Clear Sky Science · en
Gene-specific marker and trait-based evaluation of powdery mildew resistance in garden pea (Pisum sativum var Hortense L.)
Why Protecting Peas Matters
Garden peas are more than a side dish: they are a nutritious, protein-rich crop that supports both human diets and farm animals around the world. But a common fungal disease called powdery mildew can coat pea plants in a white, talc-like layer, shriveling leaves, spoiling pods, and cutting yields by half. This study set out to find pea lines that can naturally fend off this disease and to pinpoint the underlying resistance genes, so breeders can develop hardier pea varieties for farmers without relying heavily on fungicides.
When a Quiet Fungus Becomes a Big Farm Problem
Powdery mildew thrives in warm days and cool nights, especially when humidity is high—conditions often found in important pea-growing regions. The fungus lives on living plant tissue and can even reach the seeds, lowering both yield and quality. While chemical sprays can suppress outbreaks, they are costly, need repeated applications, and raise environmental concerns. A more sustainable path is to grow pea varieties that carry built-in resistance. Earlier work had identified three major resistance genes in peas, known as er1, er2, and Er3. The present study asked a practical question: among 11 garden pea lines maintained at an Indian research institute in the Himalayan foothills, which ones truly resist powdery mildew in real fields and under controlled lab conditions—and which resistance genes do they carry?
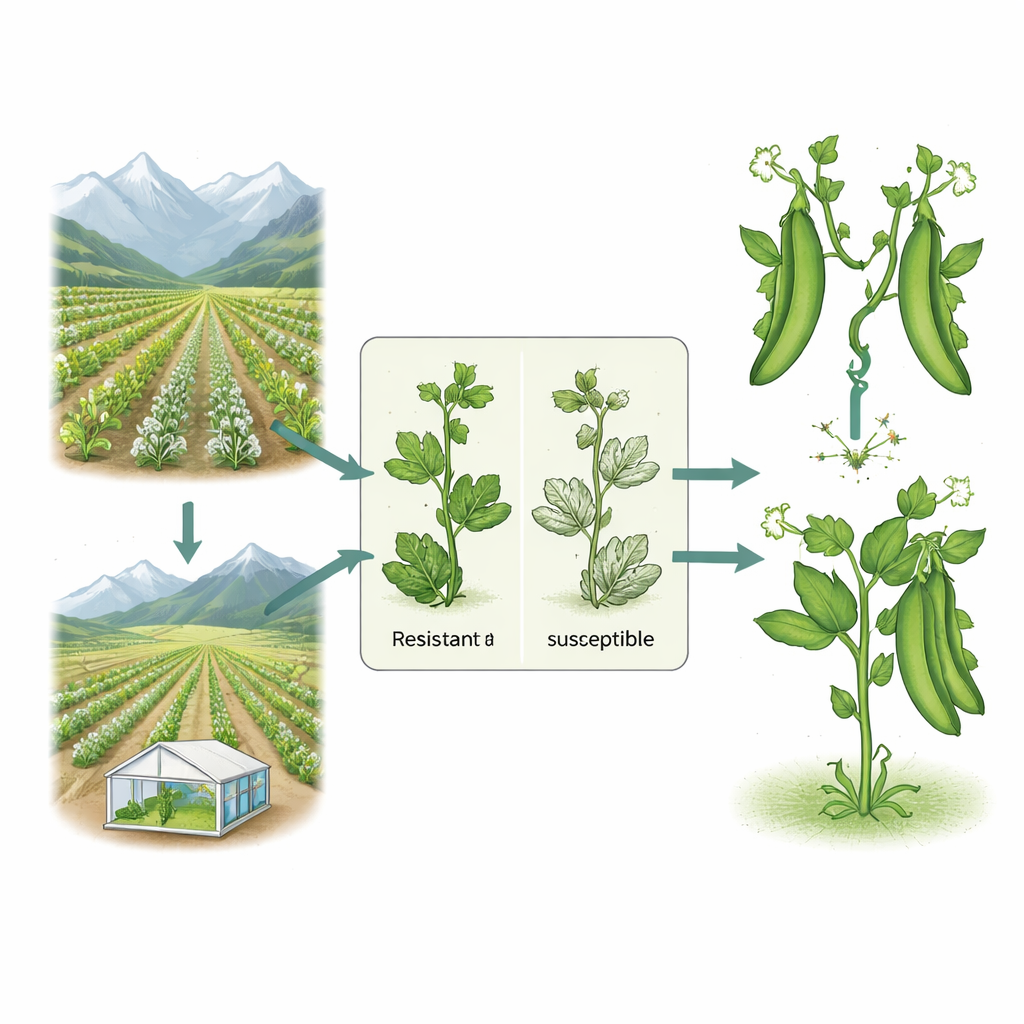
Putting Pea Lines to the Test in Fields and Lab
Researchers planted the 11 pea genotypes at two high-altitude test sites in Uttarakhand, India—Hawalbagh and Mukteshwar—during the winter growing season. They monitored the plants at two stages, when pods were developing and at the first picking, scoring how much of each plant was covered by the white fungal growth. To avoid “escapes” where plants simply miss exposure, they added extra fungus from a known susceptible variety, Arkel, which also served as a benchmark for severe disease. In the cooler, drier Hawalbagh site, disease appeared late and stayed relatively mild. At Mukteshwar, where temperatures and humidity better suited the fungus, nearly all lines eventually became infected. Two entries, VP-2020-101 and VP-2024-55, were standouts: they showed the lowest disease levels and were classed as resistant, while most others suffered moderate to high damage.
