Clear Sky Science · en
The Ies6 subunit is essential for INO80-mediated nucleosome organization
How cells keep their DNA neatly packed
Every cell in your body must cram long strands of DNA into a tiny nucleus without creating hopeless tangles. It does this by winding DNA around protein “spools” called nucleosomes, forming an organized landscape that helps genes turn on and off at the right time. This study explores how a specific protein piece, called Ies6, helps a larger machine arrange these DNA spools in an orderly fashion—work that is fundamental to genome stability and healthy cell growth.
The beads-on-a-string problem
DNA inside the nucleus looks a bit like a string of beads. Each bead is a nucleosome, and the distance from one bead to the next—the nucleosome repeat length—tends to be very regular along genes. This spacing is not accidental. It is actively set by molecular machines known as chromatin remodelers, which use energy from ATP to slide and reposition nucleosomes. One such remodeler, called INO80, is known to help place the first nucleosome after a gene’s start site and to build evenly spaced arrays further downstream. But INO80 itself is made of many parts, and until now it was unclear how crucial a single small subunit like Ies6 is for keeping nucleosomes properly arranged.
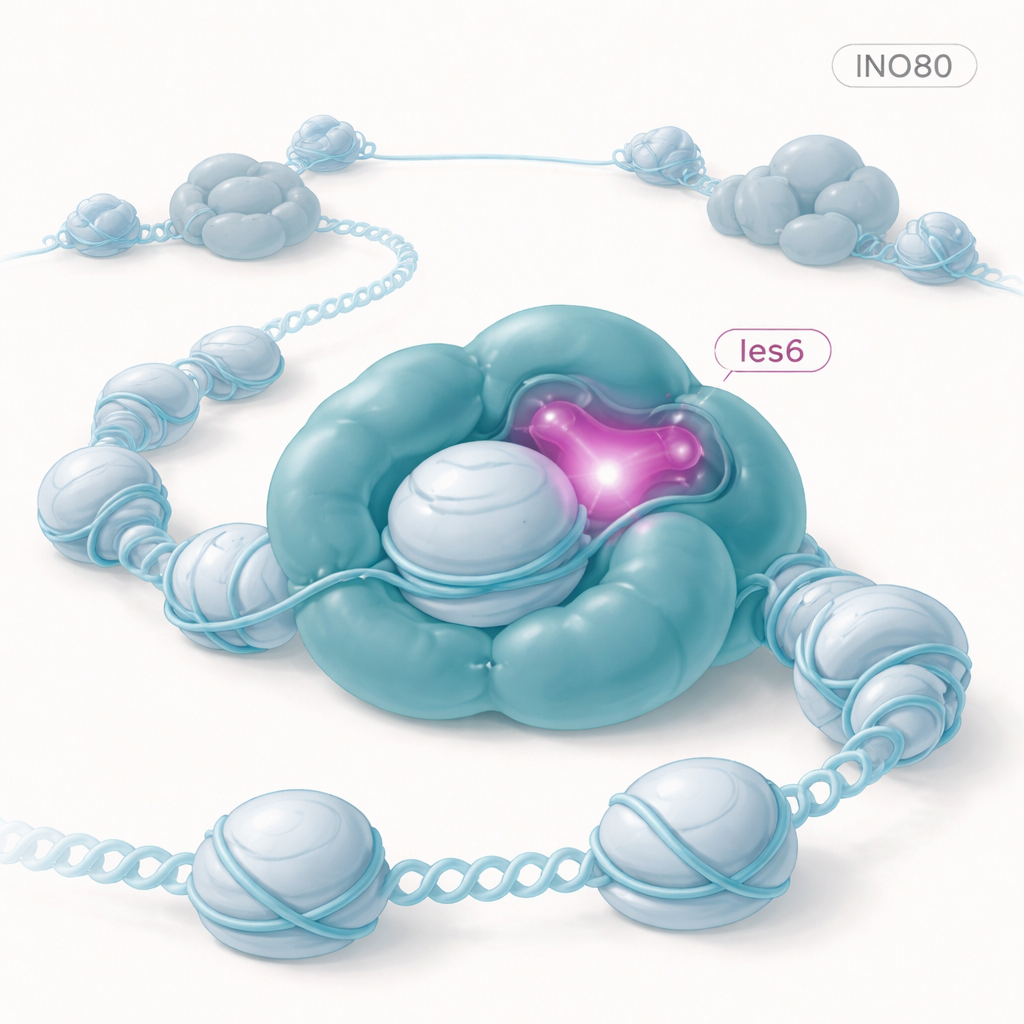
A tiny subunit with a big impact
The researchers worked in baker’s yeast, a favorite model for studying chromosome biology because many of its control systems resemble those in human cells. They deleted the gene that makes Ies6 and examined how this affected nucleosome organization across the yeast genome. Using a technique called MNase-Seq, which maps where nucleosomes sit on DNA, they found that without Ies6, nucleosomes shifted their positions and packed slightly closer together. On average, the spacing between nucleosomes shrank by about three DNA base pairs, and the normally crisp, evenly spaced arrays along genes became blurrier and less regular. These changes closely mirrored what happens when the core INO80 engine itself is removed, suggesting that Ies6 is not a minor accessory but central to INO80’s organizing power.
