Clear Sky Science · en
Gene driven analytical learning model for accurate breast cancer diagnosis
Why this research matters to patients and families
Breast cancer is now the most commonly diagnosed cancer in women worldwide, and patients with what looks like the same disease on paper can have very different outcomes. This study shows how patterns in thousands of genes, combined with a carefully designed artificial intelligence system, can help doctors tell more reliably who has cancer and how severe it may be—using only real patient data and a compact set of key genes.
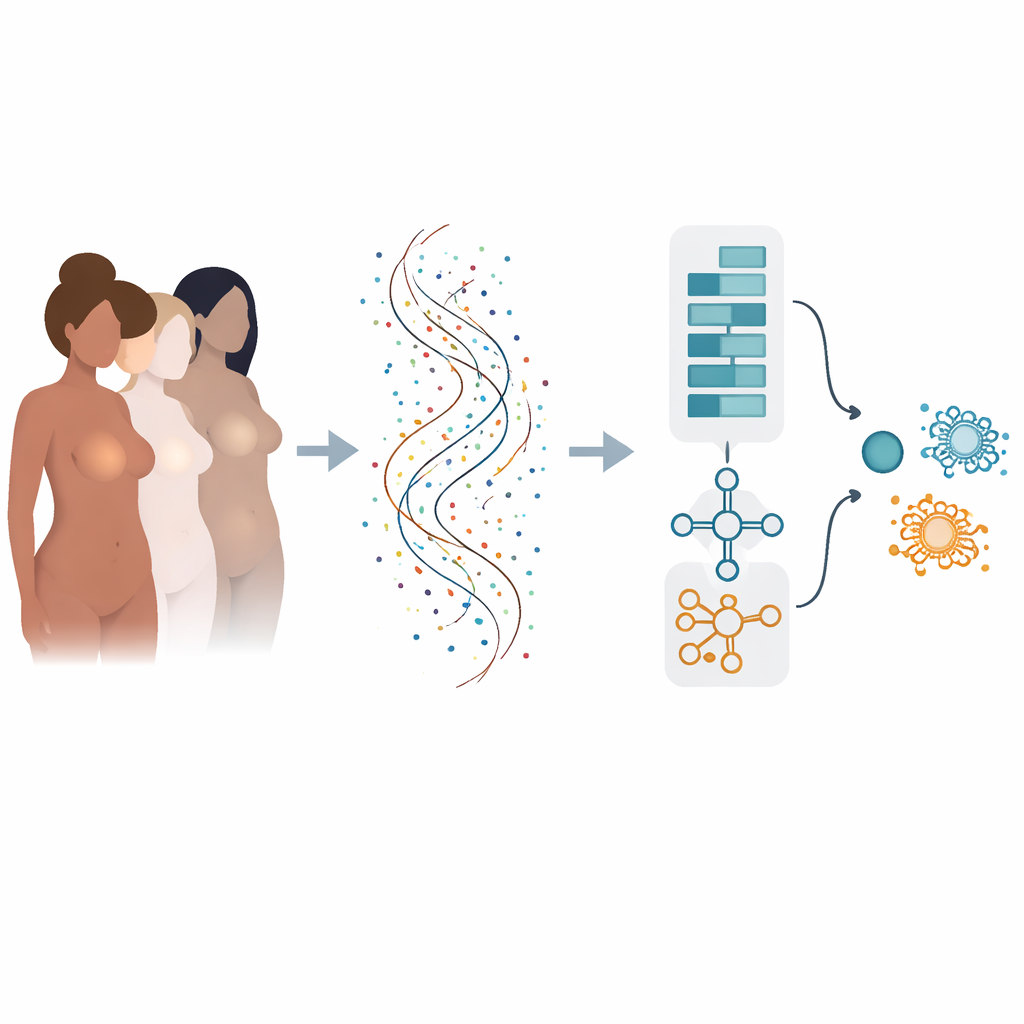
From many risk factors to the language of genes
Breast cancer risk is shaped by many influences: inherited gene changes, hormones, body weight, lifestyle, and more. Once cancer appears, its behavior is driven by which genes are switched on or off inside each tumor. Modern sequencing can measure activity in tens of thousands of genes at once, but turning this ocean of numbers into clear yes-or-no answers for diagnosis and prognosis is difficult. Traditional computer methods often look at genes one by one and can miss the way groups of genes act together, or they may perform well only on one dataset and fail when tested elsewhere.
