Clear Sky Science · en
ABE9 fused to SpRY Cas9 nickase enables precise generation of bystander free mouse models
Sharper Tools for Building Better Disease Models
Many serious diseases are caused by single-letter typos in our DNA. To understand and eventually treat these conditions, scientists often create mice that carry the same mutations as human patients. But rewriting DNA in such a precise way is surprisingly difficult: older methods can scar the genome, introduce extra mutations, and muddle experimental results. This study introduces a refined gene-editing tool, called ABE9-SpRY, designed to change one specific DNA letter while leaving neighboring letters—and the rest of the genome—largely untouched.
From Heavy-Handed Cuts to Gentle DNA Tweaks
Traditional CRISPR-Cas9 editing works like a molecular pair of scissors, cutting both strands of DNA. Cells repair these breaks imperfectly, often leaving small insertions or deletions that disrupt genes in unpredictable ways. That’s useful for shutting genes off, but not for mimicking a precise disease-causing mutation. Base editors, by contrast, do not cut the DNA. Instead, they chemically nudge one DNA letter into another—here, changing A to G—while keeping the double helix intact. This approach dramatically reduces large deletions and rearrangements, but earlier adenine base editors still suffered from three key problems: they sometimes edited nearby “bystander” letters, they could only reach DNA sites next to specific short sequence tags, and they occasionally caused edits at unintended locations in the genome.
Designing a More Selective Molecular Pencil
The researchers set out to tackle all three limitations at once. They started with ABE9, a newer base editor variant that focuses its activity on a very narrow stretch of DNA, shrinking the “editing window” so far fewer neighboring letters are accidentally changed. They then fused ABE9 to an engineered Cas9 nickase called SpRY. Unlike standard Cas9, which requires a strict “NGG” tag next to the target site, SpRY can recognize a far broader set of DNA sequences. This makes it possible to reach disease-related positions in the genome that were previously off-limits. The resulting fusion tool, ABE9-SpRY, is intended to be both highly precise and far more flexible in where it can work.
Putting the New Editor to the Test in Cells and Embryos
To see whether ABE9-SpRY lived up to its design, the team compared it to a widely used, more aggressive editor called ABE8e-SpRY. They targeted four disease-relevant positions in genes encoding ion channels (TPC1, TPC2, and TRPM4), which play important roles in heart and liver function. In mouse nerve cells grown in the lab, ABE8e-SpRY edited the target letter more efficiently but also altered many nearby bases. ABE9-SpRY, in contrast, produced fewer overall edits but a much higher fraction of “clean” outcomes—reads in which only the intended letter was changed, with no extra mutations in the local region. The same pattern held in mouse embryos. When the researchers injected editing components into fertilized eggs, ABE8e-SpRY often converted nearly all copies of the target gene but with many bystander changes. ABE9-SpRY edited fewer copies overall, yet where it did act, the DNA sequence was typically corrected exactly as planned.
