Clear Sky Science · en
Integrating multi-omics analysis identifies DNA damage-related gene CLSPN as a biomarker in gastric cancer
Why this research matters
Stomach cancer remains one of the world’s deadliest cancers, in large part because it is often found late and lacks reliable early warning signs. This study digs deep into the genetic “scratch marks” left when DNA is damaged, and identifies a gene called CLSPN as a promising marker that may help doctors detect gastric cancer earlier, understand how aggressive it is, and potentially tailor treatments more precisely.
Looking for danger signals in stomach tumors
The researchers started from a simple idea: if damage to DNA helps drive stomach cancer, then genes involved in coping with that damage might carry important clues about who gets sick and how their disease behaves. To test this, they assembled several large datasets. These included bulk RNA sequencing, which measures average gene activity across whole tumor samples; single-cell RNA sequencing, which looks at thousands of individual cells one by one; and spatial transcriptomics, which maps gene activity back onto the tissue slide. They also pulled together a long list of genes already linked to DNA damage. Using these resources, they asked which DNA damage–related genes were most strongly tied to gastric cancer and to patient survival.
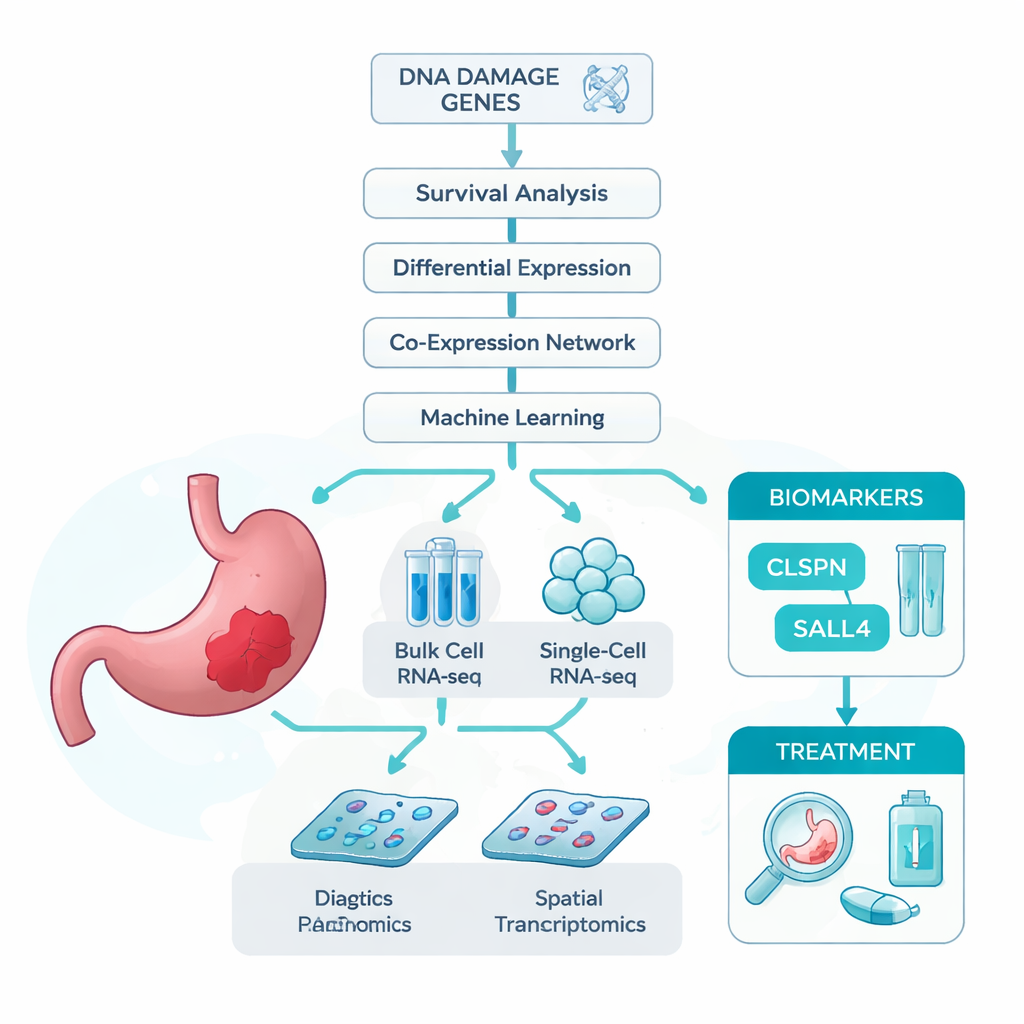
Using smart algorithms to find key genes
Because thousands of genes change in cancer, the team turned to a suite of statistical and machine learning tools to narrow the field. They first filtered genes that were both misregulated in tumors and clearly related to how long patients lived. They then grouped genes by how they rose and fell together, and cross-checked this with the DNA damage list. This multi-step screening produced 13 strong candidate genes. To see which of these mattered most for telling tumor from normal tissue, they trained seven different machine learning models, each ranking genes by how useful they were for classification. Across all models, two genes—CLSPN and SALL4—consistently rose to the top, with accuracy scores near or above 97% and areas under the ROC curve above 0.96, suggesting these genes have excellent diagnostic potential.