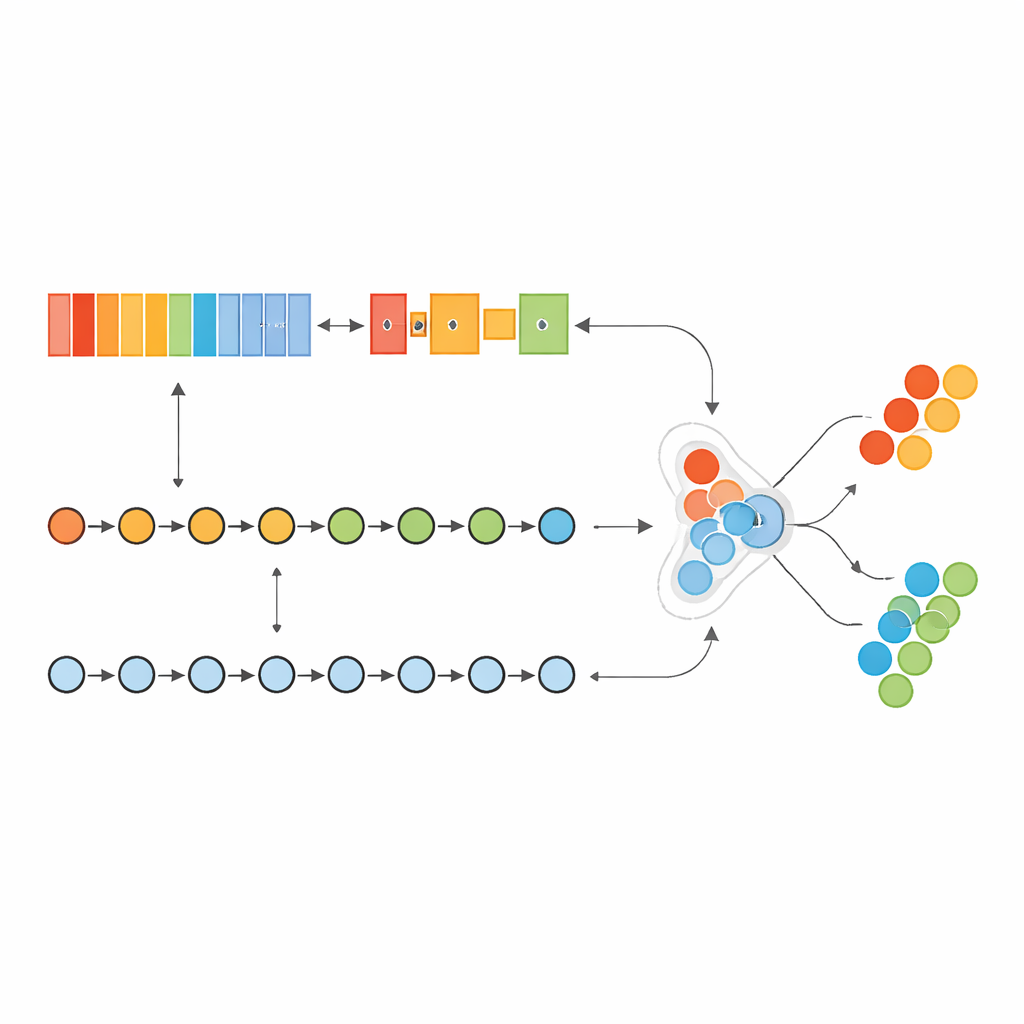
Teaching a dual-brain model to read gene patterns
The authors built a “hybrid” deep learning model that acts a bit like two specialized brains working together. One part, inspired by image analysis, scans an ordered list of genes to detect local patterns—clusters of genes whose activity together signals cancer. The other part treats the same genes as a sequence, learning how early “driver” genes and later “downstream” genes influence one another across the list. By combining these two views, the model can capture both short-range and long-range relationships within the tumor’s genetic fingerprint.
Finding a stable core set of signal genes
Instead of feeding all 17,815 measured genes into the model, the team designed a strict, “leakage-free” pipeline to select only the most informative ones. Using a standard measure of correlation within repeated cross-checking loops, they repeatedly ranked genes by how strongly their activity tracked with cancer status. They then kept only the genes that consistently rose to the top across all training splits, resulting in a stable signature of 236 genes. The researchers also mapped how these genes interact with one another, showing that many form tightly connected networks related to tumor growth, metabolism, immunity, and the surrounding tissue environment—evidence that the chosen set reflects real biology, not random noise.
Putting the model to the test
The hybrid system was trained and tuned on breast cancer samples from The Cancer Genome Atlas and then challenged with an entirely separate dataset known as METABRIC. To handle the fact that cancer samples far outnumber normal samples, the authors did not create artificial data; instead, they adjusted how much the model “cares” about mistakes on the rarer class. After an automated search for the best settings, the model reached near-perfect scores on its main dataset, correctly flagging almost all cancer cases and making virtually no false alarms. Importantly, performance stayed extremely high and very stable even when the model was applied to the external METABRIC cohort, suggesting that the approach can generalize beyond one study or hospital.
What this means for future care
In simple terms, this work delivers a finely tuned, two-part AI that reads a compact 236-gene code to tell cancerous from non-cancerous breast samples with remarkable accuracy and consistency, even under noisy conditions. While the current study looks only at gene activity and uses past patient data, its methods lay the groundwork for future tools that could combine multiple data types—such as tissue images and additional molecular layers—and provide clear explanations of which genes drive each prediction. With further validation in prospective clinical studies, such a system could become a universal backbone for precision breast cancer diagnosis, helping doctors tailor treatment using the genetic “signature” of each patient’s tumor.
Citation: Hesham, F., Abbassy, M.M. & Abdalla, M. Gene driven analytical learning model for accurate breast cancer diagnosis. Sci Rep 16, 8155 (2026). https://doi.org/10.1038/s41598-026-39430-6
Keywords: breast cancer diagnosis, gene expression, deep learning, CNN-BiLSTM, precision oncology