Clear Sky Science · en
R-GAT: cancer document classification leveraging graph-based residual network for scenarios with limited data
Why sorting cancer papers matters
Every day, scientists publish hundreds of new studies on cancer, from early detection to promising drugs. Most of this work first appears as short summaries called abstracts. Doctors, researchers, and policy makers can’t possibly read them all, yet missing an important paper could slow progress. This study tackles a simple but powerful question: can we build a fast, lightweight computer system that automatically sorts cancer-related abstracts by type of cancer, even when only a modest amount of labeled data and computing power are available?
A smarter way to read cancer research
The authors focus on four kinds of abstracts found in the PubMed database: those about thyroid cancer, colon cancer, lung cancer, and more general biomedical topics. They created a carefully checked collection of 1,875 recent abstracts, roughly equal in size across these four groups. This balance helps avoid bias toward any single cancer type. Before any modeling, the texts were cleaned: words were split into tokens, spelling was checked, related word forms were merged, and uninformative terms were removed. The cleaned abstracts were then converted into numerical form using several standard methods so that different kinds of models could be fairly compared.
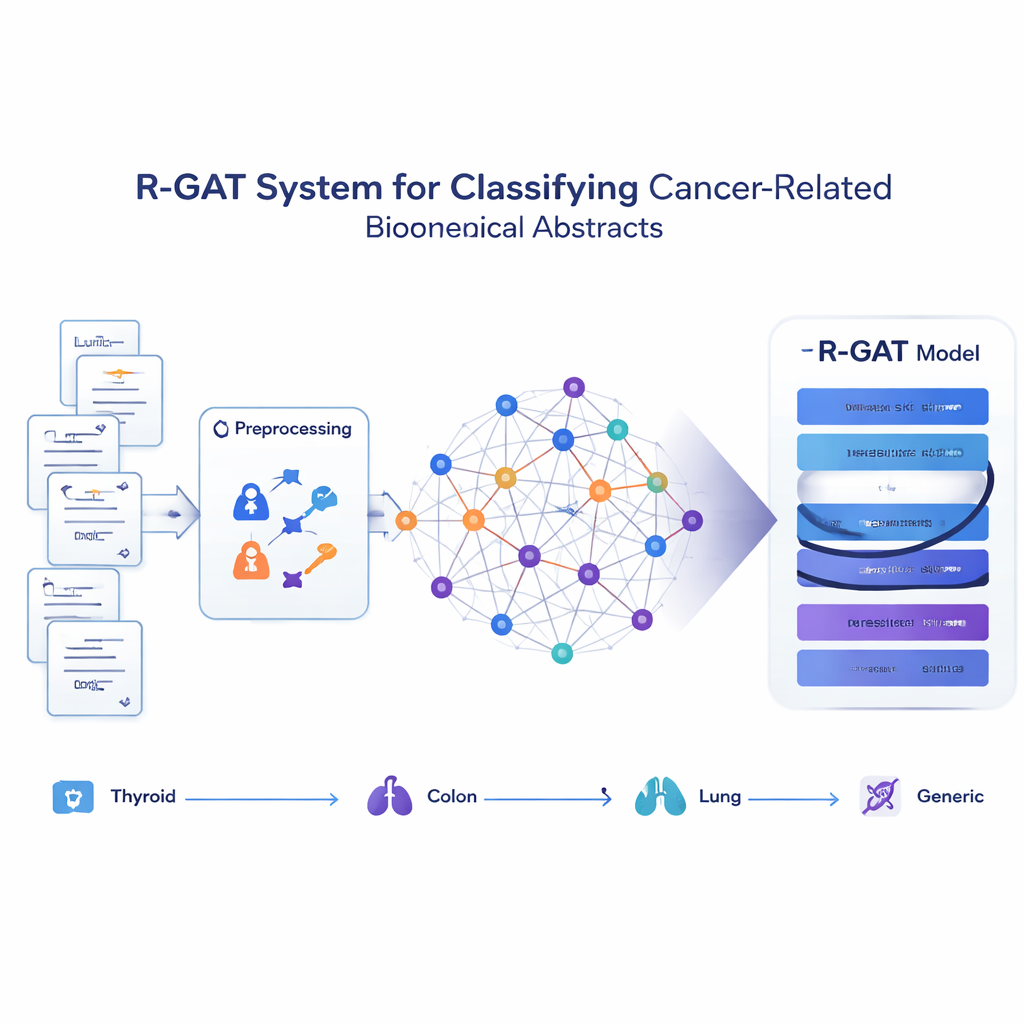
Turning papers into a network of ideas
Instead of treating each abstract as an isolated string of words, the proposed method, called R-GAT (Residual Graph Attention Network), views the whole collection as a network. In this network, each abstract is a node, and connections represent how similar two abstracts are in content. If two papers discuss closely related topics, the link between them is strong; if not, it is weak or absent. This lets the model look at an abstract in the context of its neighbors, mimicking how a human reader might understand one study better by knowing what related work says.
How the new model learns from neighbors
R-GAT builds on two key ideas from modern artificial intelligence: attention and residual connections. Attention allows the model to focus more on the most relevant neighboring abstracts in the network, rather than treating all neighbors equally. Multiple attention “heads” look for different types of patterns at the same time. Residual connections act like shortcuts that pass information around the deeper layers of the network, helping the model avoid losing important signals as it learns. After processing the graph through several attention layers and these shortcut paths, the system condenses information from the whole network into a compact summary that is fed into a final classifier predicting which of the four categories each abstract belongs to.
