Clear Sky Science · en
Novel convolutional neural network for bacterial identification of confocal microscopic datasets
Why spotting germs faster matters
When doctors try to figure out which bacteria are causing an infection, time is critical. Traditional lab tests can take many hours or even days, and they require highly trained experts to examine microscope images by eye. This study introduces a new computer vision system, called CM-Net, that can automatically read specialized microscope images and quickly tell two common, medically important bacteria apart, while also recognizing which cells are alive or dead. The work suggests a path toward faster, more reliable diagnostics that could one day be used in hospitals and research labs worldwide.
Turning glowing germs into useful pictures
The researchers started with a powerful imaging tool known as confocal laser scanning microscopy. In simple terms, this microscope uses a focused laser and fluorescent dyes to make bacteria glow in different colors depending on whether they are alive or dead. Live cells show up as green, while dead cells appear red. By scanning through the sample in very thin layers, the microscope builds crisp, detailed pictures of the bacteria on glass slides. The team worked with two well-known species that often cause hospital infections: rod-shaped Escherichia coli and round Staphylococcus aureus. These high-quality images form the raw material that CM-Net must learn to understand.
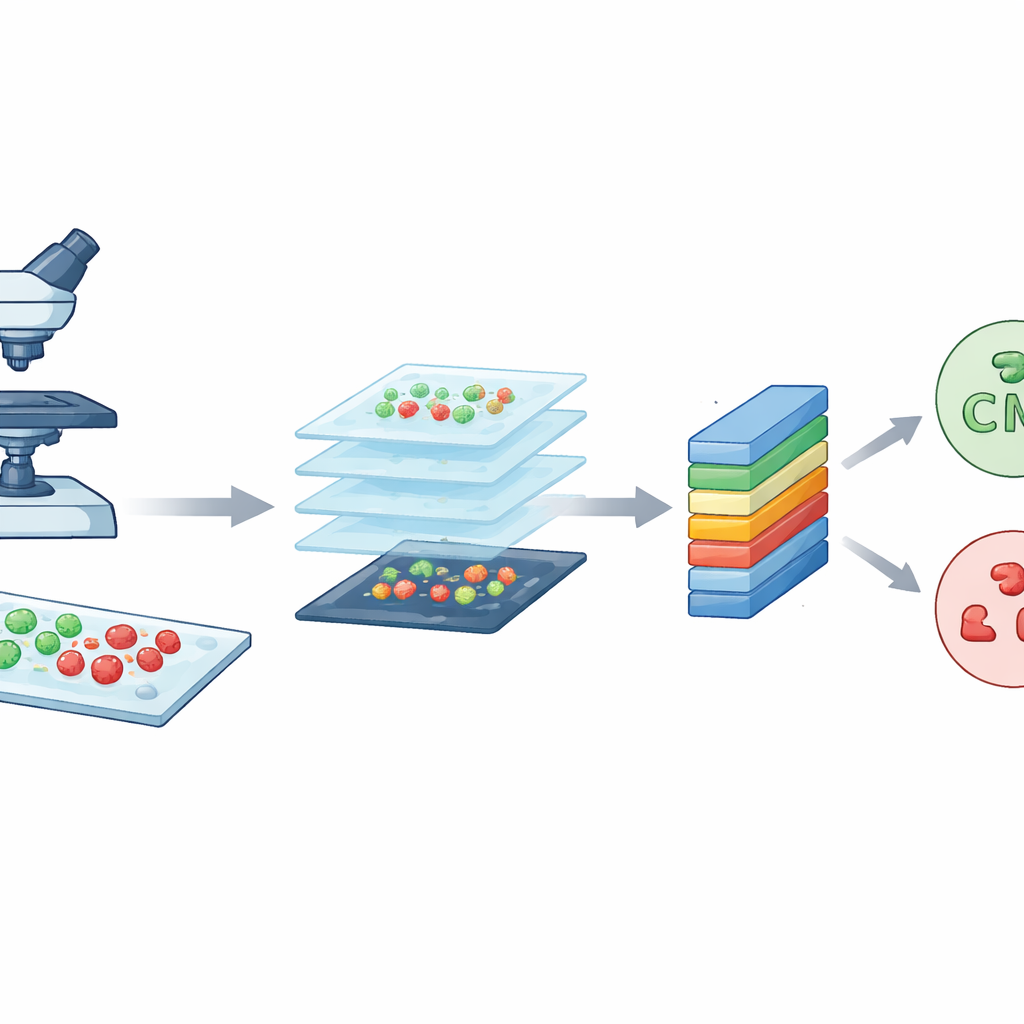
Making many small tiles from big images
Although each confocal image is rich in detail, it is also very large, about 3000 by 3000 pixels. Training a computer model directly on such huge images would be slow and demand excessive computing power. To solve this, the team sliced each large image into many smaller square tiles, each 224 by 224 pixels, a standard size in image analysis. This process, called data augmentation, both reduces the technical burden and multiplies the number of training examples. From an original set of 300 images per bacterial type, they generated 7,066 tiles in total. These tiles capture local patterns of shapes, colors, and textures from different regions of the slides, giving the model a diverse and balanced set of examples to learn from.
