Clear Sky Science · en
Structure-based modeling reveals molecular basis for CYP153A6’s novel activity toward toluene derivatives
Turning Tough Chemical Jobs Over to Nature
Chemists and industry rely on reactions that add oxygen to stubborn carbon–hydrogen bonds, for example to clean up pollutants or to build ingredients for drugs and fragrances. Doing this cleanly, without harsh chemicals or heavy metal waste, is hard. This study looks at a natural catalyst, an enzyme called CYP153A6 from a soil bacterium, and asks a simple question with big implications: can this enzyme be guided to selectively "upgrade" toluene-like chemicals—common components of fuels and solvents—into more useful and environmentally friendly products?
An Enzyme that Picks a Single Spot
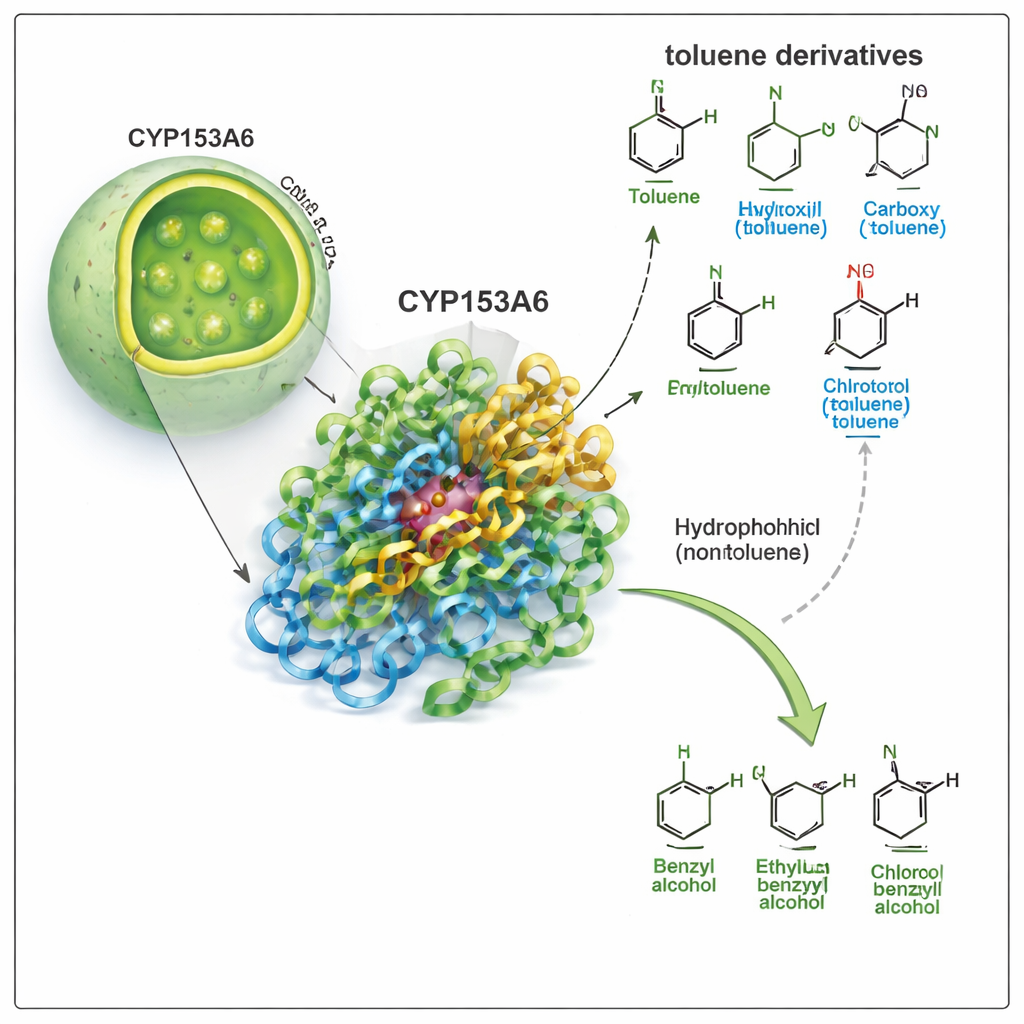
CYP153A6 belongs to the large cytochrome P450 family, nature’s versatile oxidation specialists. Unlike many industrial catalysts, this enzyme works in water, at moderate temperature, and chooses very specific positions on a molecule. Earlier work showed that CYP153A6 adds oxygen to the tail ends of simple hydrocarbon chains and certain plant-based scent molecules. Here, the authors ask whether it can also attack the “benzylic” position—the small side-chain attached to a benzene ring—in toluene and related compounds. That transformation turns toluene derivatives into benzyl alcohols, valuable building blocks for pharmaceuticals, flavors, and fine chemicals.
