Clear Sky Science · en
MyoFuse is a fully AI-based workflow for automated quantification of skeletal muscle cell fusion in vitro
Why counting muscle cells matters
When we exercise, recover from injury, or develop diseases like diabetes, our skeletal muscles constantly remodel themselves. In the lab, scientists mimic this process by growing muscle cells in a dish and watching single cells fuse into long, multi‑nucleated fibers. A simple number called the fusion index tells them how well this fusion is happening. But today that number is usually obtained by people manually counting thousands of tiny cell nuclei on a screen—slow work that can be inconsistent and prone to error. This study introduces MyoFuse, a fully AI‑based workflow that automates this counting task, aiming to make muscle research faster, more reliable, and less biased.
The challenge of seeing what is really there
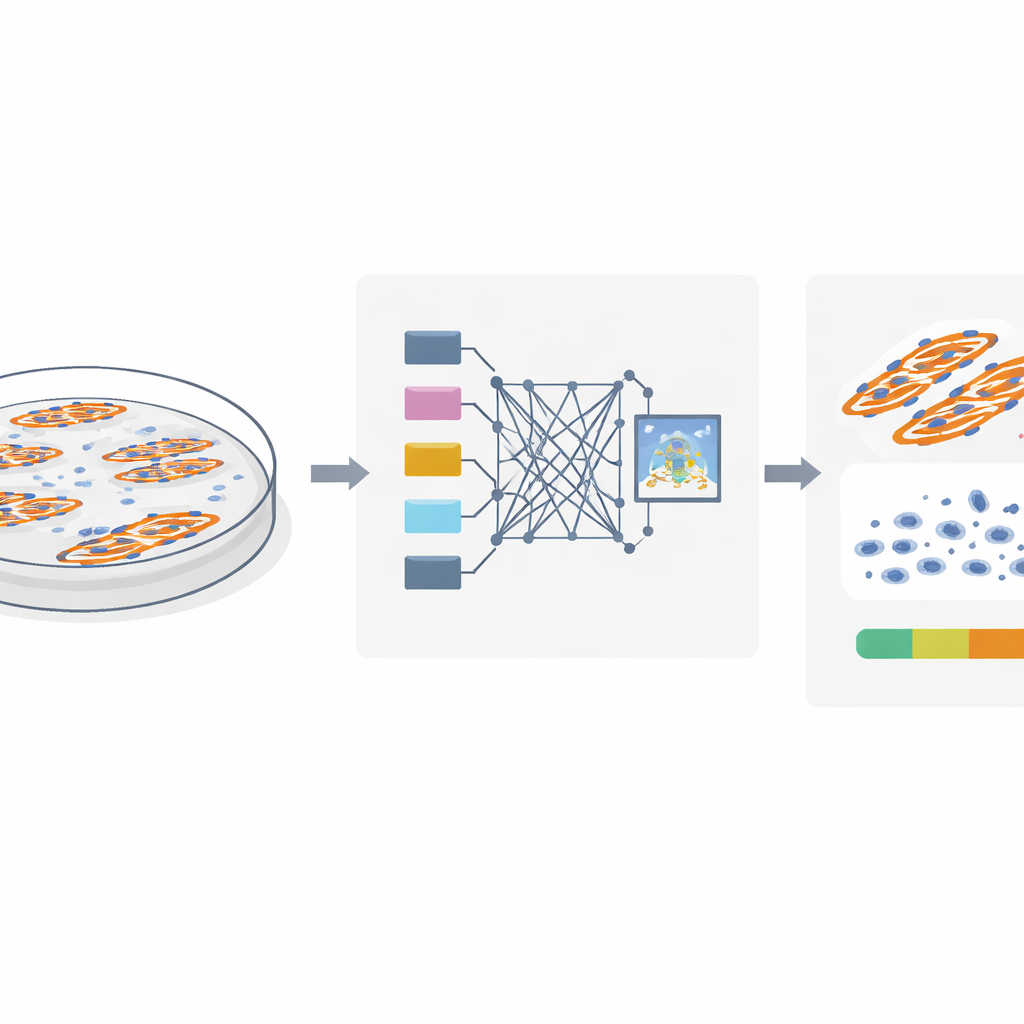
To study muscle cell fusion, researchers stain cell nuclei and muscle fibers with fluorescent dyes and take microscope images. The key question is which nuclei are truly inside fused muscle fibers and which belong to unfused cells sitting nearby. Traditional methods assume that if a nucleus overlaps a muscle fiber in a flat, two‑dimensional image, it must be inside that fiber. However, cells grow in three dimensions: a nucleus can easily lie above or below a fiber and still look like it overlaps in the image. The authors use detailed confocal imaging in both mouse and human muscle cells to show that this overlap trick can be misleading, causing many nearby nuclei to be wrongly counted as part of fused fibers and inflating the fusion index.
