Clear Sky Science · en
Evaluating the evolutionary relationship of TATA binding protein (TBP) with various folding patterns of protein domains using support vector machine (SVM)
How a DNA "on switch" protein connects to many others
The TATA‑box binding protein, or TBP, is a workhorse of our cells: it helps switch genes on by gripping DNA at many promoters. This study asks a deceptively simple question with big implications: are there other proteins, with very different jobs, that quietly share TBP’s underlying shape? By combining 3D structure comparison, sequence analysis, and modern machine‑learning tools, the authors trace hidden family ties between TBP and proteins involved in metabolism, neurotransmitter chemistry, and even cancer‑related pathways.
One key protein at the heart of gene control
TBP sits at the gateway of gene expression in organisms ranging from yeast to humans. It recognizes a short DNA sequence called the TATA box and bends the DNA to help assemble the large transcription machinery that copies genes into RNA. Because this step is so central, the fold—the three‑dimensional arrangement—of TBP’s core is highly conserved across evolution. The authors focus on a well‑studied TBP structure known as 1tba and use it as a probe to search for other proteins that may share its architectural blueprint, even if their amino‑acid sequences and everyday tasks look very different at first glance.
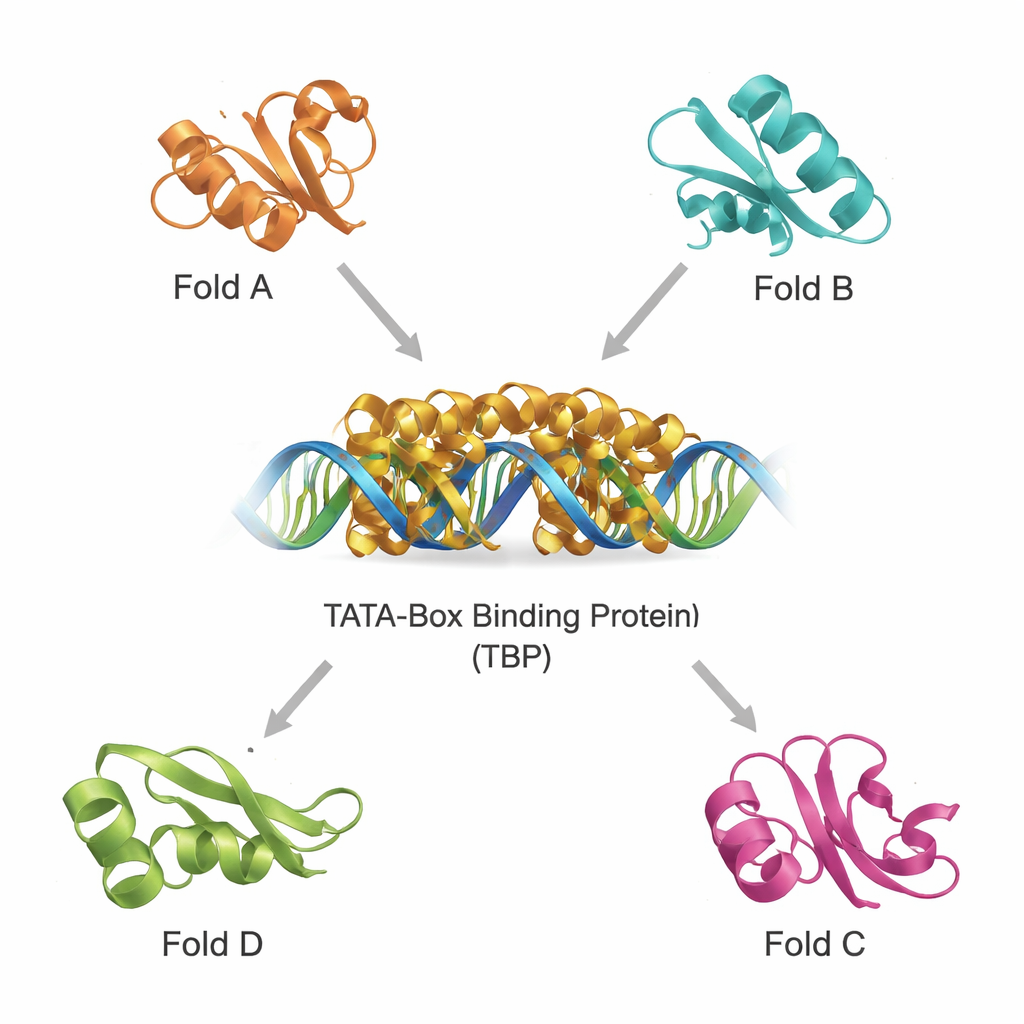
Finding structural cousins in a crowded protein universe
Modern databases contain hundreds of thousands of protein structures, making it possible to scan for distant relatives by 3D shape rather than by sequence alone. Using two powerful tools, DALI and TOP‑search, the team first pulled out proteins whose folds looked like TBP’s. They then classified these candidates with an evolutionary domain catalogue and narrowed them to a small set of structurally similar but functionally diverse examples. These include a glutamine‑making enzyme important in metabolism, a domain found in several tRNA‑handling enzymes, an enzyme with a distinctive “hot‑dog” fold involved in fatty‑acid chemistry, and proteins that help make tetrahydrobiopterin, a molecule crucial for brain function. Superimposing their structures on TBP showed that, despite different jobs, they share recognizable core motifs.