Zooming In on Leaves and DNA
Field results can be influenced by shifting weather, so the team also used a detached-leaf test to check how the fungus behaved on pea leaves in controlled settings. Leaflets from each genotype were floated on a nutrient solution in dishes and dusted with spores, then kept either in an incubator or a spore-proof chamber in a polyhouse. Under the microscope, resistant leaves showed only sparse fungal threads and few spores, while susceptible ones were blanketed by dense growth. Again, VP-2020-101 and VP-2024-55 consistently showed strong resistance across both controlled environments, closely matching what had been seen in the field. To understand why, the scientists examined the plants’ DNA with a set of gene-specific markers designed to light up when er1, er2, or Er3 are present. These markers act like genetic signposts, revealing which resistance genes are built into each line.
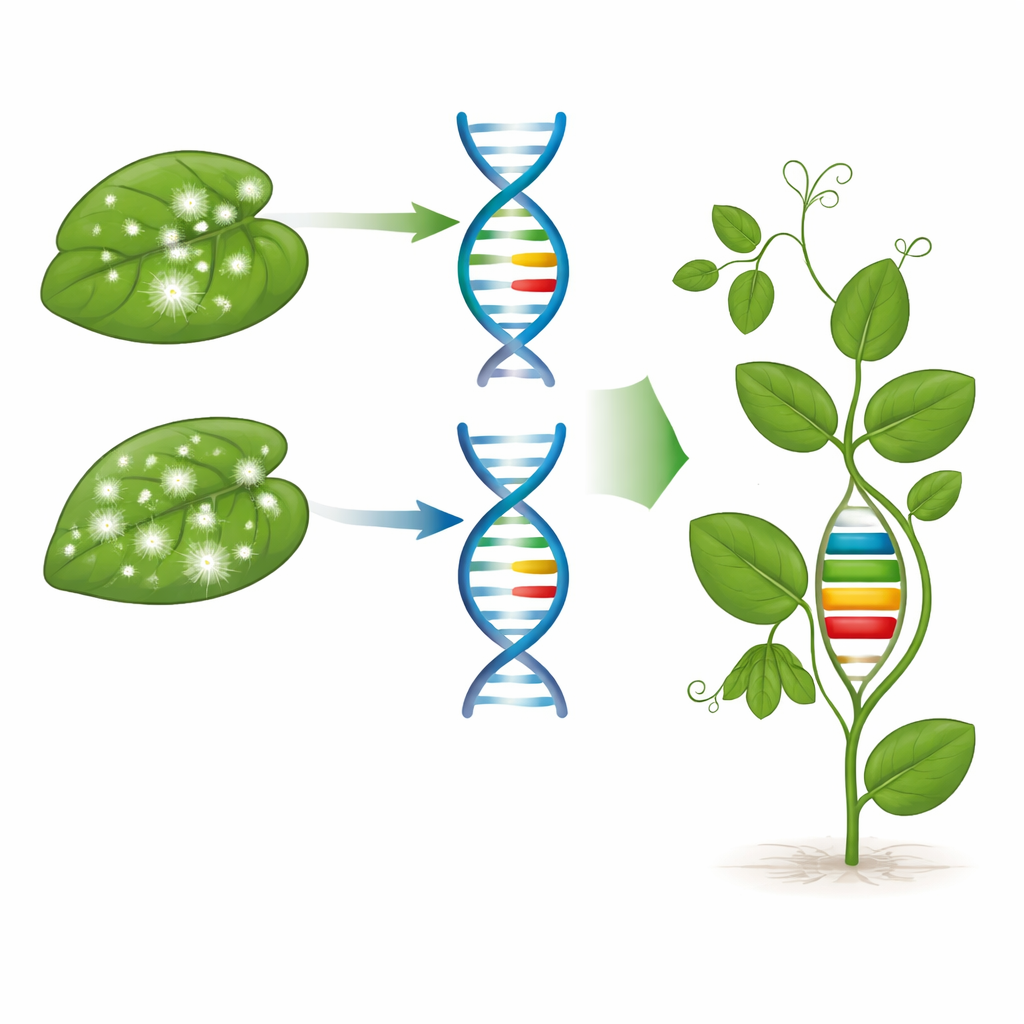
Building a Stronger Shield Inside the Plant
The DNA tests showed a clear pattern. VP-2024-55 carried a single key resistance gene, er1, which is known to block the fungus from successfully entering leaf cells and is often associated with solid, long-lasting protection. VP-2020-101, however, carried all three genes—er1, er2, and Er3—stacked together in one genetic package. This “pyramiding” of multiple resistance genes makes it harder for the fungus to evolve around the plant’s defenses, much like using several locks on a door. The molecular evidence lined up neatly with the field and leaf tests: the more complete the genetic shield, the more stable and robust the resistance under different environments.
What This Means for Future Pea Crops
For farmers and breeders, the study’s message is straightforward. Two pea lines, VP-2020-101 and VP-2024-55, offer valuable natural protection against powdery mildew, with VP-2020-101 providing the strongest and most durable resistance thanks to its three-gene shield. These lines can now serve as parents in breeding programs that aim to deliver new garden pea varieties needing fewer chemical sprays while maintaining high yields and quality. By combining careful field testing, controlled lab assays, and precise DNA tools, the researchers provide a roadmap for developing disease-resistant crops that are both productive and environmentally friendly.
Citation: Hedau, N.K., Santhiya, S., Mishra, K.K. et al. Gene-specific marker and trait-based evaluation of powdery mildew resistance in garden pea (Pisum sativum var Hortense L.). Sci Rep 16, 8784 (2026). https://doi.org/10.1038/s41598-026-38836-6
Keywords: powdery mildew, garden pea, disease resistance, plant breeding, fungal pathogens