Backup systems and a lethal partnership
Cells rarely rely on a single tool for an important job, and nucleosome spacing is no exception. Yeast carries several remodelers—such as Isw1, Isw2, and Chd1—that can also slide nucleosomes. The team tested how loss of Ies6 interacted with these other machines by combining mutations. Strikingly, yeast cells missing both Ies6 and the remodeler Isw2 could not survive, a phenomenon known as synthetic lethality. This implies that INO80 (through Ies6) and Isw2 carry out overlapping, essential tasks in shaping chromatin at specific regions, such as gene beginnings, replication origins, or ribosomal DNA. When both systems fail, the cell can no longer maintain a functional chromatin landscape.
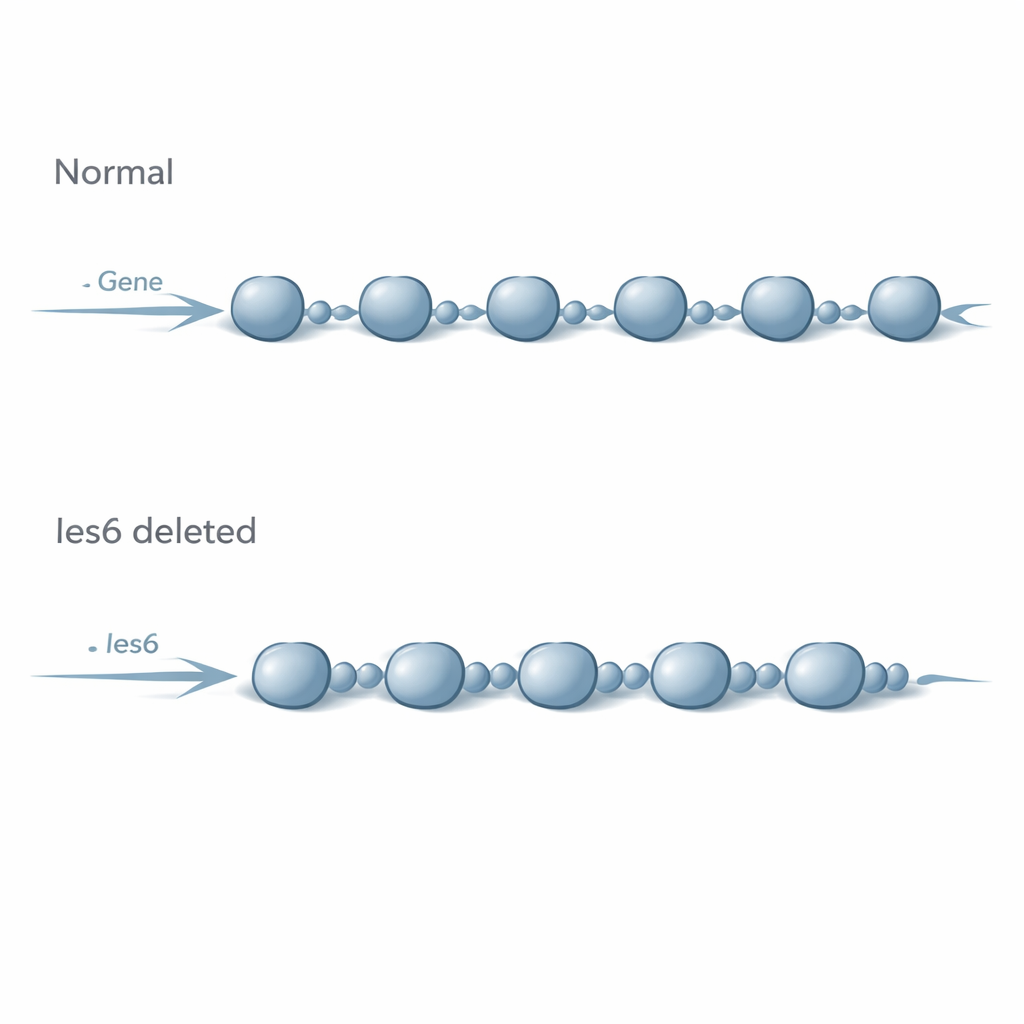
Chromatin changes without obvious gene switches
One might expect that disturbing nucleosome spacing would strongly alter which genes are turned on or off. To test this, the authors compared their chromatin data with existing RNA sequencing measurements from cells lacking Ies6 or the main INO80 subunit. Surprisingly, genes that changed their expression after these deletions did not strongly overlap with genes that showed the biggest shifts in nucleosome spacing. In other words, large-scale rearrangements of nucleosome arrays within gene bodies did not neatly predict changes in steady-state RNA levels. This suggests that INO80 and Ies6 may be more important for fine-tuning genome stability—such as preventing spurious, “cryptic” transcription or helping DNA replication proceed smoothly—than for acting as simple on/off switches for most genes.
Why this hidden organizer matters
By zeroing in on a single subunit, Ies6, this study reveals how a small structural element can be essential for the global architecture of chromatin. Ies6 enables the INO80 complex to set regular nucleosome spacing across thousands of genes, and its loss disrupts this order in a way that rivals removing the entire INO80 engine. At the same time, cells depend on partially overlapping remodelers like Isw2 to safeguard critical chromosomal regions, explaining why losing both becomes lethal. For a lay reader, the key message is that the health of our genomes depends not only on the DNA code itself, but also on the precise arrangement of DNA packaging units—and that even tiny components of the molecular packing machinery can have outsized effects on how the genome works and survives.
Citation: Singh, A.K., Mueller-Planitz, F. The Ies6 subunit is essential for INO80-mediated nucleosome organization. Sci Rep 16, 7466 (2026). https://doi.org/10.1038/s41598-026-40504-8
Keywords: chromatin remodeling, nucleosome spacing, INO80 complex, yeast genetics, genome organization