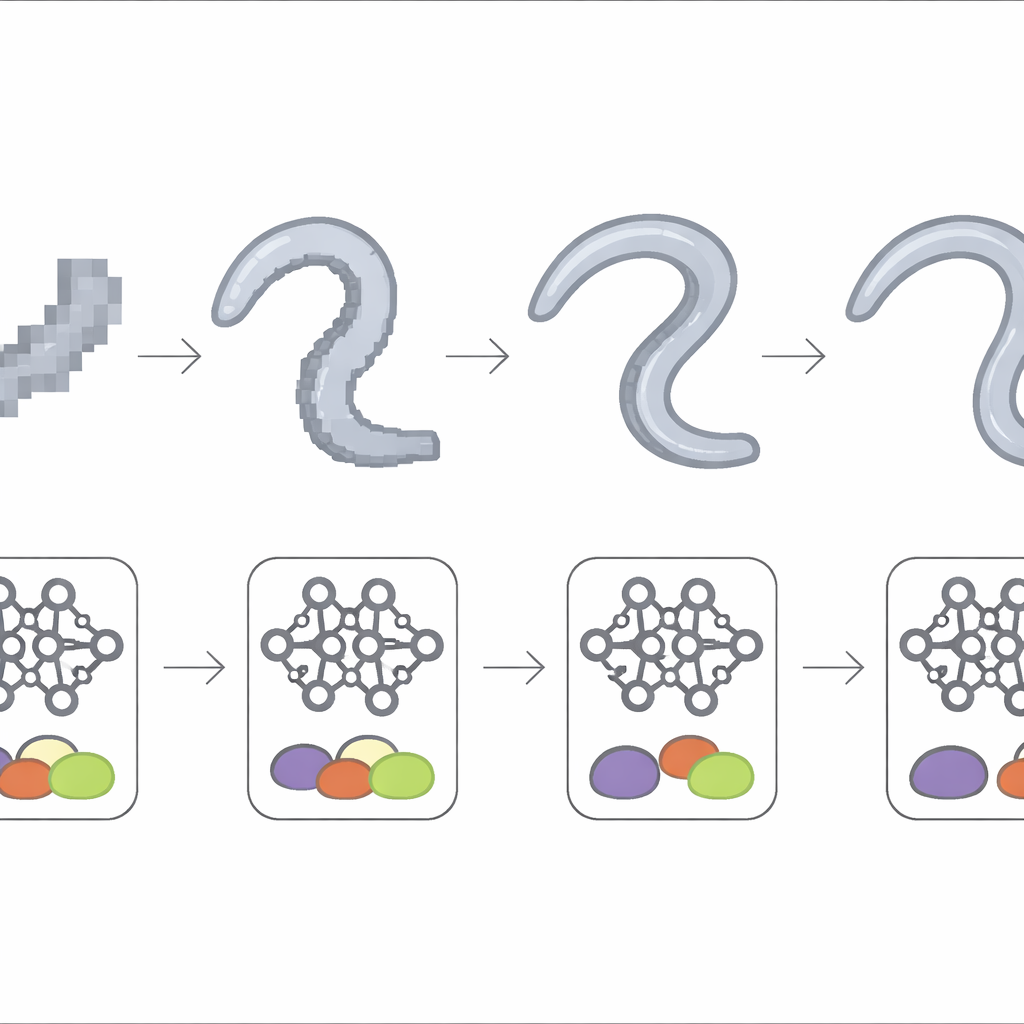
To pin down the role of image sharpness, the team gradually blurred the microscope recordings by shrinking their size by factors of two, four, eight, and sixteen, retraining the same deep model each time. Performance stayed high when the worm’s body still spanned a few dozen pixels in width, meaning the model could tolerate moderate loss of detail. Once the worm shrank to only about ten pixels wide or less, accuracy dropped sharply and became unstable from experiment to experiment. At the coarsest scales, results approached those from the dish-wide view and from the simple statistical methods, indicating that the subtle signatures of the mild motor defect had effectively vanished from the images.
What this means for future worm studies
For experiments that must distinguish only clear-cut movement defects, a broad, low-resolution view appears to be enough, and classic measurements of distance and speed work well. But when the goal is to detect slight changes in how worms bend and coordinate their bodies—such as those caused by mild genetic changes or gentle drug effects—this work shows that both high-resolution imaging and sequence-based deep learning models are needed. In plain terms, to see the quiet whispers of disease or treatment effects in these tiny animals, we must not only look closely enough but also use tools smart enough to read the subtle patterns encoded in their motion.
Citation: Peñaranda-Jara, JJ., Escobar-Benavides, S., Puchalt, JC. et al. Evaluating resolution requirements for subtle caenorhabditis elegans strain discrimination using classical descriptors and CNN–transformer models. Sci Rep 16, 8664 (2026). https://doi.org/10.1038/s41598-026-40784-0
Keywords: C. elegans locomotion, phenotype classification, image resolution, deep learning, behavioral tracking