Clear Sky Science · en
ImageNet pre-training and two-step transfer learning in chromosome image classification
Sharper views of our chromosomes
Our chromosomes carry the instructions for building and running our bodies, and doctors study their shapes to spot genetic disorders and some cancers. Today, computers can help read chromosome images, but teaching them to do this well is hard because medical images are scarce and look very different from everyday photos. This study asks a simple question with big practical impact: can computers learn better from related medical images, not just from huge collections of cat, dog, and car photos?
Why chromosome pictures matter
In hospitals, specialists arrange a person’s 46 chromosomes into a chart called a karyotype, grouped into 24 types (22 numbered pairs plus X and Y). Subtle dark and light bands along each chromosome help reveal missing or extra pieces linked to conditions such as Down syndrome or certain leukemias. Traditionally, experts classify these bands by eye, which is slow and subjective. Deep learning offers a way to automate this work, but these systems usually start from models trained on ImageNet, a massive dataset of everyday pictures. That jump—from vacation snapshots to microscope views of chromosomes—is huge, and it is not clear how well that experience really transfers.
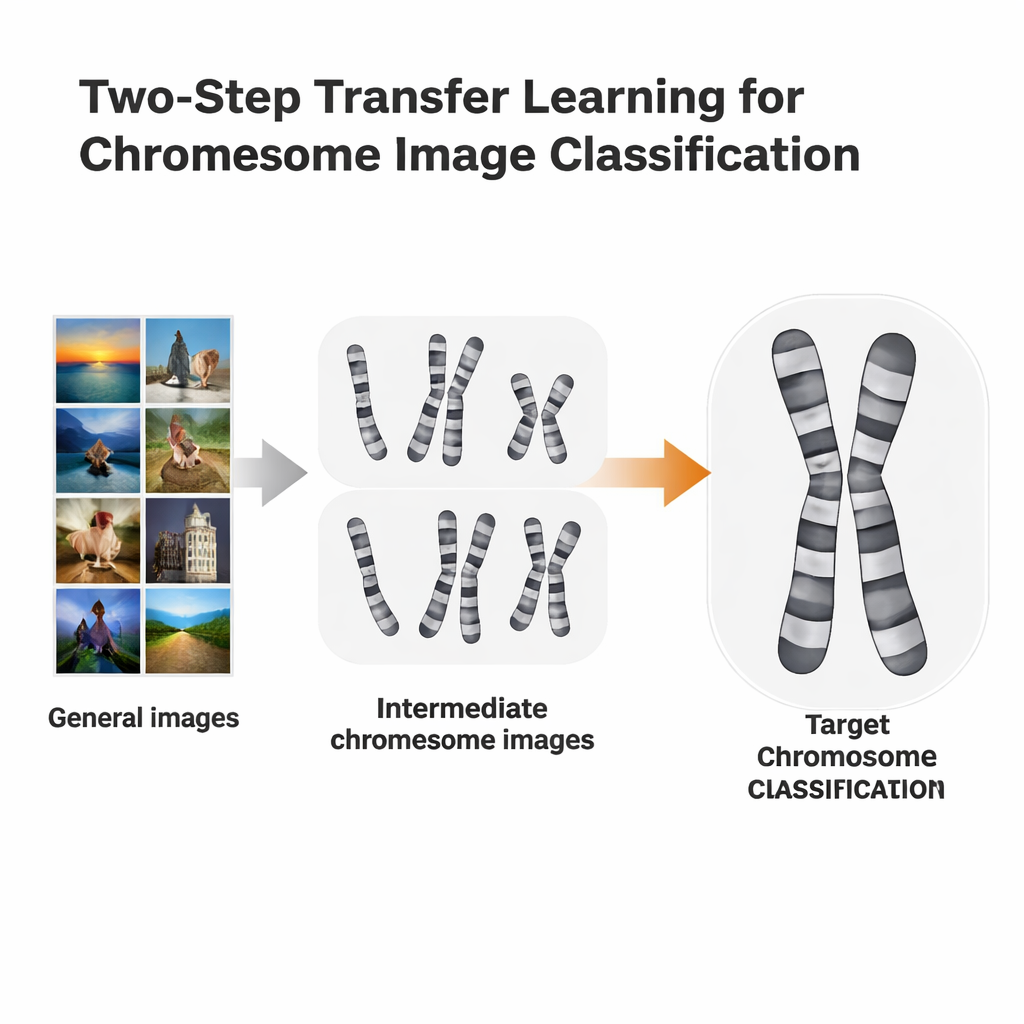
A two-step learning shortcut
The researchers tested a more tailored training route called two-step transfer learning. Instead of going straight from ImageNet to a specific chromosome task, they first fine-tuned ImageNet-trained models on chromosome images from one staining method, then fine-tuned again on a second, slightly different method. They used two open datasets: Q-band images, which are lower in quality and harder to read, and G-band images, which are cleaner and more detailed. Each dataset took turns playing the role of “stepping-stone” for the other. The idea is similar to language learning: if you already know Spanish, it may be easier to learn Italian than to jump straight from English.
Testing many computer “eyes”
To see when this extra step helps, the team trained 66 different classifiers, combining 11 popular neural network designs with three strategies: starting from scratch, fine-tuning from ImageNet only, and using two-step transfer. They measured performance using Macro-F1, a score that treats all chromosome types fairly, including the rare ones. First, they confirmed that Q-band and G-band images are statistically more similar to each other than either is to ImageNet photos, which makes them promising intermediate stepping-stones. Then they compared how well the different models learned under each strategy on both the easy (G-band) and hard (Q-band) datasets.