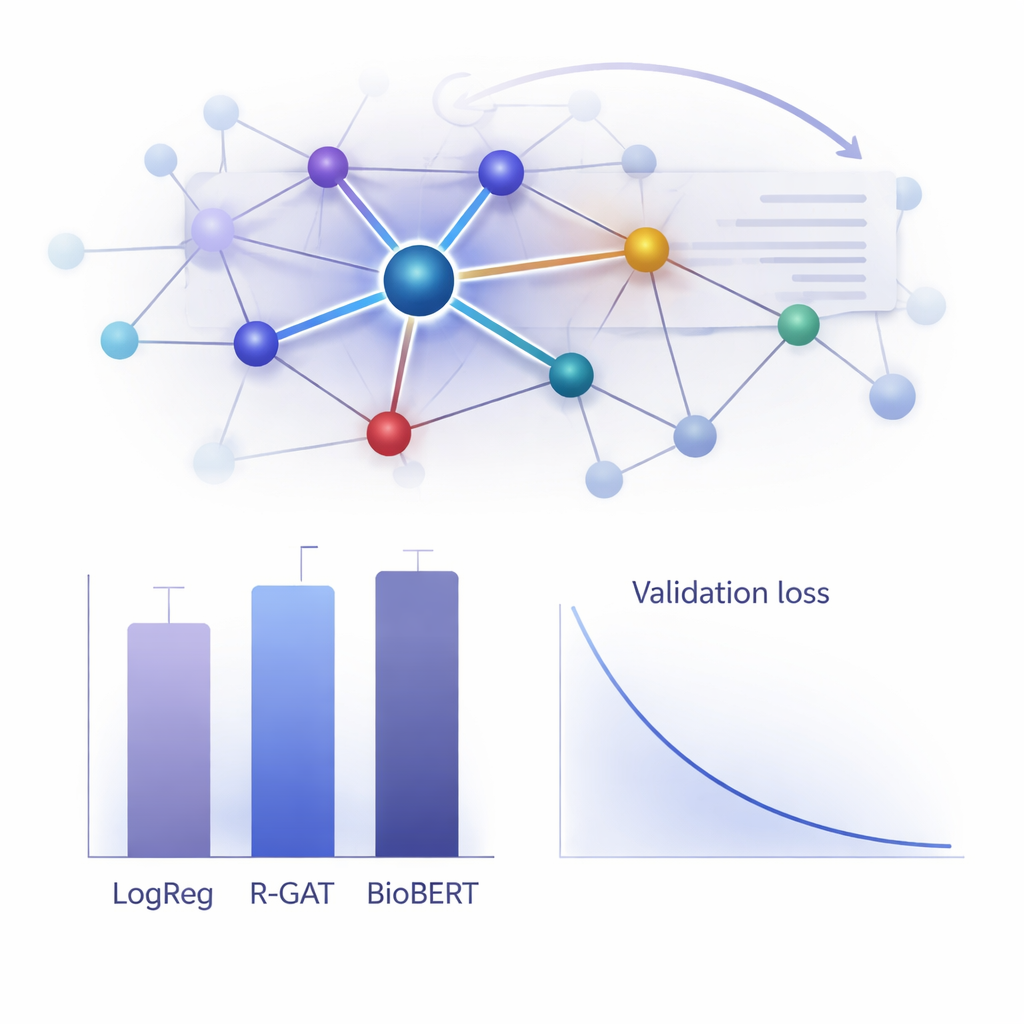
How well does it work in practice?
To judge the value of R-GAT, the authors compared it to a wide range of alternatives, from classic linear models to state-of-the-art transformer systems like BioBERT, which are popular but computationally heavy. Surprisingly, a simple logistic regression model using word-count features achieved the highest raw score on this particular dataset, and BioBERT also performed extremely well—but both had drawbacks, including dependence on specific feature choices or the need for substantial computing resources. R-GAT reached a macro F1-score of about 0.96, close to the best models, while showing very stable results across different train–test splits. Careful tests where attention or residual connections were removed showed clear drops in performance, confirming that both ingredients are crucial to the model’s robustness when data are limited.
What this means for future cancer research
For a layperson, the takeaway is straightforward: R-GAT is a practical tool that helps sort cancer research papers by cancer type with high and consistent accuracy, without requiring giant datasets or expensive hardware. It does not replace the most powerful language models on the market, but it offers a reliable middle ground—especially useful for hospitals, research groups, or public health teams that need dependable, reproducible results under tight data and budget constraints. By releasing both their model and their curated dataset openly, the authors also provide a shared benchmark others can use to build and test improved systems. In the long run, such tools could make it much easier for experts to stay on top of the cancer literature and turn new findings into better care.
Citation: Hossain, E., Nuzhat, T., Masum, S. et al. R-GAT: cancer document classification leveraging graph-based residual network for scenarios with limited data. Sci Rep 16, 6582 (2026). https://doi.org/10.1038/s41598-026-39894-6
Keywords: cancer informatics, biomedical text mining, document classification, graph neural networks, limited data learning