Clear Sky Science · en
A splicing-derived microRNA from amelogenin exon4 regulates enamel formation via control of exon4 splicing and amelogenin expression
Why a tiny RNA matters for strong teeth
Tooth enamel is the hardest substance in the human body, yet it can be surprisingly fragile when its formation goes awry. This study uncovers how a very small piece of genetic material, a microRNA called miR‑exon4, helps tooth‑forming cells build properly hardened enamel. By showing that this microRNA fine‑tunes both the main enamel protein and the timing of mineral deposition, the work links subtle RNA processing inside cells to visible enamel defects similar to those seen in a hereditary condition called amelogenesis imperfecta.
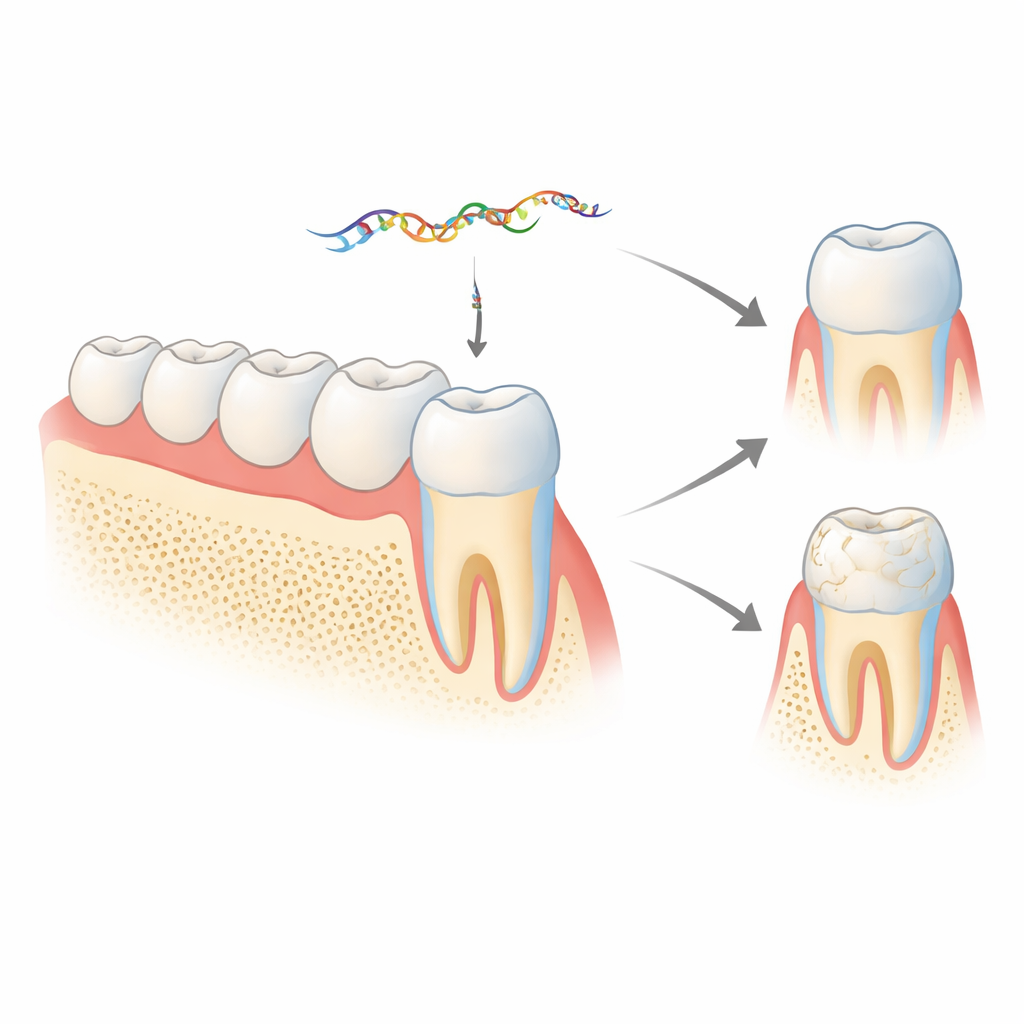
A hidden message inside an enamel gene
Enamel is built largely from a protein called amelogenin, produced by cells known as ameloblasts. The amelogenin gene (Amelx in mice) can be cut and pasted in different ways, creating several protein versions that are needed at different stages of tooth development. One short segment, called exon 4, is usually removed from the final protein‑coding message. Earlier work from this group showed that the discarded exon 4 is not waste: it is processed into a microRNA, miR‑exon4, which can regulate other genes important for bone and enamel. The new study asks what happens in living animals when this microRNA is reduced or blocked, and whether it also feeds back to control how amelogenin itself is assembled.
