Clear Sky Science · en
In Silico identification of inhalable small-molecule IL-33/ST2 antagonists for severe type-2-high asthma endotypes
Why This Matters for Breathing
For millions of people with severe asthma, even modern inhalers and advanced injections can leave them short of breath and at risk of dangerous attacks. This study explores whether familiar, pill‑like drugs could be redesigned to work as inhaled treatments that block one of asthma’s upstream "alarm" signals in the lungs. Using only computer simulations, the researchers searched for small molecules that might fit onto a key receptor involved in severe, allergy‑driven asthma, laying the groundwork for future laboratory testing rather than offering a ready‑to‑use medicine.
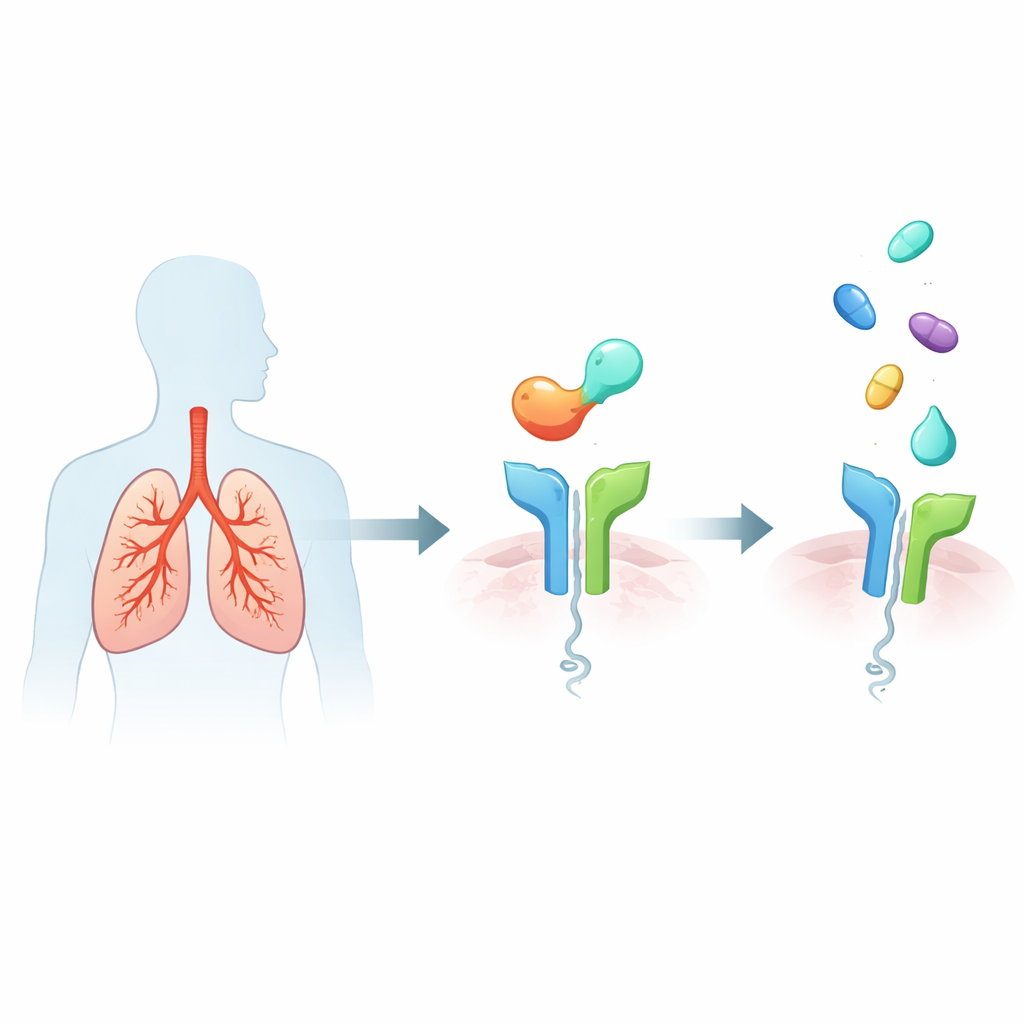
The Alarm System Inside Asthmatic Lungs
Severe type‑2‑high asthma is marked by a hyperactive immune response that fills the airways with mucus and inflammatory cells, narrows the breathing tubes, and damages lung structure over time. A major trigger is a protein called IL‑33, released by stressed airway lining cells. IL‑33 docks onto a receptor called ST2 on various immune cells, which then release powerful messenger molecules that drive hallmark asthma features such as wheezing, flare‑ups, and long‑term airway remodeling. Because higher IL‑33/ST2 activity is linked to worse symptoms and more frequent attacks, ST2 has emerged as an attractive upstream target: block ST2, and you may quiet many of the downstream inflammatory signals at once.
