Clear Sky Science · en
A polymer physics model of the interphase cell nucleus for radiobiological simulations
Why the Shape of Our DNA Home Matters
Radiation is a double-edged sword in medicine: it can kill cancer cells, but it can also damage healthy tissue. At the heart of both risks and benefits lies our DNA, packed tightly inside the cell nucleus. This paper shows how a detailed, physics-inspired 3D model of that nuclear “DNA city” can help scientists predict how radiation breaks and mis-repairs chromosomes, potentially improving cancer treatment and space radiation risk estimates.
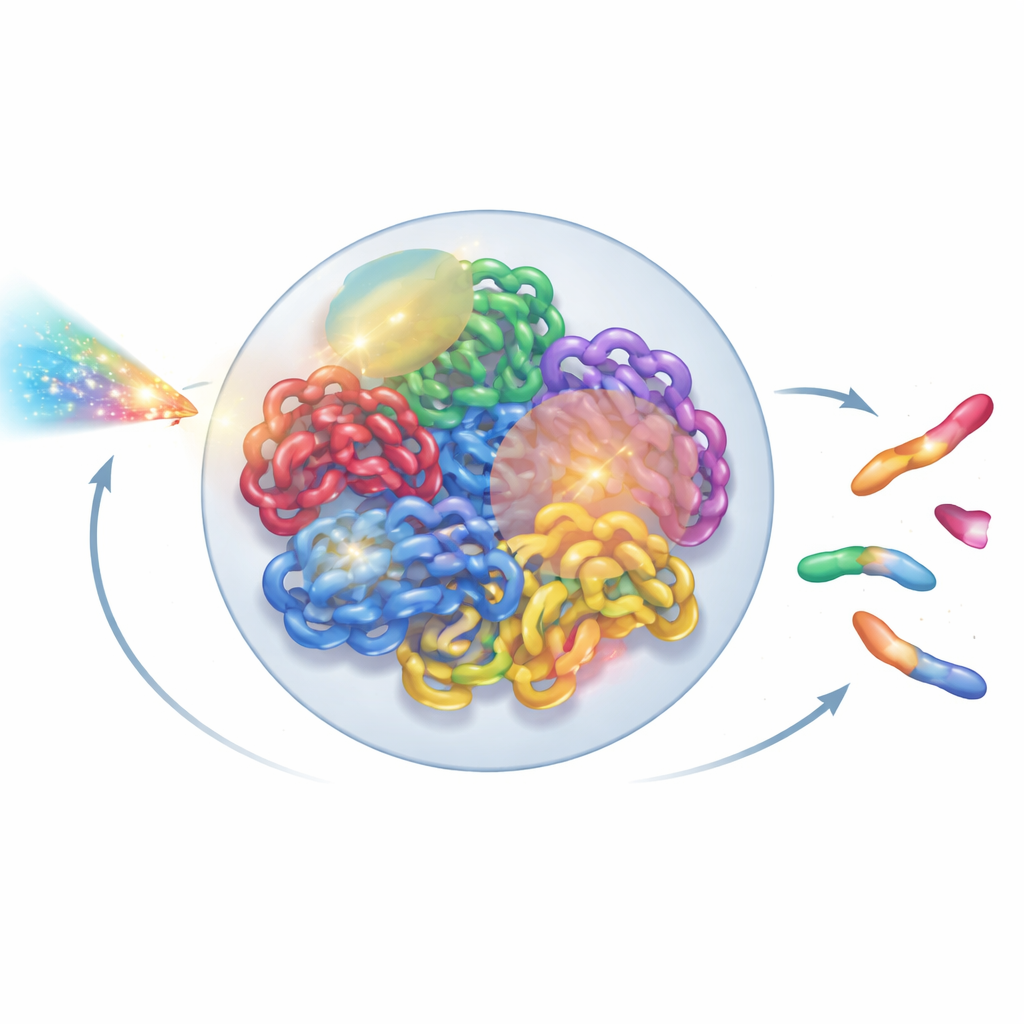
Seeing the Genome as a Living Structure
Inside each human cell, two meters of DNA are folded into a tiny nucleus only a few micrometers across. This folding is not random: each chromosome tends to occupy its own territory, which is further divided into neighborhoods and looping domains. New experimental tools, such as super-resolution microscopy and Hi-C contact maps, have revealed this intricate 3D layout in remarkable detail. Yet most radiation models still treat DNA as a simplified tangle of lines or idealized loops, glossing over this higher-order organization. The authors set out to build a whole-nucleus model that respects what biologists have learned about real chromatin architecture, while still being fast enough to run on ordinary lab computers.
Building a Virtual Nucleus with Polymer Physics
The team modeled chromosomes as flexible chains, borrowing ideas from polymer physics, the same field used to understand plastics and soft materials. They broke the problem into three relaxation stages to tame the huge computational cost. First, they placed large building blocks called chromatin domains throughout the nucleus, allowing them to repel one another, stretch along their lengths, and cluster into broader subcompartments. Second, they zoomed in within each domain to lay out the finer chromatin fibers, including looped regions held together by binding proteins. Third, they let all fibers interact freely again inside the nuclear boundary to approach a realistic final configuration. By splitting the process this way and adding hierarchical search and parallel computing tricks, they cut the runtime from an impractical four months to about three hours per virtual nucleus on a 64-core workstation.
