Clear Sky Science · en
Fine-tuned large language models with structured prompts enable efficient construction of lung cancer knowledge graphs
Why turning medical text into maps matters
Lung cancer is one of the world’s deadliest cancers, and information about how to diagnose and treat it is scattered across research papers, hospital notes, online consultations, and traditional medicine casebooks. Doctors and researchers struggle to keep up with this flood of text. This study explores a new way to automatically turn that scattered knowledge into a single, navigable "map"—a lung cancer knowledge graph—using a fine-tuned large language model and carefully structured prompts. The result aims to make complex medical knowledge easier for computers to search and for experts to use in decision support tools.
From scattered stories to connected facts
The authors focus on a simple idea: if you can reliably pull out who-does-what-to-what from medical text, you can stitch those facts into a graph. In practice, this means converting free-form sentences into tiny building blocks called triples—pairs of entities linked by a relationship, such as “lung cancer – treated by – chemotherapy.” Traditional ways of building such graphs either require armies of annotators or brittle rules that miss nuance and new discoveries. To overcome this, the team fine-tunes an existing Chinese large language model, ChatGLM-6B, so that it specializes in spotting medically meaningful triples about lung cancer in a wide range of sources, from online patient–doctor chats to structured databases and traditional Chinese medicine records. 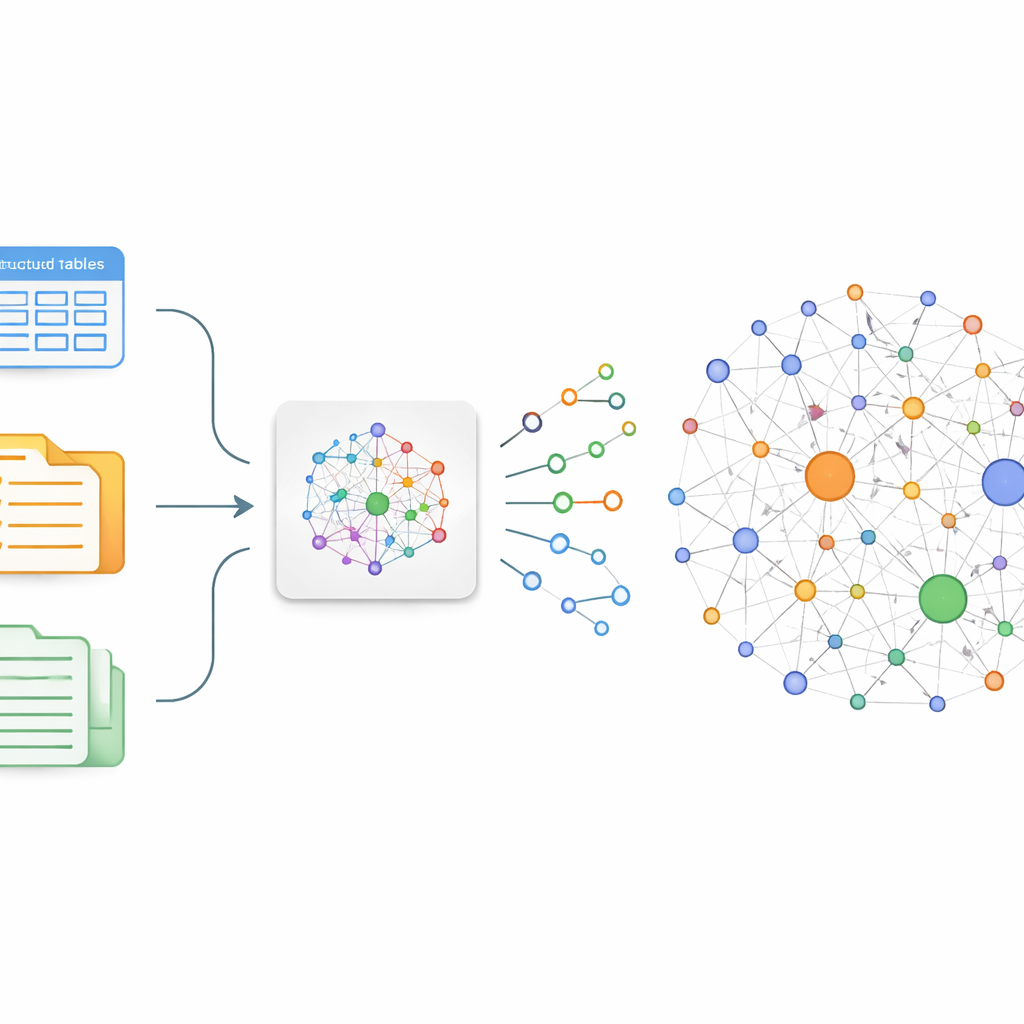
Teaching an AI to think in neat units
Simply asking a general-purpose language model to “extract information” often yields messy, chatty answers. The researchers therefore design a strict prompting scheme and then fine-tune the model on nearly 50,000 examples of good behavior. Each example shows an instruction and the exact triple-style output expected. The prompt tells the model to act like a professional text-mining expert, to produce only structured triples in a computer-readable format, and to “think step by step” when sentences contain nested details—for example, a treatment, the drug used, and its dosage. This combination of role-setting, format rules, and stepwise reasoning turns the model—now called KGLM—from a conversational assistant into a disciplined extractor of machine-ready facts.