A smarter way to read fluorescent images
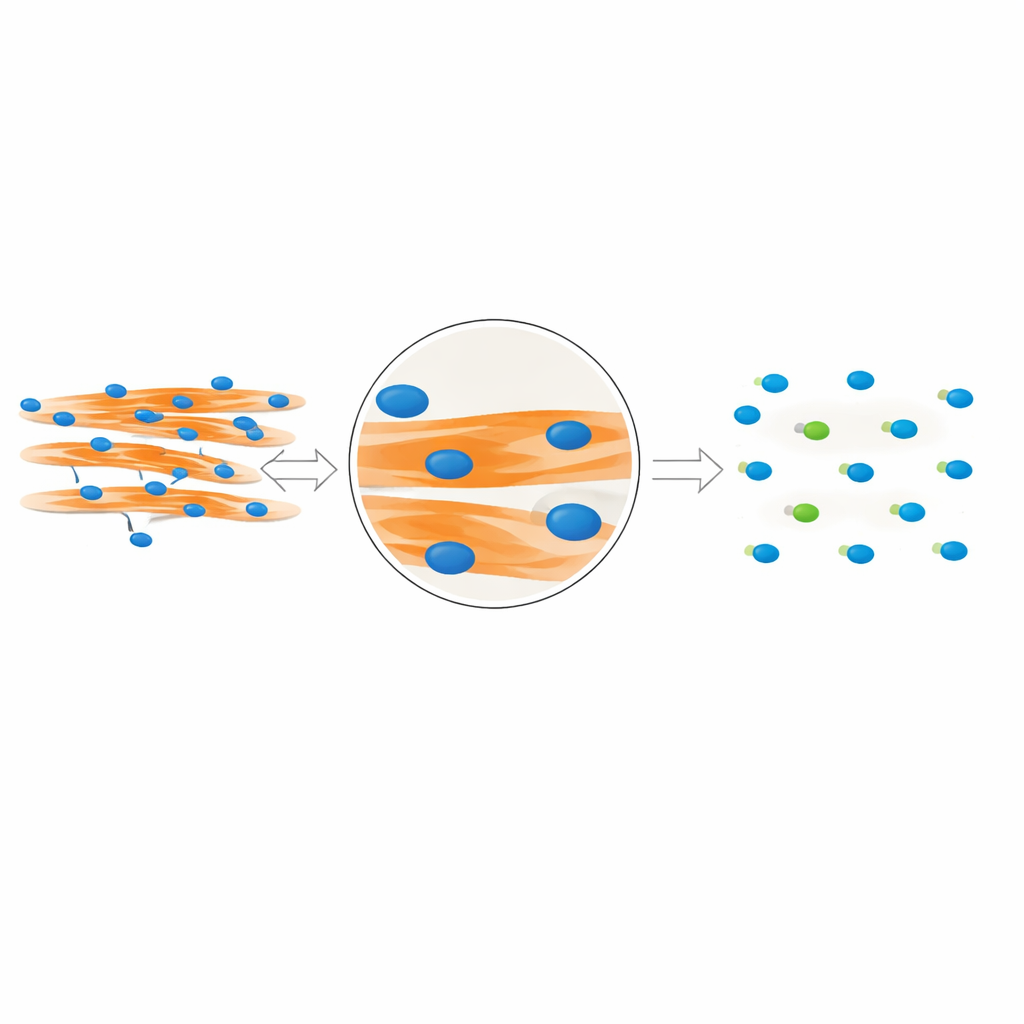
The team realized that true “in‑fiber” nuclei leave a distinct visual fingerprint. Because these nuclei physically occupy space inside the fiber, they create small dark gaps in the fluorescent signal that marks the muscle protein MyHC. In contrast, nuclei that sit above or below a fiber do not disrupt this signal. Building on this insight, the authors designed MyoFuse, a two‑step AI workflow. First, a specialized segmentation model (adapted from the open‑source tool Cellpose) precisely outlines individual nuclei, even when they are tightly packed into clusters. Second, a lightweight neural network classifier examines the surrounding MyHC signal for each nucleus and decides whether it lies inside a muscle fiber or outside, based solely on this local pattern rather than on simple overlap.
How well the AI matches human experts
The researchers rigorously tested MyoFuse on images of mouse C2C12 muscle cells and primary human muscle cells from different muscles. For both species, the AI’s count of total nuclei and its calculated fusion index agreed extremely well with careful manual annotations by experts, with near‑perfect correlations. At the level of individual nuclei, the classifier correctly distinguished nuclei inside versus outside fibers in more than 90% of cases across datasets, with performance metrics rivaling human‑level discrimination. Importantly, MyoFuse also worked well on a separate set of human cells that had never been used for training, suggesting that the approach can generalize to new samples rather than just memorizing the training images.
Revealing hidden bias in common methods
Beyond its accuracy, MyoFuse exposed systematic problems in widely used mask‑based methods that rely on simple overlap between nuclear and fiber signals. When the authors compared fusion indices from MyoFuse with those from a refined mask approach applied to the same images, the mask method consistently overestimated fusion, especially in areas where muscle fibers covered a larger fraction of the dish. Adjusting detection thresholds changed the numbers but did not remove this underlying bias; apparent improvements often came from errors canceling each other rather than better biology. The team also showed that fusion estimates can swing widely across different regions of a single well, underscoring that analyzing only a few hand‑picked fields can give a distorted view of how well cells are fusing.
What this means for future muscle research
MyoFuse offers muscle biologists a way to measure cell fusion that is both faster and more faithful to what is actually happening in the dish. By combining automated microscopy with an AI that can segment and classify hundreds of thousands of nuclei in minutes, the workflow reduces human workload, minimizes subjective choices about where to look and how to threshold images, and avoids counting nearby cells as fused muscle. The authors acknowledge that extreme imaging conditions or very different staining protocols may require retraining, but the method is openly available and designed to be adaptable. For labs studying muscle development, aging, regeneration, or metabolic disease, MyoFuse promises more robust fusion measurements—and with them, more trustworthy conclusions about how muscles grow and change.
Citation: Lair, B., Cazorla, C., Lobeto, A. et al. MyoFuse is a fully AI-based workflow for automated quantification of skeletal muscle cell fusion in vitro. Sci Rep 16, 9387 (2026). https://doi.org/10.1038/s41598-026-40047-y
Keywords: skeletal muscle, cell fusion, artificial intelligence, image analysis, myogenesis