When the extra step pays off
On the higher-quality G-band images, almost all models already performed extremely well after simple ImageNet fine-tuning, with scores around 97–98 percent. Here, the extra two-step training gave only tiny gains—often less than a percentage point—and sometimes even hurt older network designs. In contrast, on the more challenging Q-band images, the story changed. Modern, compact architectures such as ConvNeXt, Swin Transformer, Vision Transformer, and MobileNetV3 clearly benefited from the two-step route, improving by about 0.8 to 3.3 percentage points over ImageNet alone. Visual maps of where the models “looked” showed why: with two-step transfer, the networks focused more evenly along the chromosome bands on both arms, rather than just on outlines or a single region. However, very large, older networks like VGG did not gain and sometimes got worse, suggesting that smarter design beats sheer size.
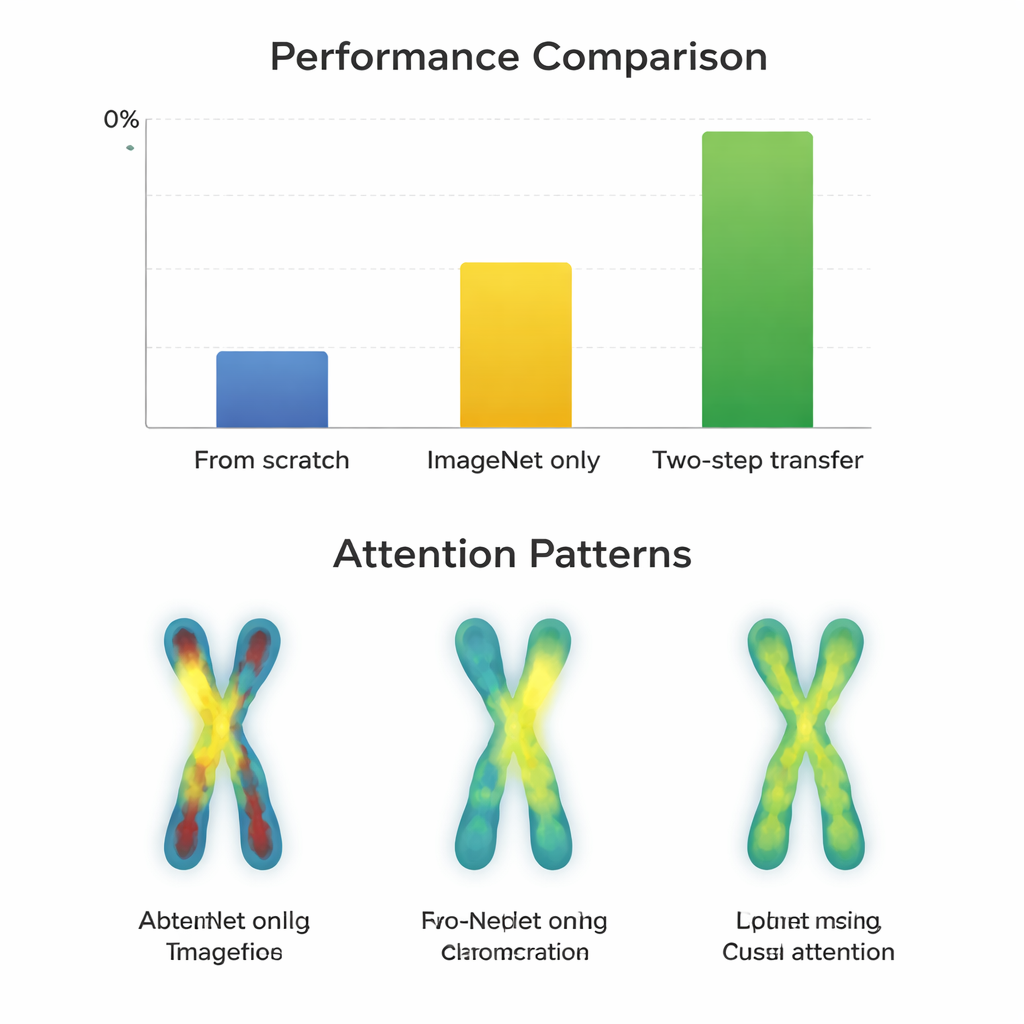
Limits set by the data itself
The researchers also examined mistakes on G-band images. Some failures were traced not to the learning strategy but to flawed input, such as chromosomes that had been cropped badly when separating overlapping shapes. In these cases, all training methods struggled, and the attention maps were scattered or fixated on misleading edges. This highlights a practical message for clinics and developers: even the best training pipeline cannot fully overcome poor image quality or preprocessing errors, especially when working with modestly sized datasets like those available for chromosome imaging.
What this means for real-world diagnosis
For non-specialists, the key takeaway is that smart reuse of related medical images can make automated chromosome reading more accurate—especially when the target data are noisy or scarce and when using modern, carefully designed neural networks. For high-quality images, standard ImageNet-based training may already be enough. But when pathologists are working with tougher datasets, an extra learning step using a closely related type of image can sharpen the computer’s “eye,” bringing performance into the 93–98 percent range. This approach could extend beyond chromosomes to many areas of medical imaging where labeled data are limited, helping bring reliable AI tools closer to everyday clinical practice.
Citation: Chen, T., Xie, C., Zhang, W. et al. ImageNet pre-training and two-step transfer learning in chromosome image classification. Sci Rep 16, 7572 (2026). https://doi.org/10.1038/s41598-026-38662-w
Keywords: chromosome classification, medical imaging AI, transfer learning, deep learning models, karyotyping