How the digital observer learns to see
CM-Net is a carefully designed deep-learning model built specifically for bacterial microscopy, rather than adapted from general photo collections. It is a kind of convolutional neural network, a class of programs that excel at finding patterns in images. CM-Net processes each tile through a series of stages. Early stages look for simple visual cues like edges and spots; deeper stages combine these into more complex patterns that distinguish rods from spheres and live from dead cells. The network uses techniques such as batch normalization, which keeps its internal signals well behaved, and a clipped form of its activation function, which prevents extreme responses that can destabilize learning. Later layers condense the extracted information and make a final decision about the bacterial type and cell state.
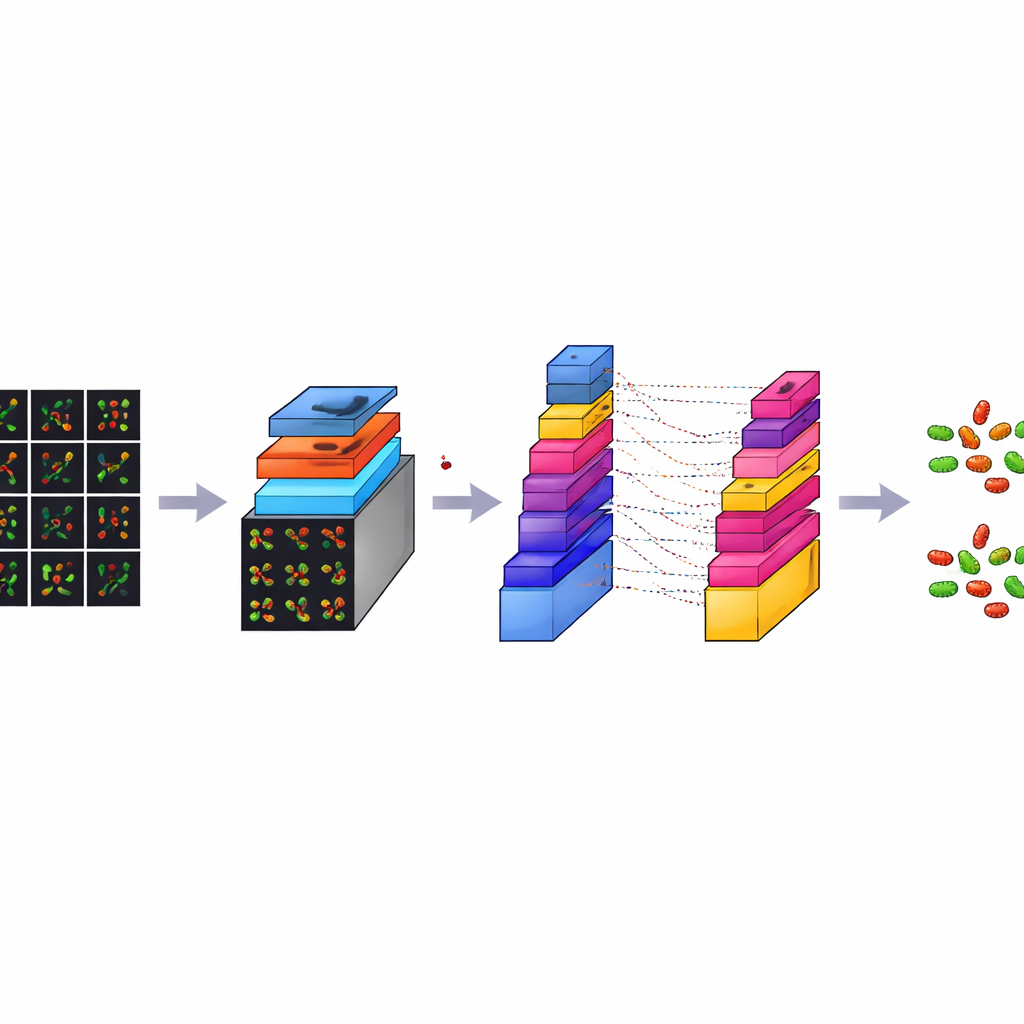
Outperforming popular off-the-shelf models
To see how well CM-Net performs, the authors trained and tested it 30 times, each time using a fresh split of the data into training and testing groups. They measured accuracy, how often the model was right overall; sensitivity, how well it detected each target; specificity, how well it avoided false alarms; and several other standard scores. CM-Net achieved about 96% accuracy on average, with sensitivity and specificity also around 96%, and a strong balance between the two classes. It also needed fewer internal parameters and less memory than several widely used pre-trained models, including GoogLeNet, MobileNetV2, ResNet18, and ShuffleNet, while still being faster to run. Visualization tools showed that CM-Net focuses its attention on the actual bacterial bodies in the images, rather than on random background features, supporting the idea that it is learning biologically meaningful cues.
What this means for future lab work
In everyday terms, the study shows that a purpose-built deep-learning system can learn to “read” complex microscope images of bacteria accurately, efficiently, and in a way that aligns with what human experts care about. For now, CM-Net has been trained on only two bacterial species and on data from a single type of microscope, so more work is needed before it can be used as a general diagnostic tool. The authors plan to extend it to more species, different cell states, and larger, more varied datasets. Still, the results suggest that systems like CM-Net could eventually help laboratories identify infections more quickly, guide treatment decisions, and open up automated analysis of microbiology experiments to users without specialized imaging expertise.
Citation: Al-Jumaili, A., Al-Jumaili, S., Alyassri, S. et al. Novel convolutional neural network for bacterial identification of confocal microscopic datasets. Sci Rep 16, 8123 (2026). https://doi.org/10.1038/s41598-026-38861-5
Keywords: bacterial image classification, confocal microscopy, deep learning, convolutional neural networks, medical diagnostics