From Radiation Tracks to DNA Breaks and Repairs
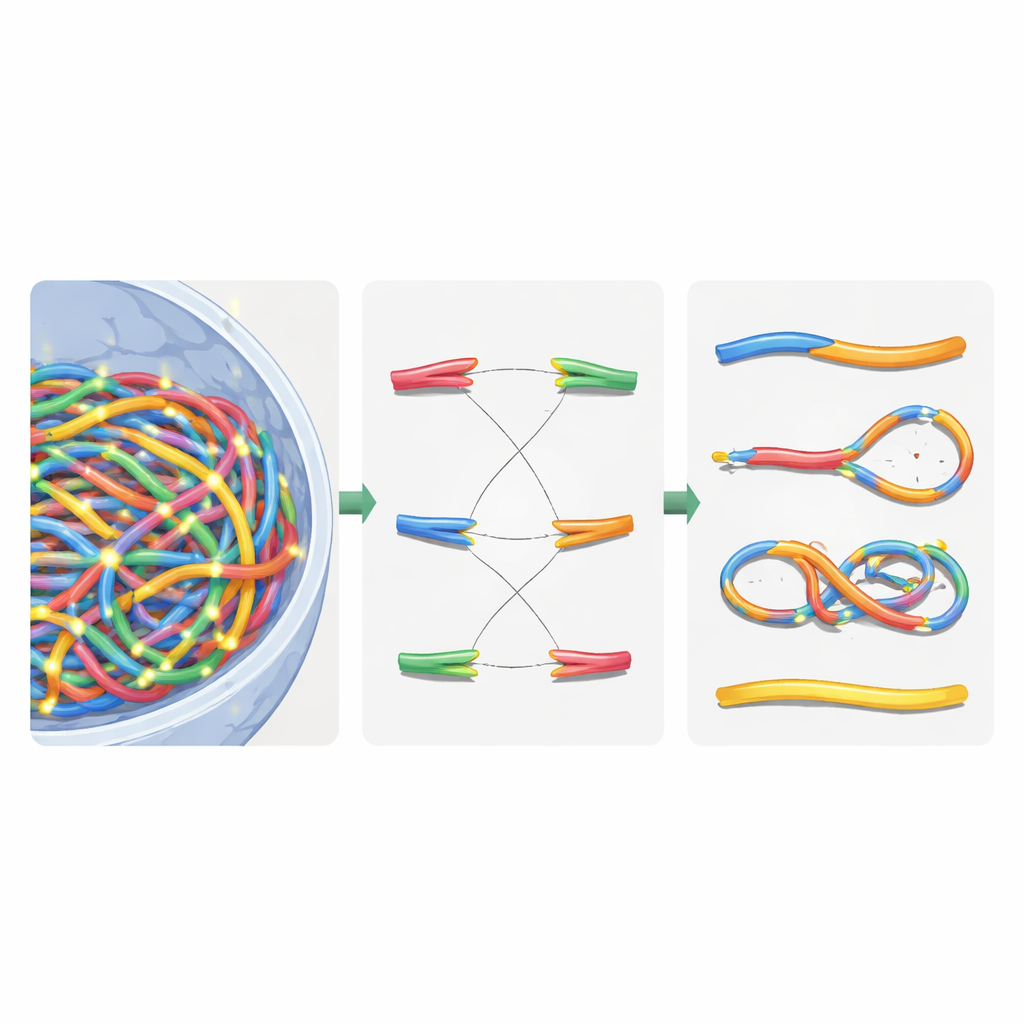
Once the virtual nucleus was built, the authors overlaid detailed radiation tracks generated by Geant4-DNA simulations. They used a hierarchical search to find, for each ionization event, which chromatin domain, fiber segment, and finally which nucleotide was closest, without ever storing all ten billion nucleotide positions in memory. Instead, they kept a reusable chromatin template and applied clever coordinate transformations on the fly. Ionizations in the DNA backbone were treated as strand breaks, and nearby breaks on opposite strands became double-strand breaks. To mimic the cell’s repair machinery, the authors introduced a distance-based rejoining rule: free DNA ends that lie close together are more likely to rejoin. Rejoining decisions were made step by step until all ends were either paired or left unrepaired, recreating both correct repairs and mis-joined fragments.
Sorting Broken Chromosomes with Network Logic
After this virtual repair, the nucleus contained a mix of reassembled chromosomes and leftover pieces. To classify what had happened, the authors translated the repaired genome into an undirected graph: chromosome segments became edges, and their ends and breakpoints became nodes. Using a standard connected-component algorithm, they could automatically recover each new chromosome-like object and count key features such as the number of centromeres (attachment points for cell division) and telomeres (natural ends). From this, they identified dicentrics (chromosomes with two centromeres), rings, and deletions. They then compared their simulations to classic experiments in human skin cells exposed to gamma rays and alpha particles, finding that total chromosome aberrations were reproduced within about 20% of measured values, a substantial improvement over earlier models.
What This Means for Patients and Astronauts
In simple terms, this work shows that “where” DNA sits in the nucleus, and “how” it folds, strongly influence how radiation damage adds up to dangerous chromosome changes. By combining realistic 3D genome architecture, efficient physics-based simulation, and graph-based readout of broken chromosomes, the authors provide a powerful bridge between structural biology and radiobiology. They argue that future refinements—such as adding detailed repair timing and using whole-genome sequencing data—could make this approach even better at predicting the biological impact of high-energy particles used in cancer therapy or encountered in deep space. Ultimately, better models of the DNA home inside our cells may help tailor safer, more effective radiation treatments and sharpen our understanding of long-term radiation risks.
Citation: Zhao, M., Huang, G., Xu, Z. et al. A polymer physics model of the interphase cell nucleus for radiobiological simulations. Sci Rep 16, 7913 (2026). https://doi.org/10.1038/s41598-026-39234-8
Keywords: radiation-induced DNA damage, chromatin 3D architecture, chromosome aberrations, polymer physics modeling, particle therapy