Zooming in on CLSPN inside the tumor
Among the two leading genes, CLSPN stood out as the most informative overall, so the authors examined it in greater detail. In single-cell data from over 30,000 cells, CLSPN was much more active in tumor cells than in nearby normal or supporting cells, pointing to a tumor-specific role. Spatial mapping of patient tissue sections showed a similar pattern: CLSPN expression clustered in regions pathologists recognized as cancer. When the team split tumor cells into “high CLSPN” and “low CLSPN” groups, the high group showed stronger activity in pathways tied to DNA copying, cell division, and repair—processes that, when overactive or misdirected, can fuel uncontrolled growth. Pseudotime analysis, which reconstructs how cells change over a notional timeline, suggested that as tumor cells progress, CLSPN levels tend to rise, hinting that CLSPN upregulation may be part of the shift toward a more malignant state.
Links to the immune system and treatment response
The study also explored how CLSPN relates to the tumor’s surroundings and to potential therapies. Tumors with higher CLSPN expression showed different mixes of immune cells, particularly macrophages and certain T and NK cells, suggesting CLSPN might help shape an immune-suppressive environment. Drug sensitivity modeling indicated that cancers with more CLSPN could be more vulnerable to several chemotherapy and targeted agents, as higher CLSPN levels were associated with lower predicted doses needed to inhibit growth. Importantly, when the authors stained real tissue samples from 70 patients, CLSPN protein levels were clearly higher in cancer than in normal stomach and were linked with larger tumors, deeper invasion, lymph node spread, and shorter overall survival.
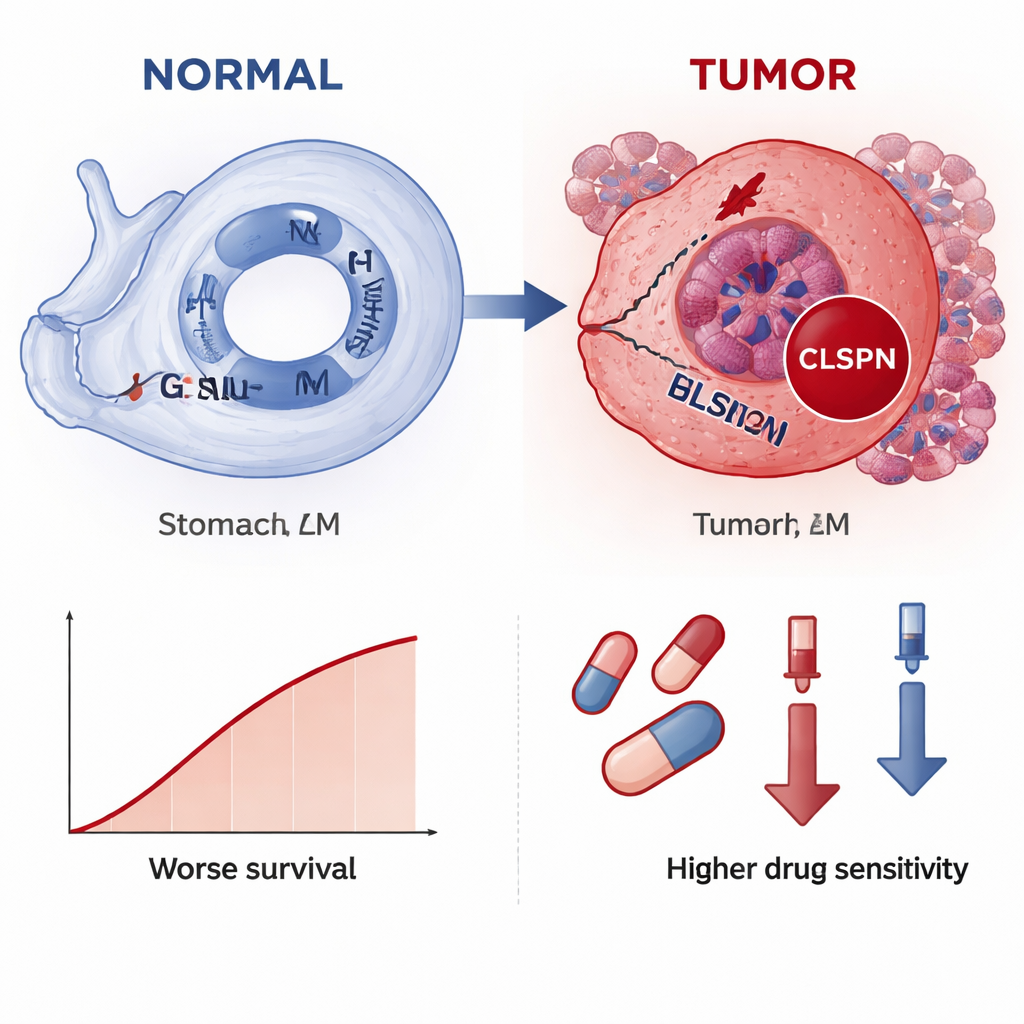
What this could mean for patients
In everyday terms, the study suggests that CLSPN behaves like a red warning light for stomach cancer: it turns on mostly in tumor cells, glows brighter in more advanced disease, and signals a higher risk of poor outcome. Because it can be detected both at the RNA level and by standard hospital staining methods, CLSPN could become a practical biomarker to help pathologists confirm gastric cancer, stratify patients by risk, and perhaps guide drug choices. While more clinical testing is needed before it can be used routinely, this work highlights how reading the footprints of DNA damage across many data layers can uncover new tools for earlier diagnosis and more personalized treatment of gastric cancer.
Citation: Ma, Q., Yang, X., Sun, N. et al. Integrating multi-omics analysis identifies DNA damage-related gene CLSPN as a biomarker in gastric cancer. Sci Rep 16, 7789 (2026). https://doi.org/10.1038/s41598-026-39387-6
Keywords: gastric cancer, DNA damage, CLSPN, biomarker, single-cell analysis