Clear Sky Science · en
Pseudo-healthy image synthesis via location-guided diffusion models for focal cortical dysplasia lesion localization
Why spotting tiny brain scars matters
For many people with epilepsy, seizures keep coming despite powerful medications. A common hidden culprit is a tiny patch of malformed brain tissue called focal cortical dysplasia. These patches can often be cured with surgery—if doctors can find them. But on standard brain scans, the lesions can be so subtle that even expert radiologists miss them. This study presents a new artificial intelligence (AI) technique that “imagines” what a person’s brain scan would look like if it were completely healthy, then uses the differences to reveal hard-to-see lesions, potentially opening the door to surgery for more patients.
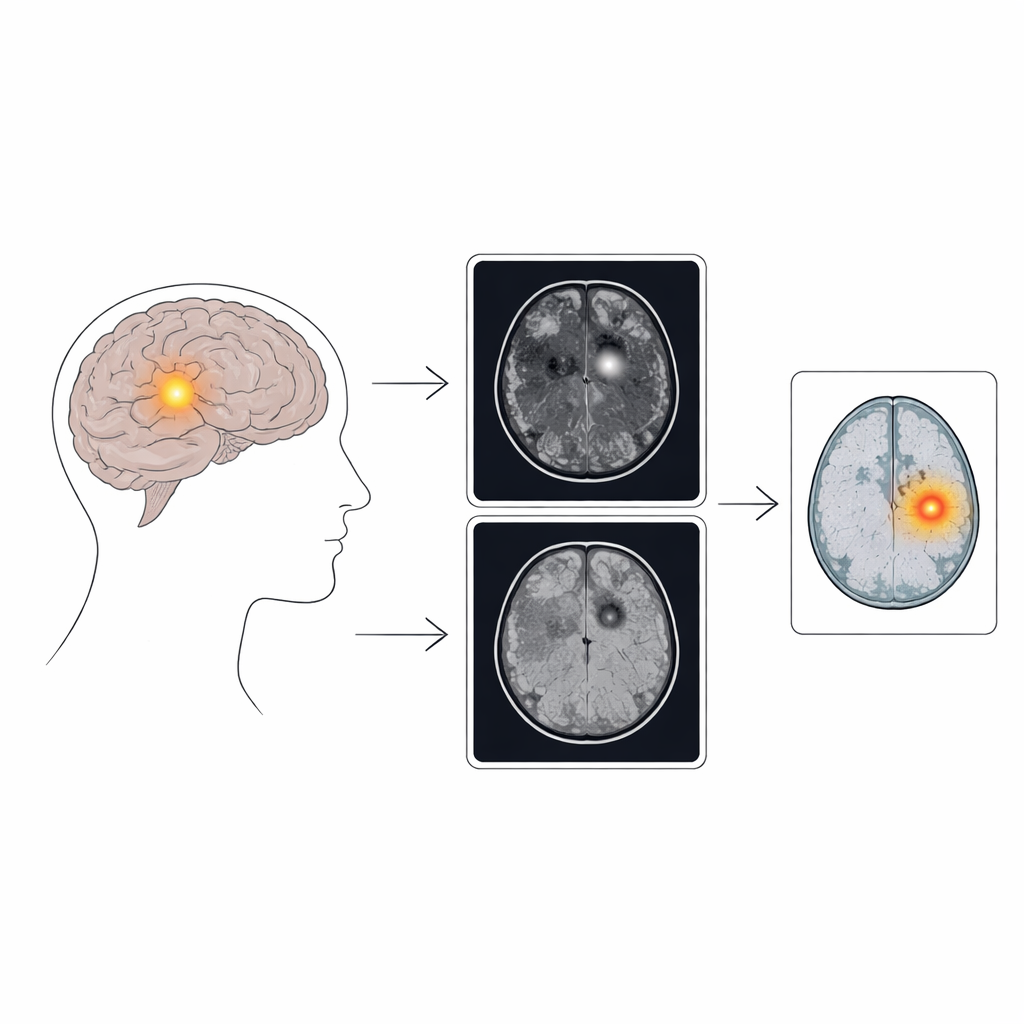
Tiny trouble spots in a big problem
Epilepsy affects more than 70 million people worldwide, and about a third continue to have seizures even after trying several drugs. In children, one of the main reasons is focal cortical dysplasia, where a small patch of brain develops abnormally. On MRI scans, these areas may show slightly thickened cortex or a blurred border between gray and white matter—changes that can be extremely easy to overlook. Because labeling these delicate abnormalities slice by slice is time-consuming and inconsistent across hospitals, there are few large, well-annotated datasets to train traditional supervised AI tools. The authors therefore focused on weakly supervised anomaly detection, an approach that learns patterns of normal tissue and flags anything that looks out of place, without needing detailed manual outlines of each lesion.
An AI that asks: what would a healthy brain look like?
The core idea of the method is to generate a “pseudo-healthy” version of a patient’s brain scan and then measure how it differs from the real image. To do this, the team built on diffusion models, a powerful class of image generators that gradually add noise to an image and then learn how to reverse that process. In this work, the model is trained to convert one MRI type (a T1-weighted image, which highlights anatomy) into another type (a FLAIR image, which is especially sensitive to certain lesion features). During the reverse process, the model is gently pushed to transform suspicious regions so they resemble healthy tissue, while leaving normal areas essentially unchanged. The difference between the original FLAIR scan and the generated, “cleaned” FLAIR scan becomes an anomaly map that highlights likely lesion locations.
