Clear Sky Science · en
Comparative plastid genomics of Hippophae reveals phylogenetic relationships and provides candidate DNA markers for taxonomic identification
Why this hardy shrub matters
Sea buckthorn is a tough shrub that thrives where many other plants fail: on cold, dry, windy slopes of the Qinghai–Tibet Plateau and beyond. Its bright orange berries are promoted worldwide as “superfruits,” and the plant is widely used to stabilize soils and restore damaged land. Yet even experts struggle to tell its closely related species and subspecies apart just by looking at them. This study asks a simple but important question: can we read the plant’s internal instruction manual—its DNA—to sort out who is who, and in the process give breeders and conservationists a powerful new way to manage this valuable resource?
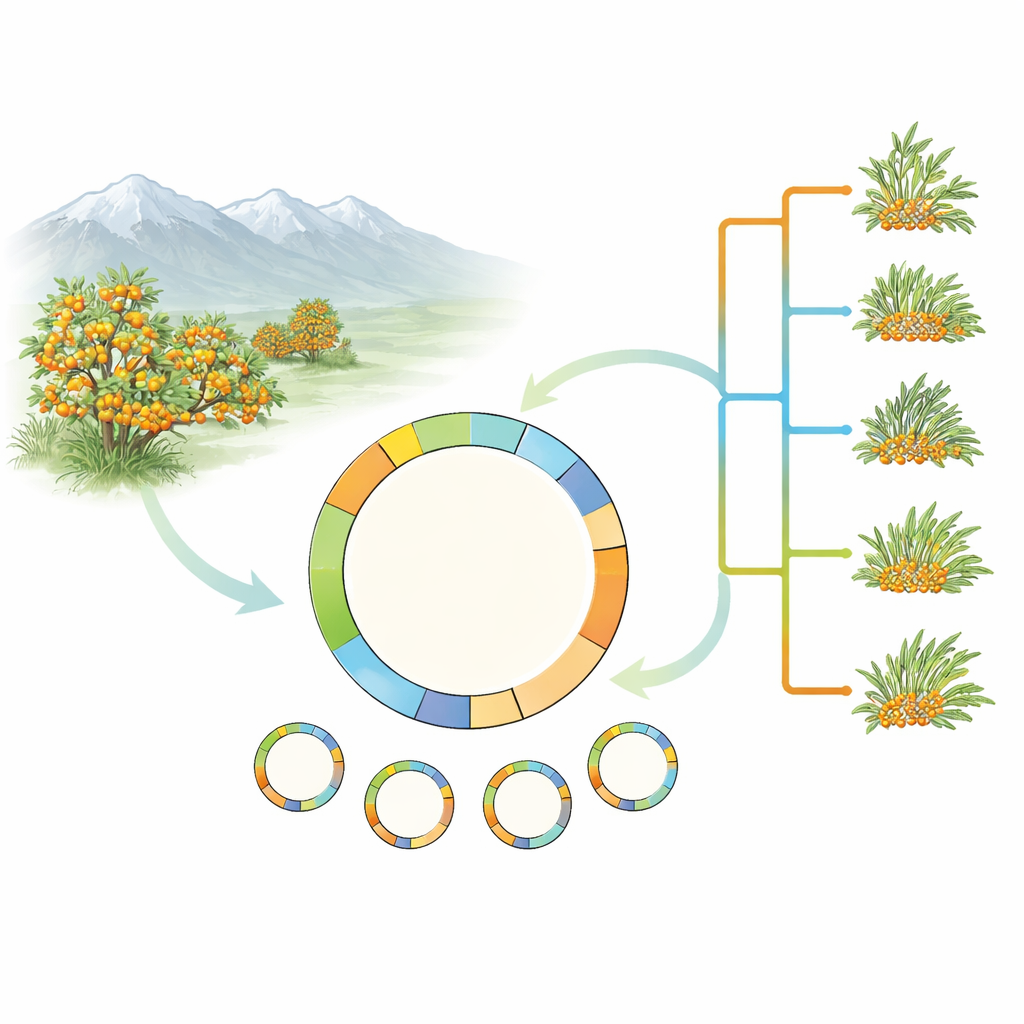
Looking inside the plant’s green engines
Rather than tackling the full complexity of the sea buckthorn genome, the researchers focused on the plant’s plastids—tiny green compartments inside cells that carry out photosynthesis. Plastids have their own small circular DNA molecule, inherited mostly from the mother, which has proved extremely useful for tracing plant family trees. The team collected and sequenced complete plastid genomes from 17 samples covering five sea buckthorn species and several forms of the widespread Hippophae rhamnoides. They also added a newly assembled plastid genome from an important cultivated type, H. rhamnoides subsp. mongolica cv. Prevoskhodnaya, and carefully rechecked earlier database entries for errors.
A shared blueprint with telling differences
At first glance, the plastid DNA of all sea buckthorn samples looked remarkably alike. Each genome was about 155,000 to 156,000 “letters” long and followed the same four-part layout found in many flowering plants: two single-copy regions separated by a pair of duplicated segments. The same set of genes was present and arranged in the same order, and even the overall balance of the four DNA letters barely varied. This structural stability suggests that, over evolutionary time, the plastid blueprint in Hippophae has been conserved. Yet when the researchers zoomed in on finer details—such as how often certain DNA “codons” are used to spell the same amino acid—they found subtle, lineage-specific patterns hinting at slow, long-term shaping of the code in different branches of the genus.