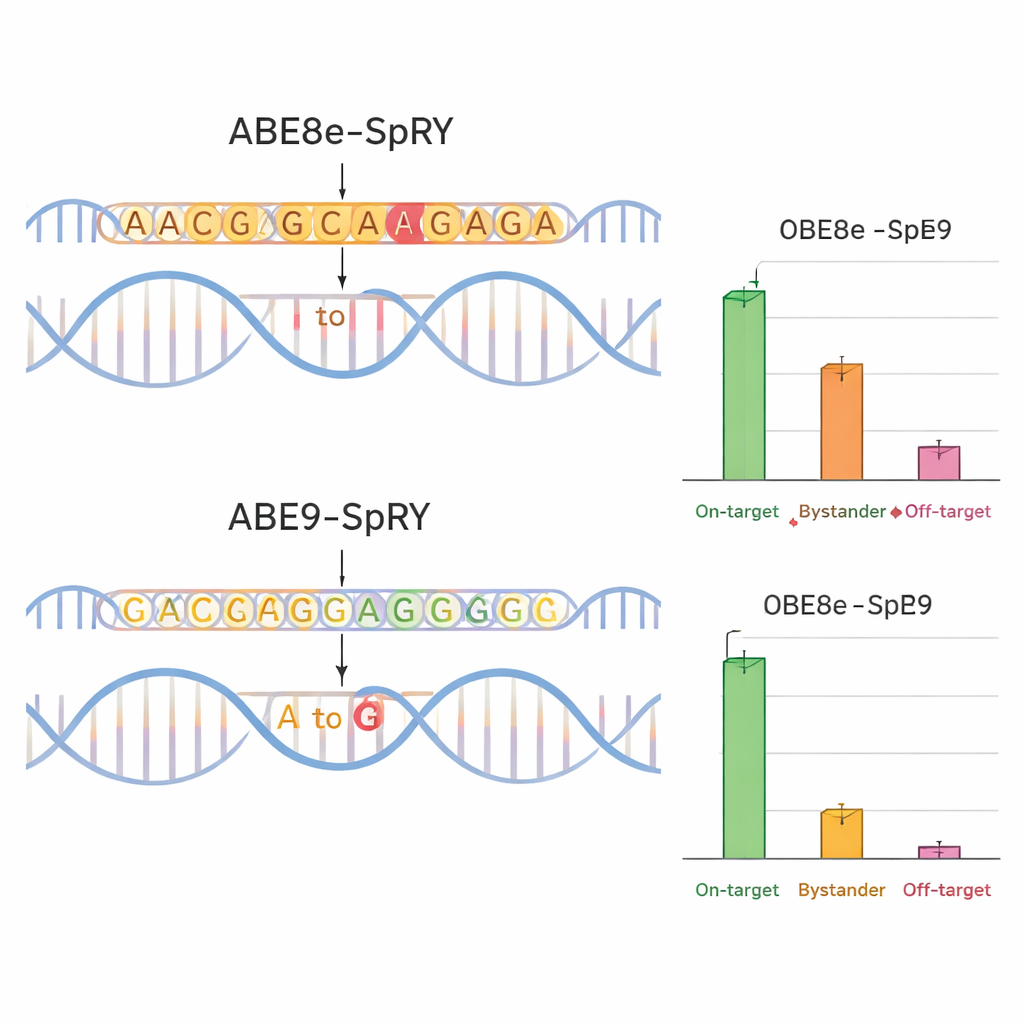
Cleaner Edits, Fewer Side Effects
Precision is not just about the local DNA neighborhood. The team also examined whether their editor altered distant regions of the genome. Using computational predictions and targeted sequencing, they measured off-target changes at likely look‑alike sites in mouse embryos. ABE8e-SpRY frequently introduced unintended A-to-G changes at these locations, sometimes at surprisingly high levels. ABE9-SpRY showed far fewer such events, with only rare off-target edits and no detectable activity in a specialized test for Cas9-independent DNA damage. Importantly, when ABE9-SpRY was used alone—not in pooled tests—to introduce two specific mutations in mice, it generated many founder animals in which the desired edit was present at high levels and passed reliably to offspring, again with minimal bystander changes or small insertions and deletions.
Extending Precision Editing to Human Stem Cells
Because patient-specific stem cells are increasingly used to study disease and test drugs, the researchers also tried ABE9-SpRY in human induced pluripotent stem cells. They targeted the human version of the TPC1 gene at a position equivalent to one of the mouse sites. Using a fluorescent reporter to enrich for edited cells, they found that ABE8e-SpRY produced higher raw editing rates but again modified multiple nearby bases. ABE9-SpRY edited fewer alleles overall, but the proportion of perfectly edited stretches—one intended change and nothing else—was markedly higher, with fewer disruptive insertions or deletions. In stem-cell work, where individual clones are expanded for detailed study, this kind of “clean” edit is usually more valuable than brute-force efficiency.
Why This Matters for Future Disease Research
For scientists trying to understand how a single DNA letter change leads to disease, extra unintended mutations can be deeply misleading. This study shows that ABE9-SpRY, while somewhat less powerful in terms of raw editing speed, can write single-letter changes into the genomes of mice and human stem cells with striking accuracy and broad targeting flexibility. That makes it a promising workhorse for building faithful animal and cell models of human genetic disorders, especially in situations where even one stray mutation could confuse the biology or derail a potential therapy.
Citation: Ong, J.K., Bhunia, S., Hilbert, B. et al. ABE9 fused to SpRY Cas9 nickase enables precise generation of bystander free mouse models. Sci Rep 16, 7463 (2026). https://doi.org/10.1038/s41598-026-40642-z
Keywords: adenine base editing, CRISPR disease models, mouse genetics, off-target effects, hiPSC gene editing