From Antibodies to Everyday‑Style Drugs
Several current therapies for severe asthma use injected antibodies that mop up inflammatory proteins in the blood or block their receptors. While effective for some, these biologic drugs are expensive, must be given by injection, and focus mainly on signals further down the pathway. In contrast, small‑molecule drugs—more like traditional pills or inhalers—could be manufactured at scale, stored easily, and potentially delivered directly into the lungs. The challenge is that ST2 binds IL‑33 through a broad protein‑protein interface, which is harder for small molecules to disrupt than a neat, deep pocket on an enzyme. The authors therefore turned to high‑resolution structural data for the ST2–IL‑33 complex and modern in silico tools to see whether any known anti‑inflammatory drugs might unexpectedly grip part of this interface.
Searching the Receptor Surface by Computer
Using the crystal structure of human ST2 bound to IL‑33, the team first removed IL‑33 to expose the receptor’s binding surface and used a pocket‑finding program to pinpoint the most promising region for a small molecule to nestle in. They then selected ten clinically relevant small molecules already used in asthma or related immune diseases—such as the leukotriene blockers Montelukast and Zafirlukast, immune‑modulating thalidomide relatives, and JAK inhibitors—plus a previously reported ST2‑focused molecule as a reference. With virtual docking software, each compound was positioned thousands of times on the ST2 surface, and the best‑fitting poses were scored by predicted binding strength. Zafirlukast surfaced as the top‑ranked scaffold, slightly outperforming Montelukast and the known ST2‑directed comparator.
Tuning a Lead Molecule on Screen
Because Zafirlukast showed the most promising balance of predicted binding and general drug‑like behavior, the authors used an AI‑guided design tool to create a modified version of this molecule. The goal was to retain its grip on ST2 while improving its predicted safety profile. The redesigned analogue had a very similar docking score and was predicted to be less toxic in silico. Detailed contact maps suggested that both the parent and modified compounds touch several of the same ST2 amino acids that normally contact IL‑33, hinting that they occupy functionally important parts of the interface. Extended molecular dynamics simulations—half a microsecond of virtual motion in water and salt—showed that the receptor and ligand both remained flexible, with the small molecule sampling different positions along the surface rather than locking rigidly into place. Calculations of interaction energies over time pointed to meaningful but dynamic association, consistent with a molecule probing a relatively flat protein‑protein surface.
