Clear Sky Science · en
SPCNNet: spiking point cloud neural network for morphological neuron classification
Why the shape of brain cells matters
Every thought, memory, and sensation you experience depends on the work of billions of neurons—electrically active cells with intricate tree‑like branches. These branches are not all shaped alike, and those differences are closely tied to what each neuron does in the brain. The paper described here introduces a new way to classify neurons by their 3D shapes using a brain‑inspired form of artificial intelligence, potentially improving how we map and understand neural circuits.
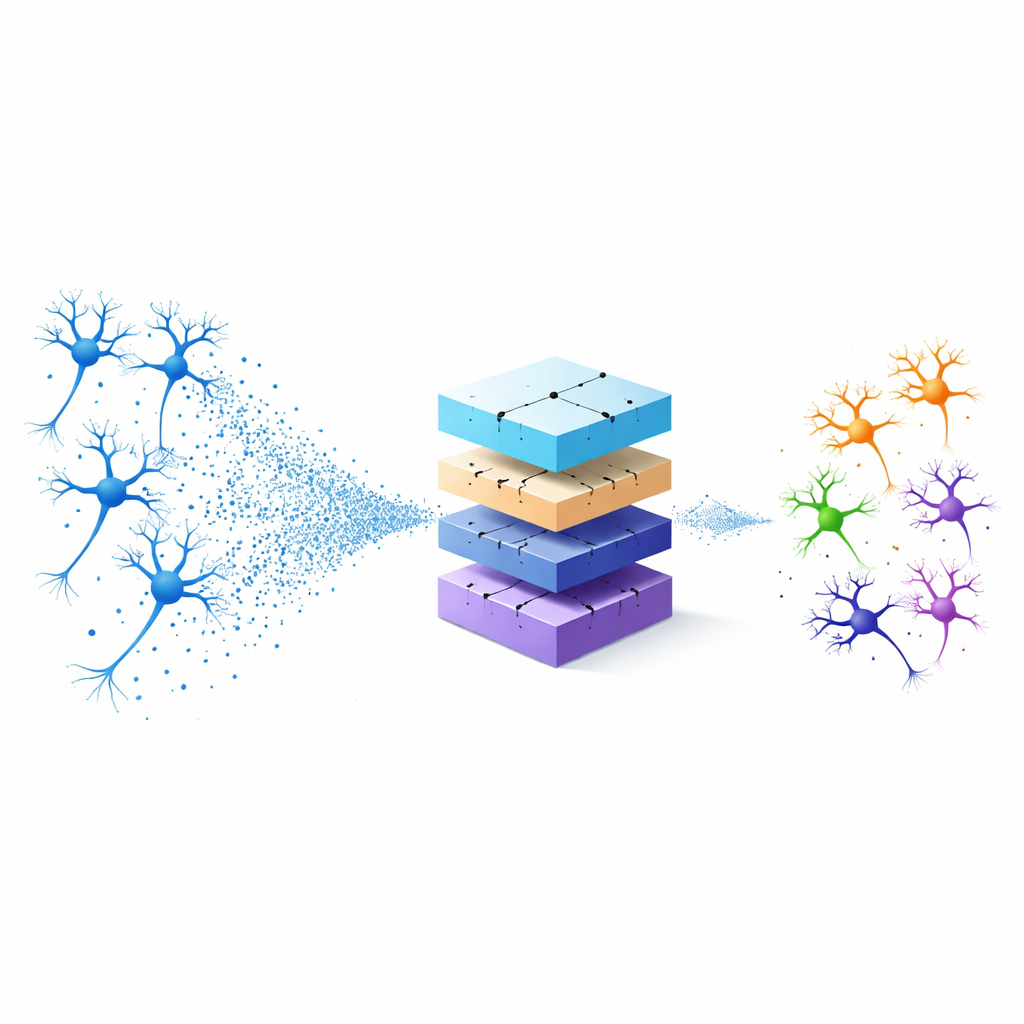
Seeing neurons as clouds of points
Traditionally, scientists have classified neurons either by hand‑crafted geometric measurements—such as how many branches they have—or by flattening 3D cells into 2D images for standard image‑recognition software. Both strategies throw away information: fixed measurements may miss subtle shape patterns, and 2D projections lose depth. The authors instead treat each neuron as a 3D “point cloud,” a set of points in space that traces its overall form. They start from a standard digital description of neurons known as SWC files and keep only the 3D coordinates and connections of each tiny segment. Using a technique called farthest point sampling, they pick a subset of points that still captures the overall structure but greatly reduces the amount of data that needs to be processed.
Letting spikes do the thinking
Most artificial neural networks use smooth, continuous signals very different from the brief electrical spikes that real neurons send to one another. In contrast, the model proposed here—called the Spiking Point Cloud Neural Network, or SPCNNet—uses artificial neurons that communicate with discrete spikes over time. After the 3D point cloud of each biological neuron is constructed and normalized, the coordinates are passed through a calibration stage that aligns them in space so that the system is not confused by rotations or ordering of the points. These aligned values are then converted into spike trains using a simplified model of electrical activity, turning spatial information about the neuron’s shape into patterns of spikes that unfold over a short simulated time window.