Blending many voices into one clear graph
Raw triples from text are only part of the story. The same disease or drug often appears under different names—"chronic obstructive pulmonary disease" versus "COPD," for instance. To avoid clutter and confusion, the authors design a fusion stage that merges equivalent entities across three data streams: unstructured web text, semi-structured clinical cases, and existing medical knowledge graphs. First, a fast string-based similarity check flags obvious matches. When that is not enough, a deeper semantic similarity model (Sentence-BERT) compares meanings in context. Entities judged to be duplicates are collapsed into a single canonical node, with shorter names preferred and other forms stored as aliases. Experts then review edge cases and remove misleading or low-quality statements, yielding a cleaner and more coherent lung cancer knowledge graph stored in a Neo4j database. 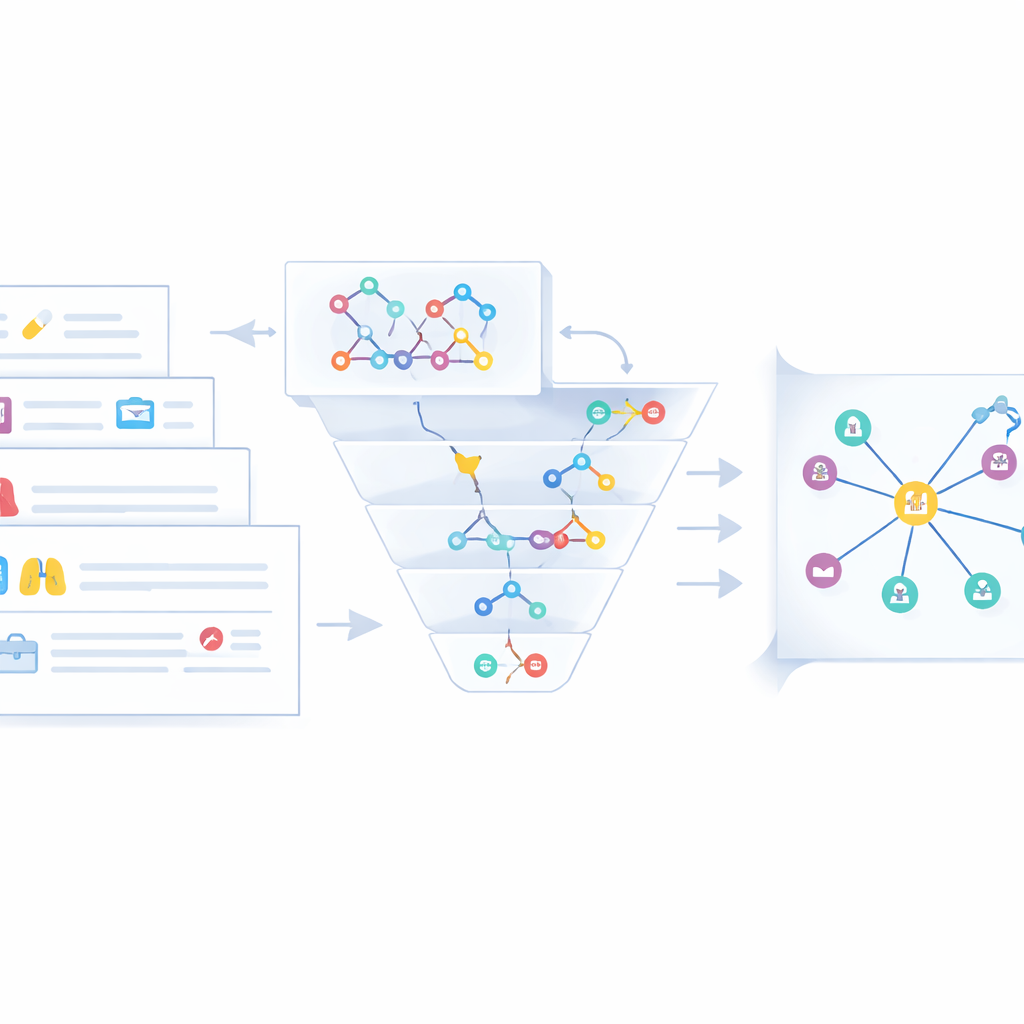
How well does this knowledge map work?
To gauge performance, the team compares KGLM to standard deep learning approaches based on BERT and convolutional networks, as well as to the original, unfine-tuned ChatGLM model. On the task of relation extraction—deciding which entities are linked and how—the fine-tuned, prompt-guided KGLM achieves an F1 score of about 0.82, outperforming all tested baselines and improving roughly 25 percent over the starting model. Ablation tests show that each prompt component matters: removing the expert role, the strict triple format, or the “think step by step” guidance all degrades accuracy, especially for complex sentences with nested attributes or traditional Chinese medicine terminology. A panel of clinical and informatics experts also judges the resulting graph to be more accurate, usable, and clinically relevant than graphs built without fine-tuning or structured prompts.
What this means for future medical tools
In plain terms, the study shows that with the right training and instructions, a large language model can efficiently turn messy, real-world lung cancer text into a structured, searchable web of facts. This lung cancer knowledge graph, while still a research prototype and limited to Chinese-language sources and a single disease area, points toward a future where constantly updated “knowledge maps” could support decision-support systems, education tools, and research exploration. The authors stress that such graphs must be carefully validated and regularly updated, and are not ready to guide care without expert oversight. Still, their results suggest that fine-tuned language models plus smart prompting can make the daunting task of organizing medical knowledge more scalable and timely.
Citation: Zhou, C., Gong, Q., Luan, H. et al. Fine-tuned large language models with structured prompts enable efficient construction of lung cancer knowledge graphs. Sci Rep 16, 9505 (2026). https://doi.org/10.1038/s41598-026-38959-w
Keywords: lung cancer, knowledge graph, large language model, relation extraction, medical AI