A regulatory chain inside tooth‑forming cells
The researchers first confirmed in mouse teeth that miR‑exon4 participates in a regulatory chain they had previously mapped in cells grown in the lab. In normal enamel organs, miR‑exon4 keeps two upstream genes, Nfia and Prkch, in check. When these are held low, levels of a key transcription factor, RUNX2, rise. Using mice that either lacked the amelogenin gene, received extra miR‑exon4, or were treated with a miR‑exon4 blocker, the team showed that lowering miR‑exon4 boosts Nfia and Prkch and reduces RUNX2, while adding miR‑exon4 has the opposite effect. This confirmed that the miR‑exon4–Nfia/Prkch–RUNX2 pathway operates in vivo within developing teeth.
From disrupted signals to weaker enamel
To see how these molecular shifts affect actual enamel, the scientists inhibited miR‑exon4 in mouse pups for one week during active tooth formation. Three‑dimensional X‑ray imaging revealed that treated animals had a clear drop in highly mineralized enamel in both incisors and molars. Heat maps and stained sections showed that the start of mineral buildup along the enamel layer was delayed and that the early mineralization phase was shortened, leading to rougher surfaces and blurred boundaries between enamel and underlying tissues. At the same time, RUNX2 protein levels in ameloblasts fell, while amelogenin protein—including versions that contain exon 4—increased. This pattern mirrors earlier models in which overproduction of a long amelogenin form with exon 4 leads to enamel defects, suggesting that too much of this isoform, triggered by loss of miR‑exon4, can directly disturb normal mineralization.
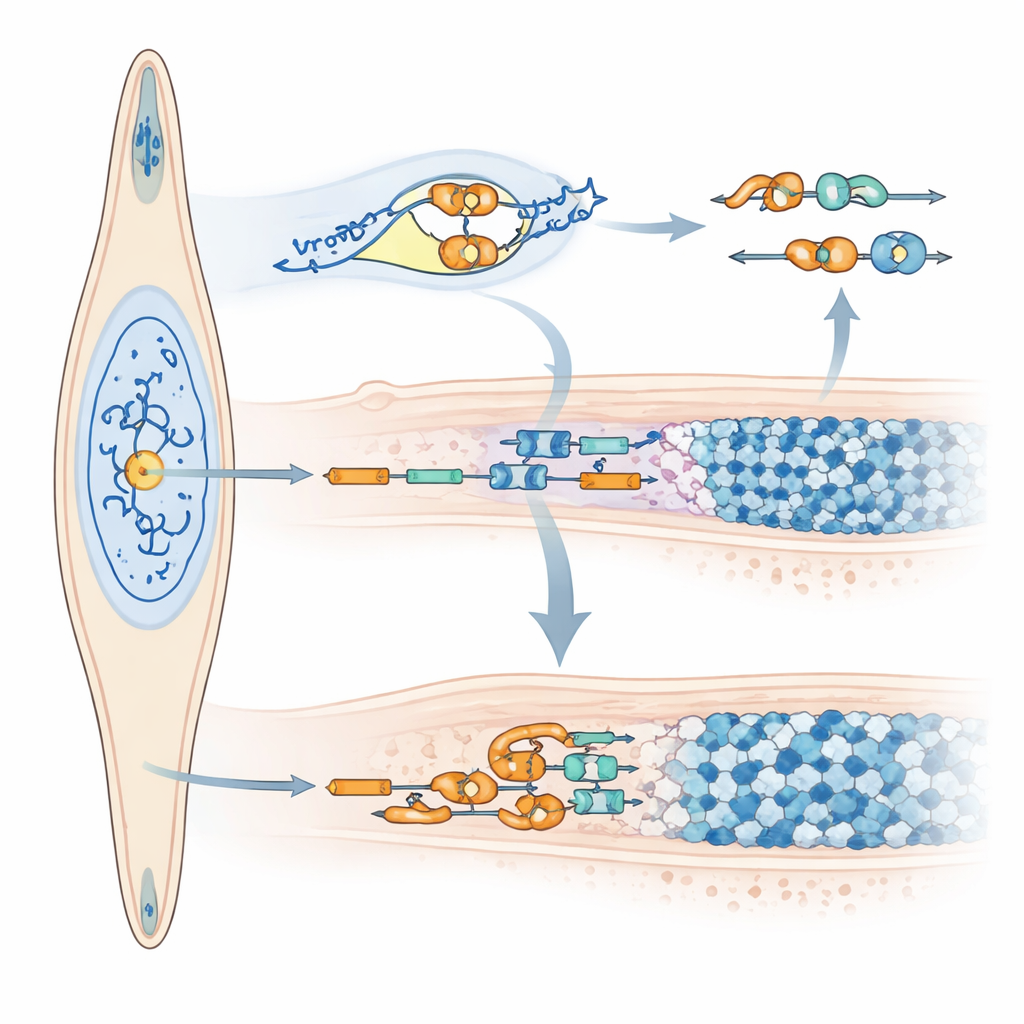
How the microRNA reshapes the enamel message
