Clear Sky Science · en
Novel marker genes for simultaneous detection of Salmonella, EHEC O157:H7, and Cronobacter
Why this matters for everyday food safety
From salad greens to burgers to baby formula, the foods we trust can occasionally harbor dangerous germs. Three of the most worrisome culprits—certain strains of E. coli, Salmonella, and a lesser-known bacterium called Cronobacter—can cause severe illness, especially in young children and vulnerable adults. This study shows how scientists are using massive DNA databases and clever lab tests to spot all three of these threats at once, quickly and accurately, before contaminated food reaches your table.
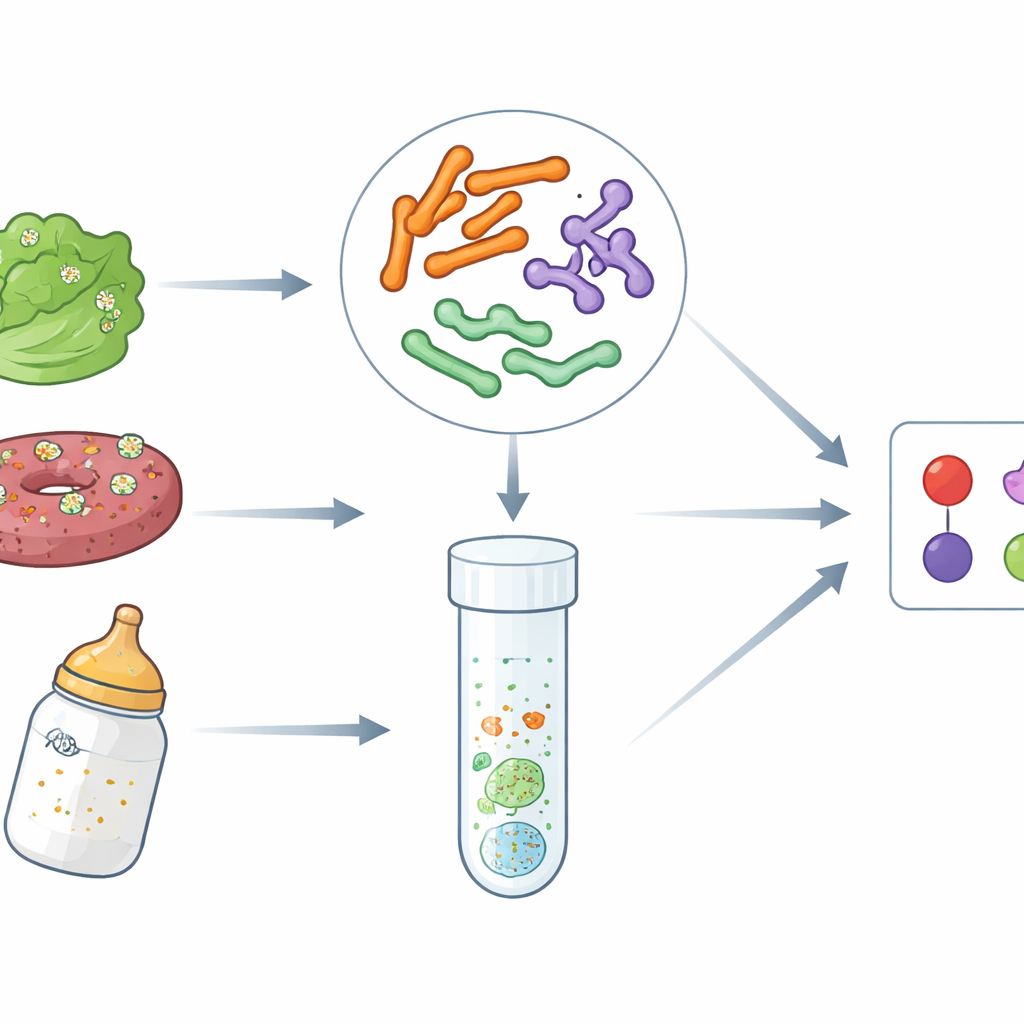
Hidden dangers in common foods
Food-borne illnesses have risen over the past decade, driven in part by global supply chains and the sheer volume of processed and ready-to-eat foods. EHEC O157:H7, a highly toxic form of E. coli, can trigger bloody diarrhea and kidney failure. Salmonella causes millions of cases of food poisoning every year, and Cronobacter, while less famous, can be deadly for newborns, especially in connection with powdered infant formula. Traditional lab methods often focus on one germ at a time and require days of culturing, which slows down outbreak investigations and routine screening. The authors set out to build faster DNA-based tests that can flag these three pathogens together in a single, streamlined procedure.