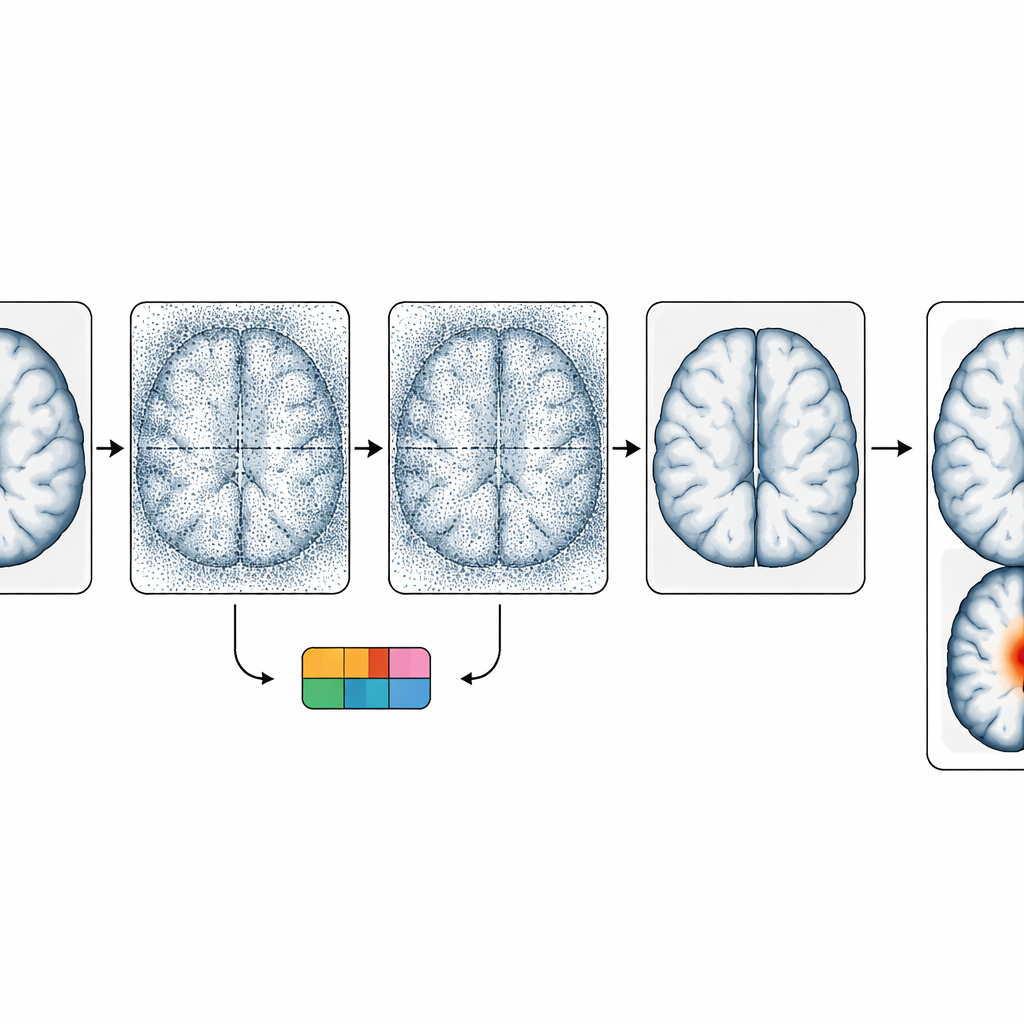
Using two types of scans and rough location hints
Different MRI sequences show focal cortical dysplasia in different ways. T1 images better reveal shape changes in the cortex, while FLAIR tends to make water-rich, abnormal tissue stand out with brighter signal and blurred boundaries. The authors exploit this complementarity by feeding the T1 image as a guide when generating the FLAIR image, encouraging the model to use structure from one modality and signal changes from the other. They also add a second source of guidance: a classifier trained to recognize which broad brain region (such as frontal or temporal lobe) contains the abnormality, or whether the scan is normal. This regional information steers the diffusion process to focus its “healing” efforts on areas where lesions are most likely, improving the chances of revealing small epileptic foci without heavily altering the whole brain.
Fixing color shifts and testing on real patients
Generative models can subtly change image brightness or contrast, which risks confusing doctors or hiding true abnormalities. To counter this, the researchers apply histogram matching, a standard image-processing technique that forces the generated FLAIR image to have the same overall intensity distribution as the original scan. This keeps the look and feel of the image familiar while preserving the lesion-related differences the model introduces locally. The method was tested on a public dataset from the University Hospital Bonn, containing MRI scans of 85 patients with focal cortical dysplasia type II and 85 healthy controls. After careful preprocessing and training, the new approach outperformed four competing anomaly-detection methods, achieving a high image-level recall (finding lesions in most affected scans) and better alignment with expert lesion maps at the pixel level.
What this could mean for people with epilepsy
The study shows that AI can be used not just to classify brain scans, but to generate realistic “what if healthy?” images that make hidden lesions stand out. Without requiring labor-intensive, voxel-by-voxel labels, the method combines multimodal MRI, rough location cues, and careful intensity correction to detect subtle epileptic scars more reliably than several existing tools. While it is not perfect—the differences between scan types can still introduce false alarms, and some lesions may remain too similar to normal tissue—the approach moves the field closer to dependable, automated support for radiologists. In the long run, such techniques could help identify surgical targets earlier and more consistently, improving outcomes for people living with drug-resistant epilepsy.
Citation: Li, Y., Pan, Y., Zhang, X. et al. Pseudo-healthy image synthesis via location-guided diffusion models for focal cortical dysplasia lesion localization. Sci Rep 16, 8101 (2026). https://doi.org/10.1038/s41598-026-38981-y
Keywords: epilepsy, brain MRI, focal cortical dysplasia, medical imaging AI, anomaly detection