Clear Sky Science · en
The DNA methylation landscape of naturally short-lived killifish
Tiny fish with big clues about aging
Why do some animals race through life while others linger for years? This study tackles that puzzle using annual killifish, tiny African fish that live for only a few months yet show many of the same aging symptoms as humans. By mapping how chemical tags on their DNA change over time, the researchers ask whether these short-lived fish follow the same “epigenetic” aging patterns seen in people and other mammals—and whether those patterns could one day help predict health and lifespan in a simple laboratory animal.
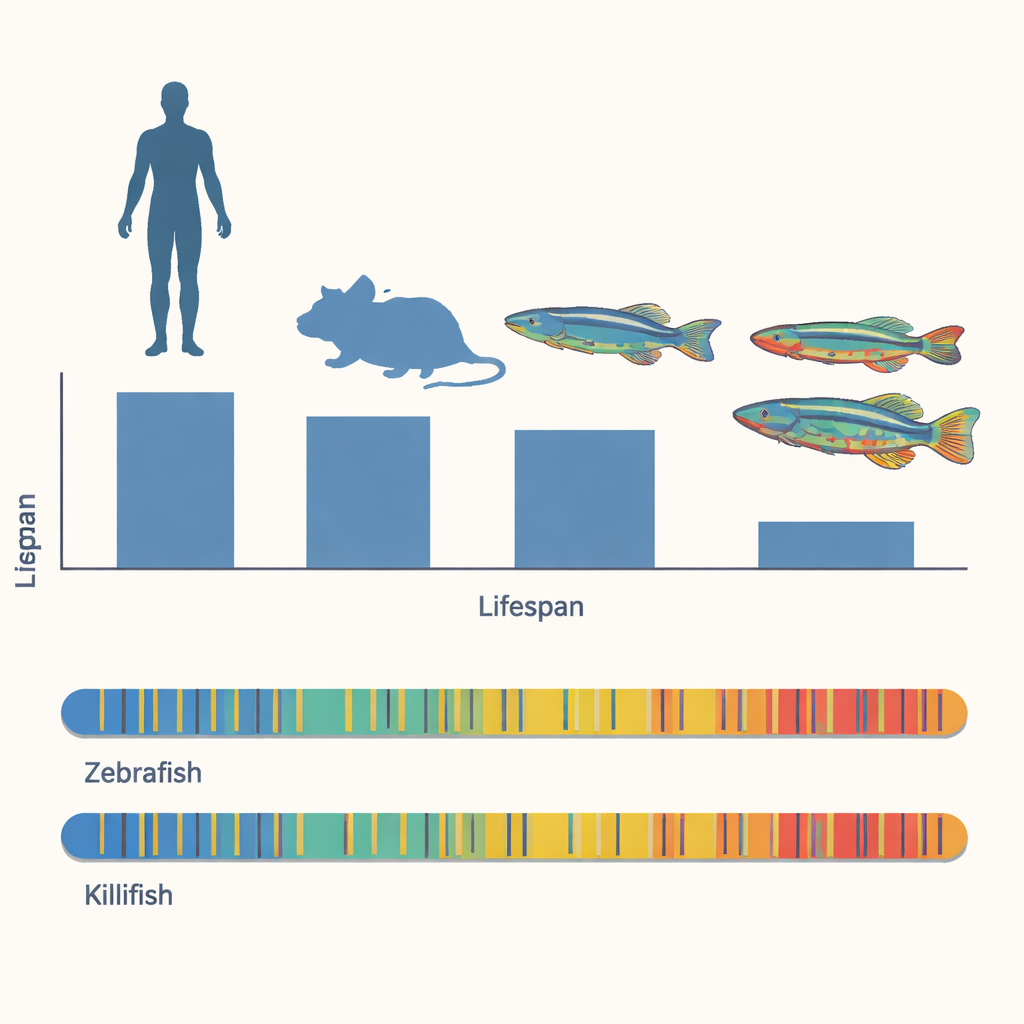
Short lives in an unpredictable world
Annual killifish have evolved to survive in temporary puddles that appear during the rainy season and vanish soon after. Their eggs wait in the dry mud, then hatch and grow at remarkable speed once water returns. One species, Nothobranchius furzeri, typically lives only three to six months; its close cousin N. orthonotus can reach about ten months under similar conditions. Both species develop familiar signs of aging—reduced fertility, slower swimming, changes in skin and eye color, memory problems, immune decline, gut imbalance, and tumors. Because these changes unfold in less than a year, the fish offer a rare chance to study the biology of aging on a fast-forward schedule.
Looking beyond the genetic blueprint
The researchers wanted to know whether the killifish’s rapid aging is written into the structure of its genome or instead into how that genome is regulated. They first compared basic genome features of the two killifish with those of humans, mice, zebrafish, and the worm C. elegans. Despite their dramatically shorter lifespans, the killifish genomes are similar in size and gene layout to zebrafish. Both fish devote about half of their DNA to repetitive sequences, especially mobile genetic elements known as transposable elements, and show comparable distributions of the DNA letters where methyl groups—the chemical tags in question—can attach. These broad similarities suggest that the killifish’s short life cannot be explained simply by having a smaller or more compact genome.
DNA tags that shift with tissue and time
To probe regulation instead of raw sequence, the team produced high-resolution maps of DNA methylation—the pattern of methyl chemical groups attached to cytosine bases—across the entire genomes of both species. They analyzed brain, liver, and heart from young and old N. furzeri, and brains from young, old, and very old N. orthonotus. Overall, the killifish methylation landscape resembled that of mammals: most sites were either heavily tagged or barely tagged, and regions near gene start sites tended to have fewer tags. Differences between tissues were much stronger than differences between ages. Brain, liver, and heart each carried distinct methylation signatures, and brain-specific low-tag regions often sat near genes important for nerve cell identity, hinting that these patterns help define and maintain each organ’s function.
Subtle aging fingerprints and mobile DNA
Age-related changes were present but modest. Across the genome as a whole, methylation levels remained largely stable with age in both killifish species. However, closer inspection revealed thousands of specific regions where methylation did shift between young and old animals. Many of these changes occurred within or near transposable elements—the mobile DNA sequences that make up a large fraction of the killifish genome. Different tissues showed partially overlapping sets of age-sensitive elements, suggesting both shared and organ-specific aging effects. In the longer-lived N. orthonotus, detailed brain analyses uncovered groups of methylation sites whose levels changed in a way that tracked the fish’s age, and these sites could be combined into a statistical model that roughly predicted whether an individual was young, old, or very old.
What these tiny fish tell us about growing old
The study shows that even in a vertebrate that lives only months, there are recognizable DNA methylation changes as animals age, much like those used to build “epigenetic clocks” in humans and mice. Yet the shifts in killifish are relatively small and spread across many sites rather than concentrated in a few key pathways. This may reflect the compressed lifespan of these fish: there may be less time for gradual epigenetic drift to build up. By delivering the first comprehensive methylation maps for annual killifish, the work lays essential groundwork for turning these animals into a fast, flexible system for testing how genes, environment, and potential treatments influence the pace of biological aging.
Citation: Steiger, M., Singh, N., Tyers, A.M. et al. The DNA methylation landscape of naturally short-lived killifish. Sci Rep 16, 7173 (2026). https://doi.org/10.1038/s41598-026-39352-3
Keywords: epigenetic aging, DNA methylation, annual killifish, transposable elements, lifespan biology