Untangling the family tree
Using 78 protein-coding genes from the plastid DNA, the team built evolutionary trees that place sea buckthorn in the wider rose-plant family and then zoom in on relationships within Hippophae. The analyses confirmed that sea buckthorn as a group forms a single natural lineage, and that H. rhamnoides and its subspecies are also tightly knit. Intriguingly, one species, H. tibetana, consistently falls inside the H. rhamnoides group in the plastid tree, even though earlier work with nuclear DNA had placed it nearer the base of the genus. This mismatch between nuclear and plastid histories hints at past hybridization or other complex evolutionary events and points to the need for future studies that combine whole nuclear and plastid datasets.
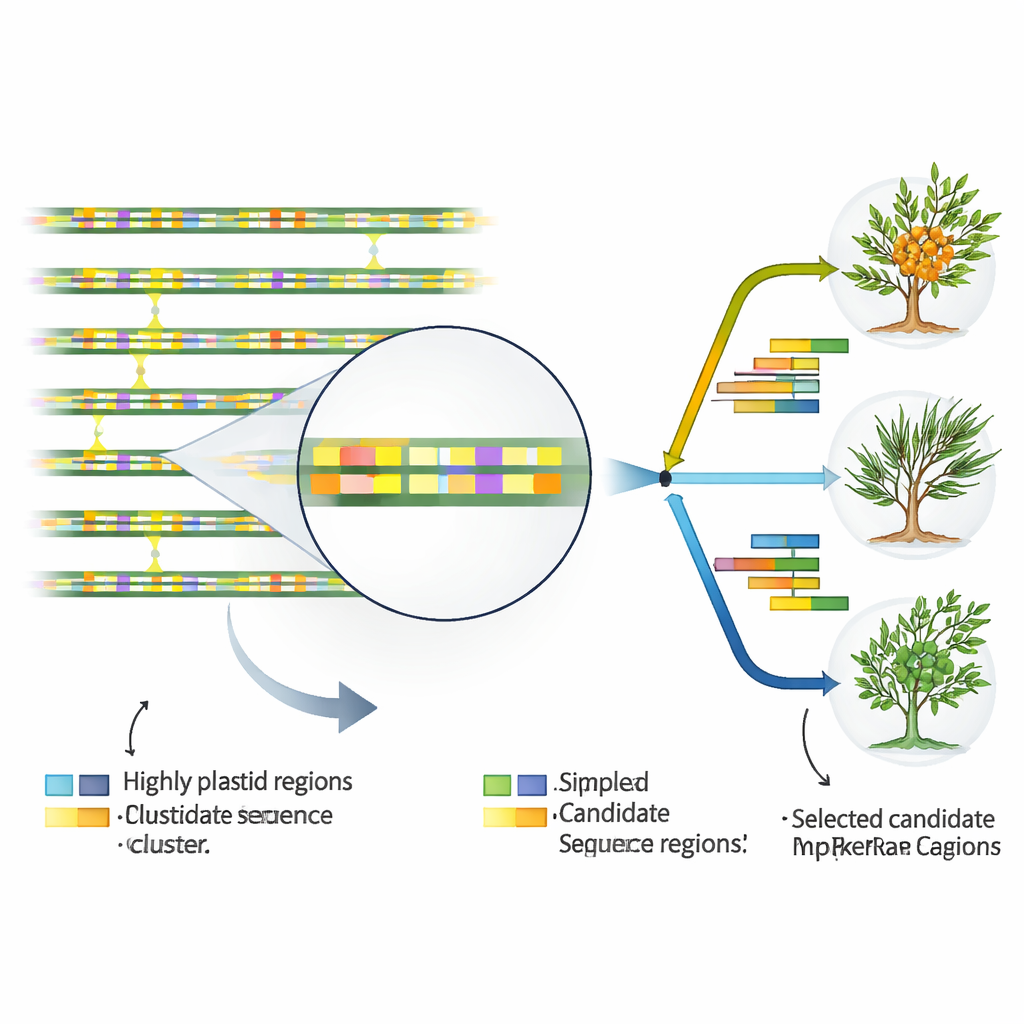
Finding the genomic hotspots that mark identity
To turn plastid DNA into everyday tools for taxonomists and breeders, the authors searched for stretches of sequence that change more quickly than the rest. Comparing all 17 plastid genomes, they pinpointed 46 especially variable regions, almost all lying between genes or within noncoding introns rather than in gene bodies themselves. They also mapped dozens of tiny repeated motifs known as simple sequence repeats, which are particularly rich in the letters A and T and cluster in noncoding regions. Some of these repeats and variable segments showed clear differences among species and even among subspecies of H. rhamnoides. A few regions stood out as hotspots in both species-level and subspecies-level comparisons, making them prime candidates for practical DNA markers that can be targeted with simple laboratory tests.
From DNA patterns to real-world uses
By charting both the stable framework and the small but informative differences in sea buckthorn plastid genomes, this study delivers a toolbox for reliable DNA-based identification. The proposed marker regions could help distinguish look-alike species, verify the origin of commercial berry products, protect wild genetic diversity, and guide the selection of parents in breeding programs aimed at nutrition, medicine, and land restoration. In plain terms, the authors show that a carefully read chloroplast instruction manual can reveal who is who in this rugged shrub’s extended family, clearing the way for smarter conservation and more targeted use of one of the world’s hardiest fruit crops.
Citation: Asakura, N., Noda, M., Takahashi, Y. et al. Comparative plastid genomics of Hippophae reveals phylogenetic relationships and provides candidate DNA markers for taxonomic identification. Sci Rep 16, 7943 (2026). https://doi.org/10.1038/s41598-026-40776-0
Keywords: sea buckthorn, plastid genome, DNA markers, plant taxonomy, genetic diversity