Testing Which Molecules the Enzyme Likes
The team first ran biotransformation experiments using engineered E. coli cells that produce CYP153A6, either as whole cells or as cell-free extracts. They exposed the enzyme to a panel of toluene derivatives, changing the group attached to the aromatic ring (such as methyl, chlorine, methoxy, hydroxyl, or nitro) and its position (para, meta, or ortho). The enzyme performed very well on apolar or only slightly polar compounds, such as p-cymene, p-xylene, p-methylanisole, and p-chlorotoluene, consistently converting the benzylic methyl group into a benzyl alcohol. Strikingly, it completely ignored more strongly polar molecules like p-cresol, p-methyl benzyl alcohol, and p-nitrotoluene, even though these are structurally similar. This pattern suggested that the enzyme’s pocket is strongly hydrophobic and disfavors polar groups.
Building a 3D Map of the Molecular Machine
Because the three-dimensional structure of CYP153A6 had not been solved experimentally, the researchers built a high-quality computer model using a closely related P450 whose structure is known. They validated this model by checking its geometry and running long molecular dynamics simulations to see whether the protein stayed stable over time, which it did. The simulations revealed a buried active site connected to the outside by narrow tunnels through which substrates enter and products leave. These tunnels are shaped and gated by flexible loops on the protein surface, helping explain how the enzyme can guide hydrophobic molecules down to the reactive heme iron center while allowing water and oxidized products to escape.
Watching Molecules Dock, Move, and React
With the model in hand, the authors simulated how different toluene derivatives bind inside CYP153A6. They used an approach called ensemble docking, trying many subtly different protein shapes and several electronic states of the heme iron, to identify which setups best distinguished real substrates from non-substrates. The winning scenario used the iron in its resting ferric state and no extra water molecules in the pocket, and it correctly favored known substrates. Follow-up simulations tracked, frame by frame, how each molecule sat relative to the heme. Good substrates like p-cymene and p-xylene stayed close to the iron with their benzylic methyl group pointed at it, and their predicted binding energies were strongly favorable. Poor substrates tended to drift away, adopt awkward orientations, or interact unfavorably with the mostly oily side chains that line the pocket.
