Clear Sky Science · en
Attention-guided enhanced deconvolution enables reference-free cell type estimation in spatial transcriptomics
Seeing Cells in Place
Modern biology can read the activity of thousands of genes at once, not just in isolated cells but directly within thin slices of tissue. This "spatial transcriptomics" view reveals where different cells live and interact, but each measurement often blends signals from many neighboring cells. The study introduces a new computational method, called AGED, that can untangle these mixtures and estimate which cell types are present where—without needing a separate, carefully matched single-cell reference dataset. 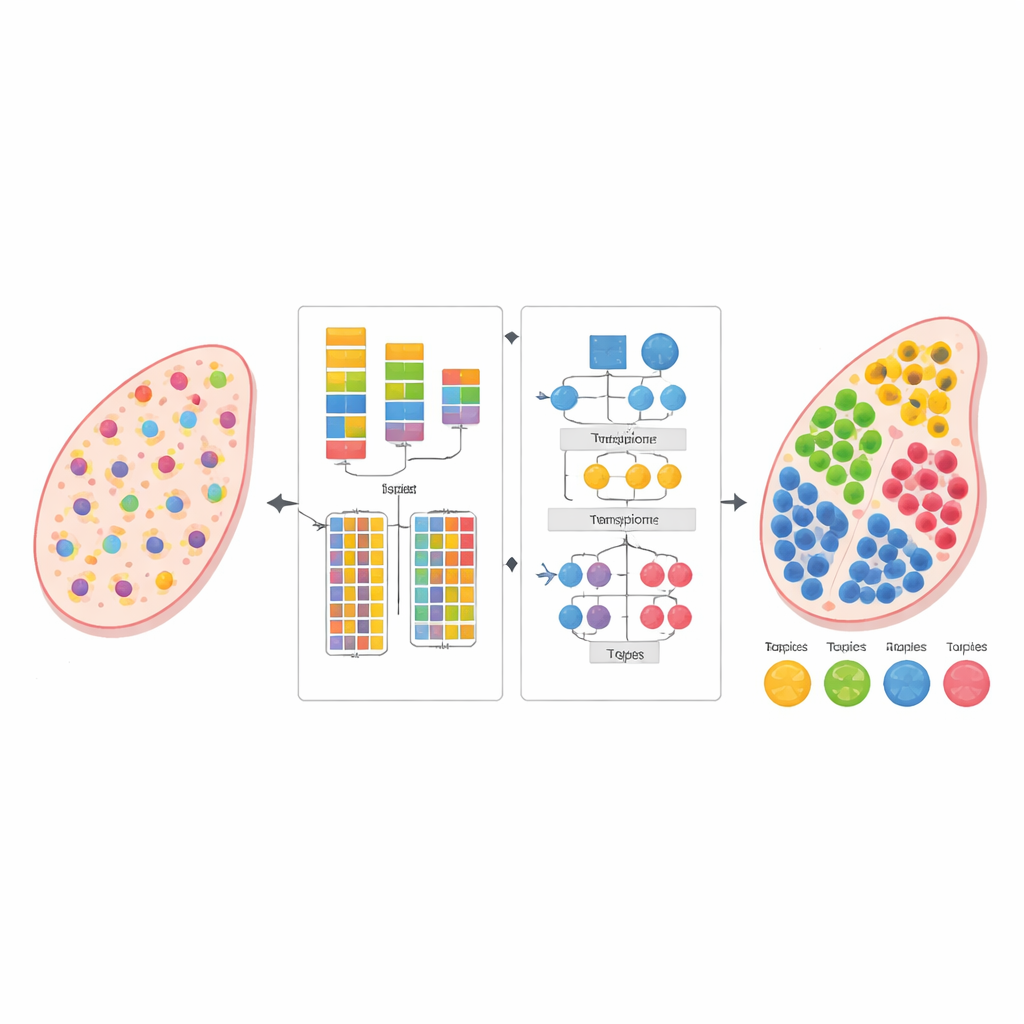
Why Mapping Cells in Tissues Is Hard
Spatial transcriptomics platforms measure gene activity across a grid of spots laid over a tissue slice. Because most spots capture several cells at once, researchers must mathematically decompose the mixed signals to recover the underlying cell types and their proportions. Existing tools often rely on external single-cell reference atlases of the same tissue. Those atlases can be missing for rare tissues, special disease states, or unusual experimental conditions, and even when available they may not match perfectly, introducing biases. Reference-free methods avoid this dependency, but current approaches struggle with complex spatial patterns, subtle gene relationships, and the challenge of deciding how many distinct cell types to look for in the first place.
A Two-Step Strategy for Untangling Mixtures
The authors designed AGED as a two-stage framework that combines ideas from statistics and modern deep learning. In the first stage, the method tests a range of possibilities for how many cell types might be present in the tissue. It uses a fast attention-based neural network, known as a Performer, to learn candidate decompositions and then scores them using several criteria at once: how well the model reconstructs the observed gene counts, how clearly the inferred cell groups separate from each other, and how diverse those groups are. A curve-fitting procedure finds an "elbow point" where adding more cell types brings little benefit, allowing the method to automatically select a suitable number instead of relying on a user’s guess.
Guided Attention to Capture Biology
Once the number of cell types is set, AGED’s second stage refines the solution with a richer attention-based architecture. It starts from a statistical topic model that treats each tissue spot as a mixture of hidden "themes"—here standing in for cell types—and each cell type as a characteristic gene pattern. These initial themes provide global structure. The model then layers several attention mechanisms on top: one connects the statistical themes to the neural network, another pools information from neighboring spots in physical space, and a third links themes to genes directly. A gating system lets the model decide, for each case, how much to trust the prior statistical patterns versus the local data. Additional constraints encourage sparse solutions, reflecting the biological reality that most tissue locations are dominated by only a few main cell types. 
Putting the Method to the Test
The researchers evaluated AGED on several types of data. In simulated mouse olfactory bulb tissue, the method recovered four known anatomical layers and matched the true cell compositions more closely than widely used reference-based and reference-free tools, achieving both high correlation with ground truth and low reconstruction error. In human pancreatic ductal adenocarcinoma, AGED automatically chose a twenty-cell-type solution that aligned with pathologist-annotated regions such as tumor, duct, and normal pancreas, outperforming other methods on a structural similarity measure that compares inferred maps to the visible tissue structure. In human thymus tissue, AGED accurately separated key cell populations and captured a biologically expected negative relationship between two specialized epithelial cell types—a pattern that competing approaches failed to reproduce. Additional analyses on other datasets and at single-cell-like resolution further supported the method’s robustness.
What This Means Going Forward
To a non-specialist, AGED can be viewed as a smart unmixing engine for complex tissues: it learns how many distinct cell communities are present, where they are located, and which genes define them, all from the spatial data itself. By weaving together interpretable statistical models with flexible attention-based neural networks, the framework offers both accuracy and insight, even when no suitable reference atlas exists. This makes it a practical tool for exploring tissue organization in health and disease, from brain layers to tumors and immune organs, and points toward a broader strategy for using prior knowledge to guide powerful but opaque machine-learning models in biology.
Citation: Yang, X., Wang, Y. & Chen, X. Attention-guided enhanced deconvolution enables reference-free cell type estimation in spatial transcriptomics. Sci Rep 16, 8097 (2026). https://doi.org/10.1038/s41598-026-39703-0
Keywords: spatial transcriptomics, cell type deconvolution, deep learning, tissue architecture, reference-free analysis