Teaching machines to recognize hidden protein families
To move beyond case‑by‑case inspection, the authors built machine‑learning models that could automatically flag TBP‑like folds. They assembled large sets of protein sequences known to belong to TBP or to each of the related folds, along with a broad “background” set of unrelated proteins. Each protein was converted into simple numerical summaries: how often each amino acid appears, and how frequently each possible pair of amino acids occurs in sequence. These profiles fed support vector machines (SVMs) and random‑forest models, which learned to separate one fold type from all others. Using rigorous cross‑validation, the models reached very high accuracy—often above 95 percent—even when trained on only parts of the sequences corresponding to conserved regions.
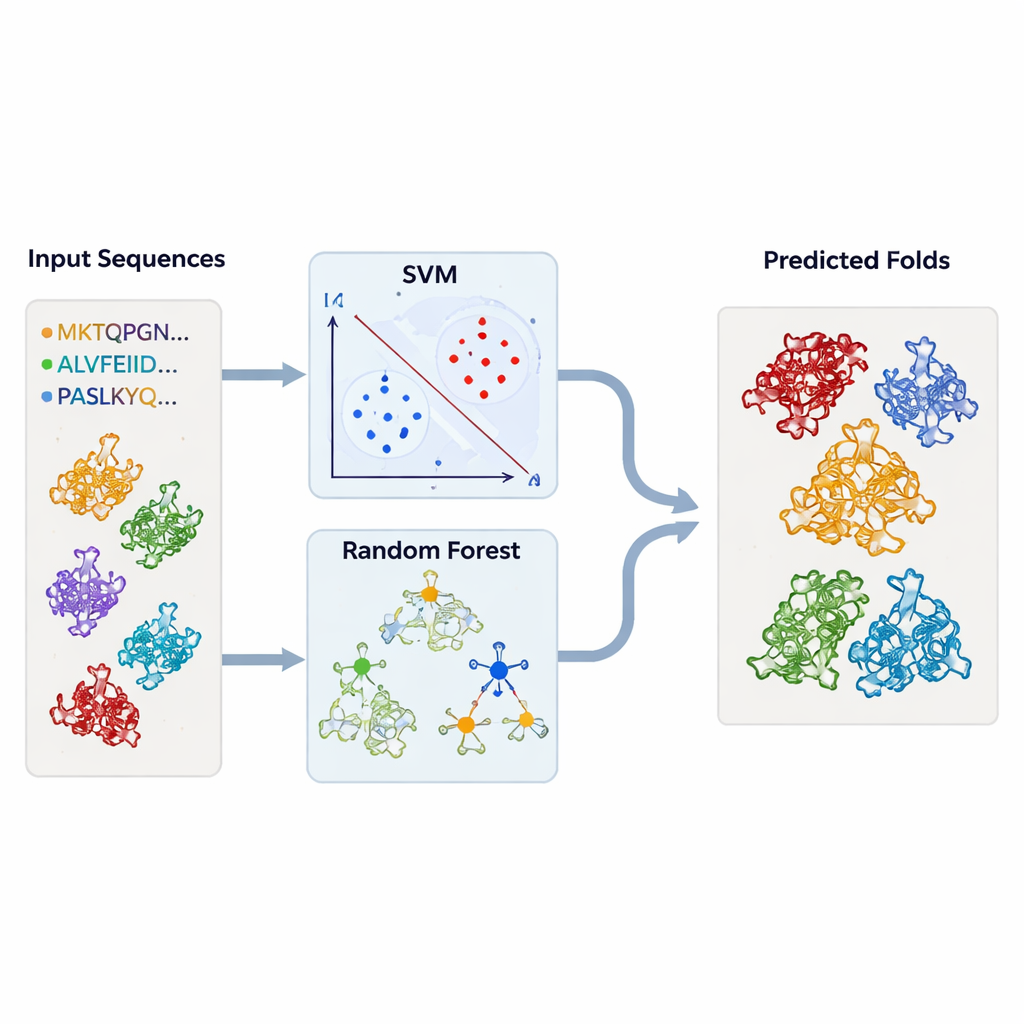
Testing the models on thousands of unknown structures
Armed with these trained classifiers, the team returned to the structural databases. They ran thousands of protein chains—retrieved from DALI and TOP‑search—through their models to see which ones bore the statistical hallmarks of TBP‑like or related folds. The SVM and random‑forest approaches largely agreed and picked out many candidates that structural tools had also flagged as similar. In some cases, enzymes with apparently unrelated activities nonetheless clustered strongly with TBP or with one another, reinforcing the idea that evolution can repurpose the same underlying framework for many different biochemical roles.
Why these hidden connections matter
The study concludes that TBP shares deep structural ancestry with several enzyme families, including glutamine synthetase‑like proteins and editing domains of tRNA‑processing enzymes. Even when sequences have drifted and functions have diverged, these proteins retain common architectural motifs, suggesting descent from a shared ancestor. For a non‑specialist, the key message is that nature tends to recycle successful designs: one fold can be adapted repeatedly to solve very different problems, from turning genes on to fine‑tuning metabolism and brain chemistry. By combining 3D structure comparison with machine learning, the authors provide a practical toolkit to uncover such relationships, helping biologists predict what uncharacterized proteins might do and pointing drug developers toward new, evolution‑guided targets in disease‑relevant pathways.
Citation: Selvaraj, M.K., Kaur, J. Evaluating the evolutionary relationship of TATA binding protein (TBP) with various folding patterns of protein domains using support vector machine (SVM). Sci Rep 16, 7696 (2026). https://doi.org/10.1038/s41598-026-38883-z
Keywords: TATA-box binding protein, protein evolution, machine learning, protein structure, support vector machine