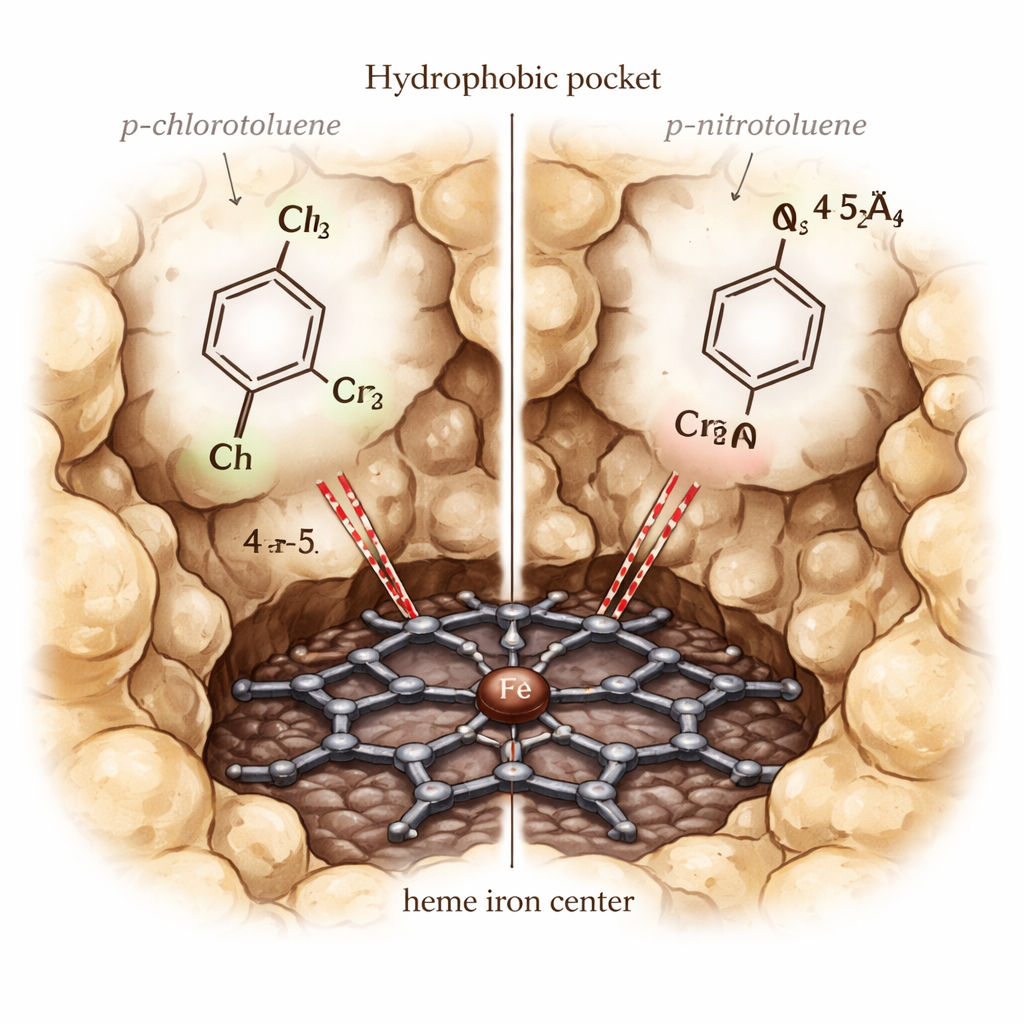
Why One Look-Alike Reacts and Another Does Not
One puzzle stood out: p-chlorotoluene is efficiently oxidized by CYP153A6, while p-nitrotoluene is not, even though both fit into the pocket and bind with similar strength in the simulations. To resolve this, the authors carried out more detailed quantum mechanics/molecular mechanics (QM/MM) calculations on the highly reactive “Compound I” state of the enzyme. These calculations showed that the nitro group draws so much electron density away from the benzylic carbon that it becomes very hard to oxidize, despite being geometrically close enough. In contrast, the chloro group allows better electronic communication between the heme and the substrate, reflected in higher spin density on the aromatic ring and methyl group and a more reactive alignment for the critical hydrogen-abstraction step.
What This Means for Cleaner Chemistry
In everyday terms, CYP153A6 behaves like a tiny, oily glove that holds certain fuel-like molecules in just the right way for oxygen to be added at a specific point—turning them into more water-soluble, useful, and ultimately more degradable products. The study shows that the enzyme prefers non-polar substituents and that both shape and electron flow determine whether a given toluene derivative will be transformed. By clarifying how CYP153A6 recognizes and activates these molecules, the work lays a blueprint for engineering improved versions that could help clean up aromatic pollutants or manufacture complex chemicals more sustainably.
Citation: Wei, Y., Donzella, S., Foiadelli, S. et al. Structure-based modeling reveals molecular basis for CYP153A6’s novel activity toward toluene derivatives. Sci Rep 16, 7570 (2026). https://doi.org/10.1038/s41598-026-38986-7
Keywords: biocatalysis, cytochrome P450, toluene hydroxylation, enzyme engineering, green chemistry