Finding “barcodes” in a sea of DNA
To do this, the team first needed reliable genetic “barcodes”—short pieces of DNA that are present in nearly all strains of a target germ but absent from others. Instead of searching a handful of genomes, they tapped into an enormous public collection of 1.46 million bacterial genomes from 50 different genera. For EHEC O157:H7, they compared thousands of E. coli genomes and sifted through more than 100,000 accessory gene families to find DNA sequences unique to this dangerous subtype. After several rounds of filtering and cross-checking against over a million non-E. coli genomes, they landed on a gene called z0340 as a highly specific marker for EHEC O157:H7. Using a similar strategy on more than half a million Salmonella genomes and nearly a million other bacterial genomes, they identified another gene, sbcC, as a reliable marker for the Salmonella group as a whole.
Turning markers into real-world tests
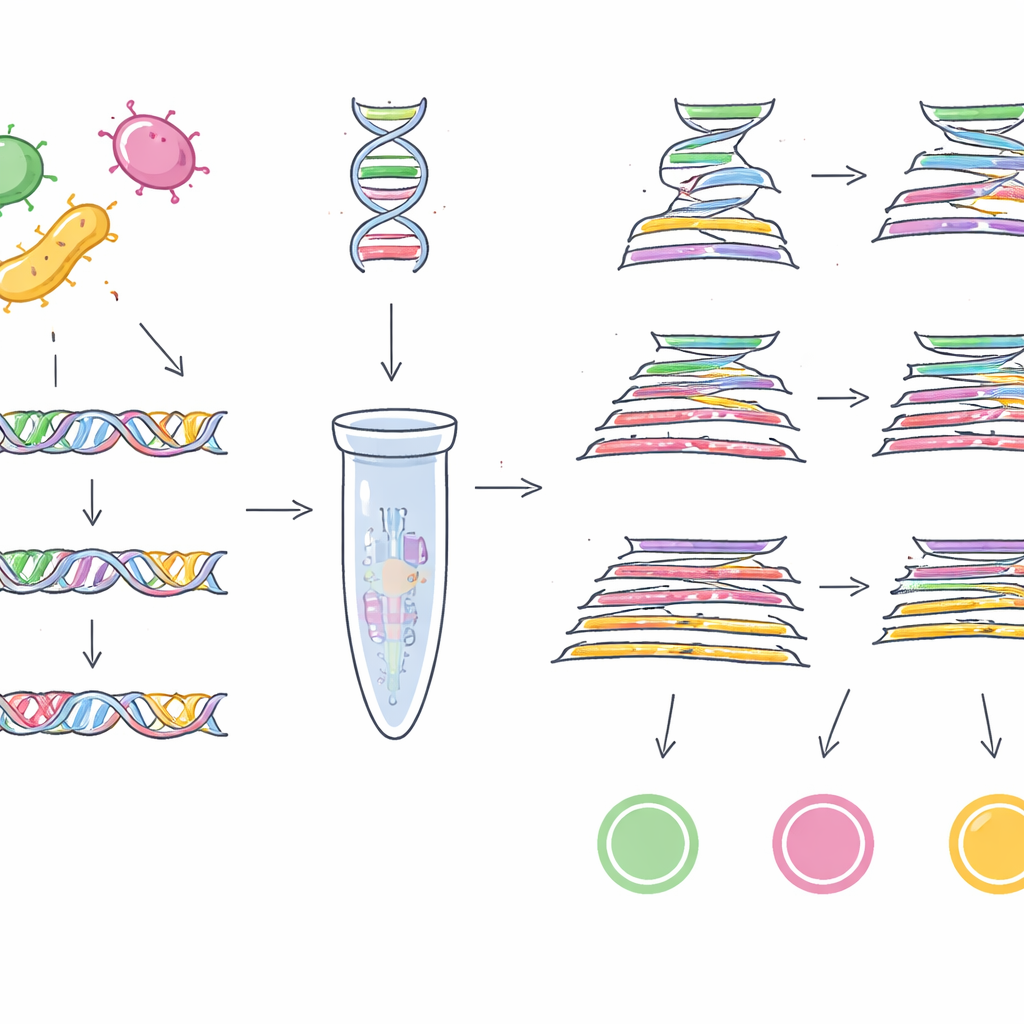
With these two new barcodes in hand—plus a Cronobacter-specific gene, ygcB, that the group previously discovered—the researchers designed laboratory tests that can detect all three pathogens in one go. They built multiplex PCR assays, which act like molecular photocopiers targeting specific stretches of DNA, and a more sensitive TaqMan quantitative PCR version that measures how much target DNA is present in real time. When these tests were challenged with 23 strains of the three pathogens and 100 strains of other common bacteria, they correctly picked out only the intended targets every time, showing 100% specificity. The basic multiplex PCR could detect as little as 1 picogram of DNA per microliter, while the TaqMan version went down to 0.5 picograms, indicating high sensitivity.
Putting the methods to the food test
Lab accuracy is only useful if the tests work in messy, real-world samples. To check this, the scientists deliberately contaminated lettuce, ground beef, and powdered infant formula with known amounts of EHEC O157:H7, Salmonella, or Cronobacter. After standard enrichment steps to allow any bacteria present to multiply, they