Beyond changing how much amelogenin is made, miR‑exon4 also alters how the amelogenin message is cut and spliced. Short‑term blocking of miR‑exon4 reduced RNA molecules that still contained exon 4 without changing total amelogenin levels, indicating that exon 4 was being removed more often. The team linked this shift to changes in several splicing regulator genes (SRSFs), with some moving up and others down when miR‑exon4 was reduced. In cell models carrying a specially engineered version of the amelogenin gene that produces less miR‑exon4, exon 4 was likewise skipped more frequently. Crucially, the microRNA itself was found inside the cell nucleus, where splicing occurs, and biochemical tests showed that it associates with the amelogenin precursor RNA at a specific control point in the neighboring intron. These findings support a dual role for miR‑exon4: indirectly shaping exon choice by adjusting splicing factors, and directly binding near exon 4 to influence whether it is kept or removed.
What this means for enamel health
Taken together, the study paints miR‑exon4 as a small but central coordinator of enamel formation. When present at the right level, it supports proper RUNX2 activity, keeps amelogenin production in balance, and helps ensure that exon 4 is included or excluded at the right stages. When miR‑exon4 is missing or reduced, this balance shifts: signaling pathways are disturbed, exon 4 is handled incorrectly, amelogenin isoforms become skewed, and early enamel mineralization is weakened. These insights help explain how certain mutations in the amelogenin gene can cause inherited enamel disorders, and they highlight nuclear microRNAs as important players in shaping the hardest tissue in the body.
Citation: Shemirani, R., Duong, T., Kim, R. et al. A splicing-derived microRNA from amelogenin exon4 regulates enamel formation via control of exon4 splicing and amelogenin expression. Sci Rep 16, 11044 (2026). https://doi.org/10.1038/s41598-026-40706-0
Keywords: dental enamel, amelogenin, microRNA, RNA splicing, amelogenesis imperfecta