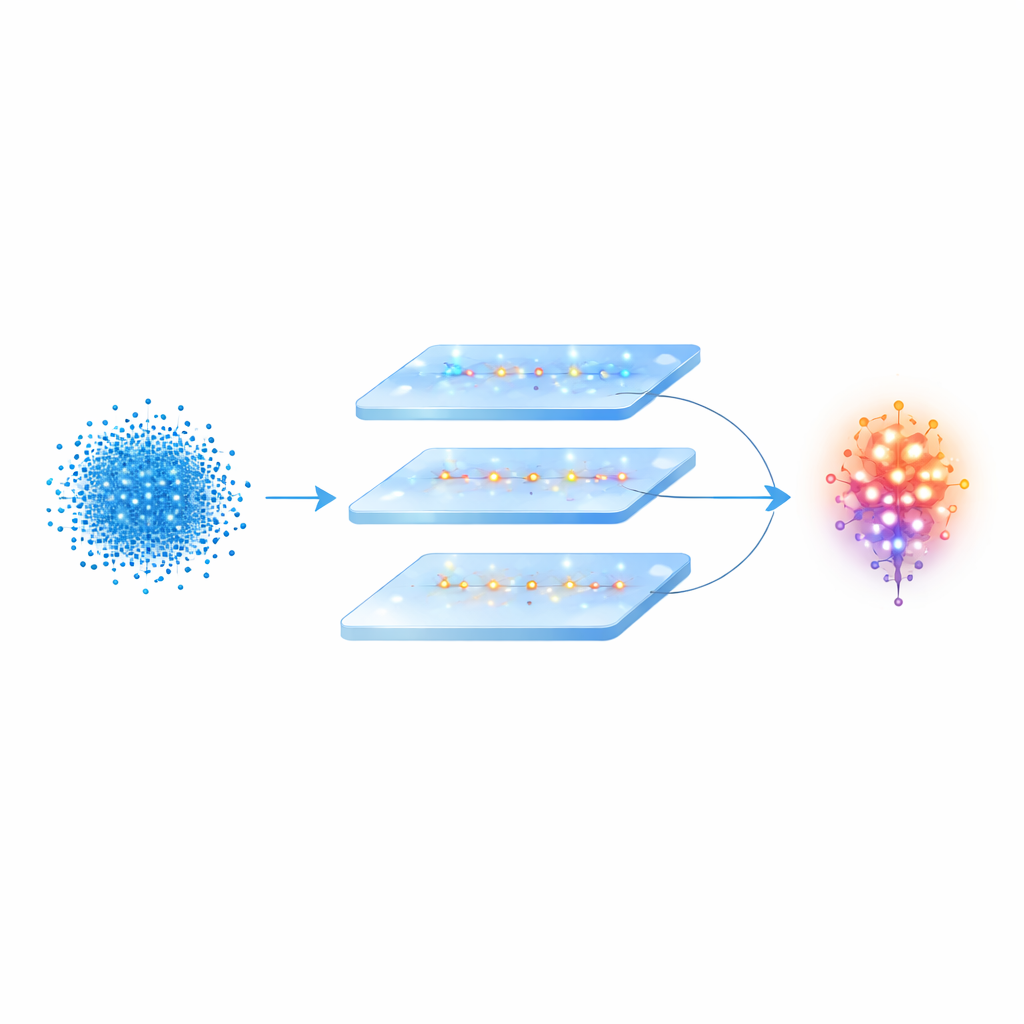
Teaching the network to recognize cell types
Once the neuron shapes have been encoded as spike trains, SPCNNet applies a series of operations to extract informative features. Convolution‑like layers examine all sampled points and gradually build up higher‑dimensional representations of the neuron’s overall form, while a pooling step compresses this information into a compact summary. Fully connected layers then map this summary to a small number of possible neuron types, and a final decision stage outputs the most likely class. The authors trained and tested their model on two carefully constructed datasets drawn from the public NeuroMorpho database: one of three types of neurons in the tiny worm C. elegans, and another of four neuron types in the olfactory bulb of zebrafish, as well as on a larger and more imbalanced collection called NeuMorph.
How well the new approach performs
Across these datasets, SPCNNet proved both accurate and efficient. On the worm neurons, it reached test accuracies of about 85 percent, rivaling or slightly trailing the best traditional deep‑learning methods that rely on hand‑engineered geometric features. On the more challenging zebrafish neurons—larger cells with thousands of segments—SPCNNet clearly outperformed competing approaches, again achieving around 85 percent test accuracy while many 3D image‑based or point‑cloud methods lagged well behind. Careful experiments showed how performance depended on key design choices such as how many points were sampled from each neuron, how long the spike simulation ran, and how many examples were processed at once. Additional ablation tests demonstrated that both the farthest point sampling and the spiking neuron units were crucial to the model’s success.
What this means for brain research
By treating each neuron as a 3D point cloud and processing it with spike‑based computation, SPCNNet offers a way to classify neurons that is closer in spirit to how the brain itself handles information. The method avoids the need for hand‑designed measurements or 2D projections and instead learns directly from the full 3D structure, while also promising lower energy consumption thanks to its sparse spiking activity. Although the current version uses only position and connectivity and leaves out other details such as branch thickness or cell type tags, it already matches or surpasses many established techniques and scales well to larger, imbalanced datasets. With further refinement, this approach could become a powerful tool for automatically cataloging the diverse shapes of neurons, helping neuroscientists build richer maps of the brain’s cellular landscape.
Citation: Lin, X., Yu, M. & Wang, X. SPCNNet: spiking point cloud neural network for morphological neuron classification. Sci Rep 16, 7989 (2026). https://doi.org/10.1038/s41598-026-38839-3
Keywords: neuron morphology, spiking neural networks, 3D point clouds, cell type classification, computational neuroscience