Clear Sky Science · en
Implementation of SARS-CoV-2 genomic surveillance during the COVID-19 pandemic through an academic–public health collaboration in southeast Michigan
Why tracking variants close to home matters
During the COVID-19 pandemic, most of us heard about new variants from national headlines. But the virus did not spread in the same way everywhere. This paper describes how scientists, hospitals, and public health officials in southeast Michigan built a local system to watch the coronavirus evolve in real time. By reading the virus’s genetic code from thousands of patient samples, they could see which variants were spreading, where they were taking hold, and which communities were being hit hardest—information that can guide faster and more targeted responses in future outbreaks.
Building a neighborhood early‑warning system
The team brought together Wayne State University, the Detroit Health Department, Henry Ford Health, a fleet of mobile health units, and the state health department. Their shared goal was to create a regional “early‑warning system” for COVID-19 based on the virus’s genetic fingerprints. Hospitals, public clinics, and mobile vans collected nose swabs from people who tested positive. These samples were barcoded, safely stored in a central biobank, and then routed to a university laboratory equipped to handle large numbers of tests. Careful data agreements and privacy protections made it possible to pair each viral sequence with basic information about the patient and their neighborhood without revealing identities.
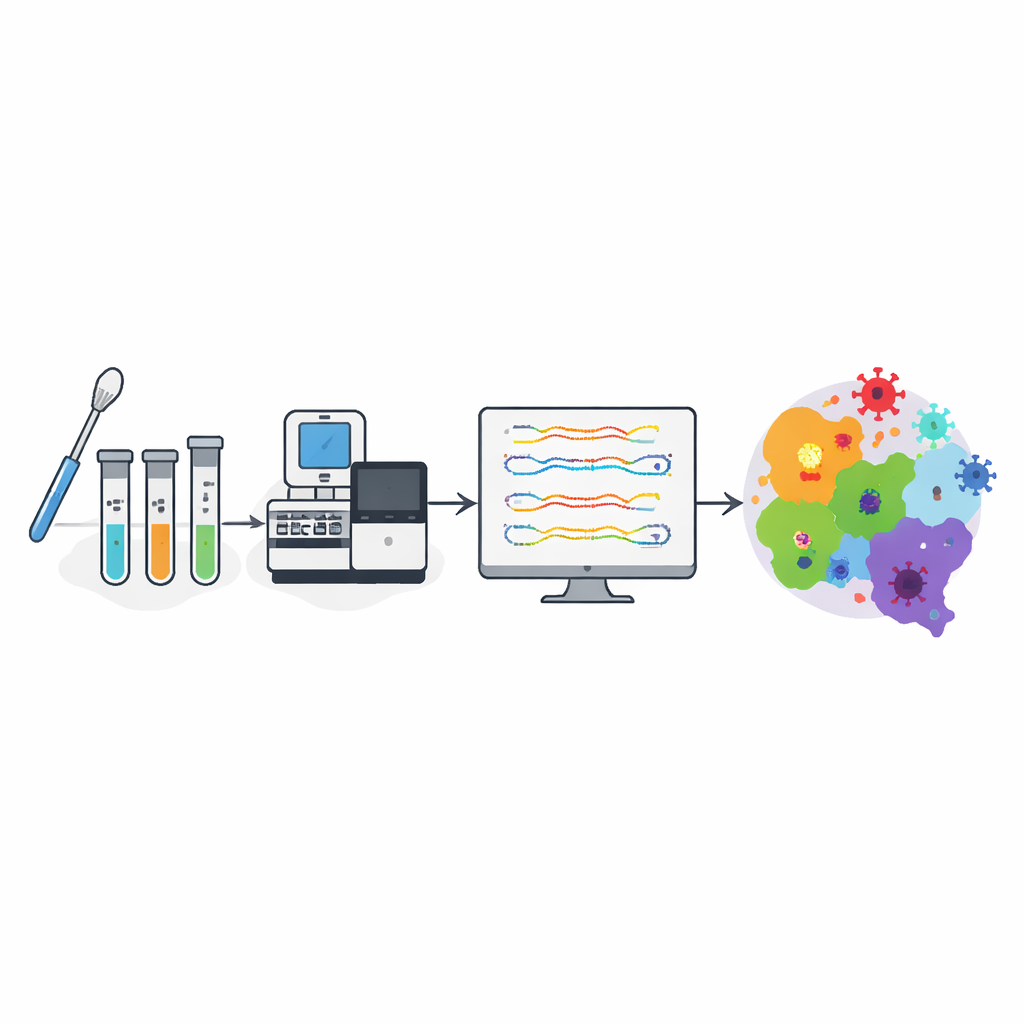
From swab to genetic map
Inside the lab, technicians in biosafety hoods heat‑treated the samples, extracted viral genetic material, and confirmed infection using a standard PCR test. The same material then went through high‑throughput machines that read out the entire genetic code of SARS-CoV-2 from each patient. Powerful computers compared these sequences to reference genomes and used specialized software to assign each virus to a known variant group. This pipeline turned raw swabs into organized genetic data—showing which versions of the virus were circulating, when they appeared, and how they changed over time.
Where and in whom the virus hit hardest
Between early 2022 and mid‑2024, the program collected over 7,500 samples and successfully sequenced more than 6,200 of them, most from patients in the Henry Ford Health system. These cases came from nearly 300 ZIP codes across southeast Michigan. Omicron was by far the dominant variant, accounting for about two‑thirds of sequenced infections, and the pattern of variants closely matched what was seen in statewide data. Older adults were more likely to appear in the dataset and more likely to die, reflecting their higher risk of severe disease. Infections were somewhat more common in women, but deaths were slightly more common in men. When researchers compared racial and neighborhood patterns, they found that Black residents had higher infection and death rates than White residents—yet once infection, age, and variant were taken into account, it was neighborhood disadvantage, not race alone, that best explained higher death risks.

Watching the virus change over time and space
Because each virus carried a time stamp and a location, the team could draw maps and timelines of the pandemic in their region. They saw early waves dominated by older lineages, followed by a brief surge of the Delta variant, and then a long period where Omicron and its offshoots took over. Omicron showed the widest geographic reach, including into northern parts of the state, but the highest infection rates clustered in and around the Detroit metro area. When the researchers compared their data to national and global databases, they found strong agreement in overall variant trends but also clear signs of local quirks and sampling gaps, underscoring why regional surveillance adds value beyond national totals.
What this model means for the future
In plain terms, this paper shows that a city and its surrounding communities can build their own “genetic radar” for tracking a virus, even during a fast‑moving crisis. The southeast Michigan program connected hospitals, mobile clinics, and public health agencies into a working system that could see dangerous variants arriving, follow them as they spread, and link them to real‑world outcomes like hospitalization and death. Although the authors acknowledge limits—such as uneven sampling and the focus on one health system—they argue that the basic framework is sustainable and adaptable. With adequate support, similar partnerships could be used to monitor influenza, RSV, Mpox, or the next unknown pathogen, giving local leaders an earlier and clearer view of trouble before it becomes a full‑blown emergency.
Citation: Raychouni, R., Zhang, X., Bauer, S.J. et al. Implementation of SARS-CoV-2 genomic surveillance during the COVID-19 pandemic through an academic–public health collaboration in southeast Michigan. Sci Rep 16, 8680 (2026). https://doi.org/10.1038/s41598-026-39974-7
Keywords: SARS-CoV-2 variants, genomic surveillance, COVID-19 epidemiology, public health data, Detroit Michigan