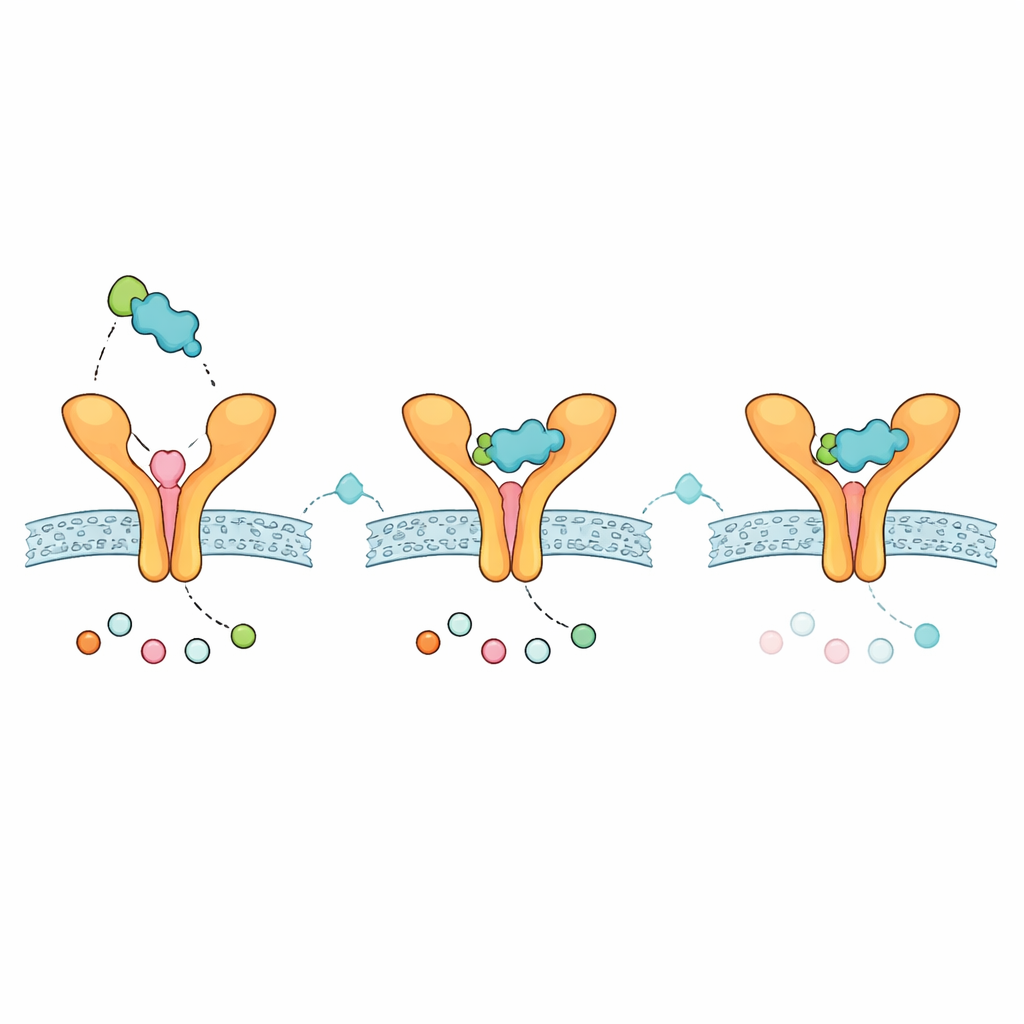
Clues About How a Future Inhaler Might Behave
To explore whether the modified Zafirlukast could plausibly function as a medicine, the team ran standard computational checks of absorption, distribution, metabolism, excretion, and toxicity. The analogue satisfied common oral drug‑likeness rules, showed moderate size and fat‑solubility, and was predicted to stay out of the brain—features that may be favorable for a lung‑focused treatment with limited whole‑body side effects. Solubility estimates varied by model, implying that careful formulation would be needed, especially for non‑oral routes such as inhalation. The compound was also predicted to interact with certain liver enzymes, suggesting that potential drug–drug interactions would need attention if it were ever used alongside other medications.
What This Work Really Shows
Rather than delivering a new asthma drug, this study demonstrates a step‑by‑step computational strategy for finding and refining small molecules that may interfere with the IL‑33/ST2 alarm system at the surface of airway cells. It highlights Zafirlukast and an AI‑optimized relative as plausible starting points for laboratory experiments, showing that they can occupy key parts of the ST2 interface and have broadly acceptable predicted safety and drug‑like properties. However, all of the evidence is virtual: the work does not prove that these molecules block IL‑33 from binding ST2, dampen inflammatory signaling, or help patients breathe more easily. Those answers will require biochemical tests, cell studies, animal models, and eventually clinical trials. For now, the study offers a computational roadmap and a short list of candidate scaffolds for scientists aiming to turn the body’s own asthma alarm switch into a practical, inhalable drug target.
Citation: Sun, G., Liu, Q., Yu, M. et al. In Silico identification of inhalable small-molecule IL-33/ST2 antagonists for severe type-2-high asthma endotypes. Sci Rep 16, 7996 (2026). https://doi.org/10.1038/s41598-026-39027-z
Keywords: severe asthma, IL-33 ST2 pathway, small-molecule antagonists, in silico drug design, inhaled therapeutics