applied their multiplex PCR and TaqMan qPCR assays. In every case, the methods correctly detected the introduced pathogen and did not produce false alarms in negative controls. The performance was consistent across all three food types, suggesting that fats, plant components, or complex ingredients did not noticeably interfere with the detection.
Limitations and future improvements
Despite the strong results, the authors note that their validation panel still represents only a tiny fraction of the microbial diversity found in nature. For example, they tested only one EHEC O157:H7 strain in the lab, even though their genome-based screening covered tens of thousands of such strains. They also worked with relatively high levels of contamination compared with the extremely low numbers of bacteria that can still cause disease. Future work will need to test many more real-world isolates, examine naturally contaminated foods, and add internal controls to guard against substances in food that can inhibit DNA amplification.
What this means for consumers
In everyday terms, this study shows that carefully chosen DNA barcodes can let food safety labs screen for multiple high-risk germs at once, with high accuracy and in less time than traditional culture-based methods. By mining millions of genomes, the researchers identified three genetic signatures—one for EHEC O157:H7, one for Salmonella, and one for Cronobacter—that appear to be both highly specific and stable. The resulting tests could strengthen routine monitoring of foods like salad greens, meats, and infant formula, catching contamination earlier and more reliably. Beyond these three pathogens, the same genome-first approach could be used to design rapid tests for many other dangerous microbes, offering a powerful new tool to keep the global food supply safer.
Citation: Zhang, H., Xiong, P., Lu, Z. et al. Novel marker genes for simultaneous detection of Salmonella, EHEC O157:H7, and Cronobacter. Sci Rep 16, 9362 (2026). https://doi.org/10.1038/s41598-026-38990-x
Keywords: foodborne pathogens, multiplex PCR, genomic markers, Salmonella and EHEC, Cronobacter detection