Clear Sky Science · en
ConvAHKG: Action-based hybrid knowledge graph with a dual-channel convolutional approach for drug repurposing
Finding New Uses for Old Medicines
Bringing a brand-new drug to market can take more than a decade and cost billions of dollars, and many candidates fail along the way. This study explores a smarter shortcut: using data and artificial intelligence to discover new diseases that existing, already-approved drugs might safely treat. The authors introduce a framework, called ConvAHKG, that weaves together many kinds of biomedical information into a single connected map and then uses a modern neural network to spot promising drug–disease matches—offering a faster, cheaper route to new therapies.
A Rich Map of How Drugs and Diseases Interact
At the heart of this work is a “hybrid knowledge graph,” a giant map that connects drugs, diseases, proteins, side effects, chemical structures, and biological pathways. Instead of storing just simple yes–no links (for example, “drug A treats disease B”), the map records how a drug acts on a protein—whether it activates, inhibits, or binds to it—and how proteins are involved in diseases, such as serving as biomarkers or changing in activity. The graph contains over 11,000 entities and 59 types of relationships, including detailed information on drug classifications, side effects, protein–protein contacts, and chemical substructures. By capturing these many layers of context, the graph can reflect more of the real biological complexity behind treatment effects and adverse reactions. 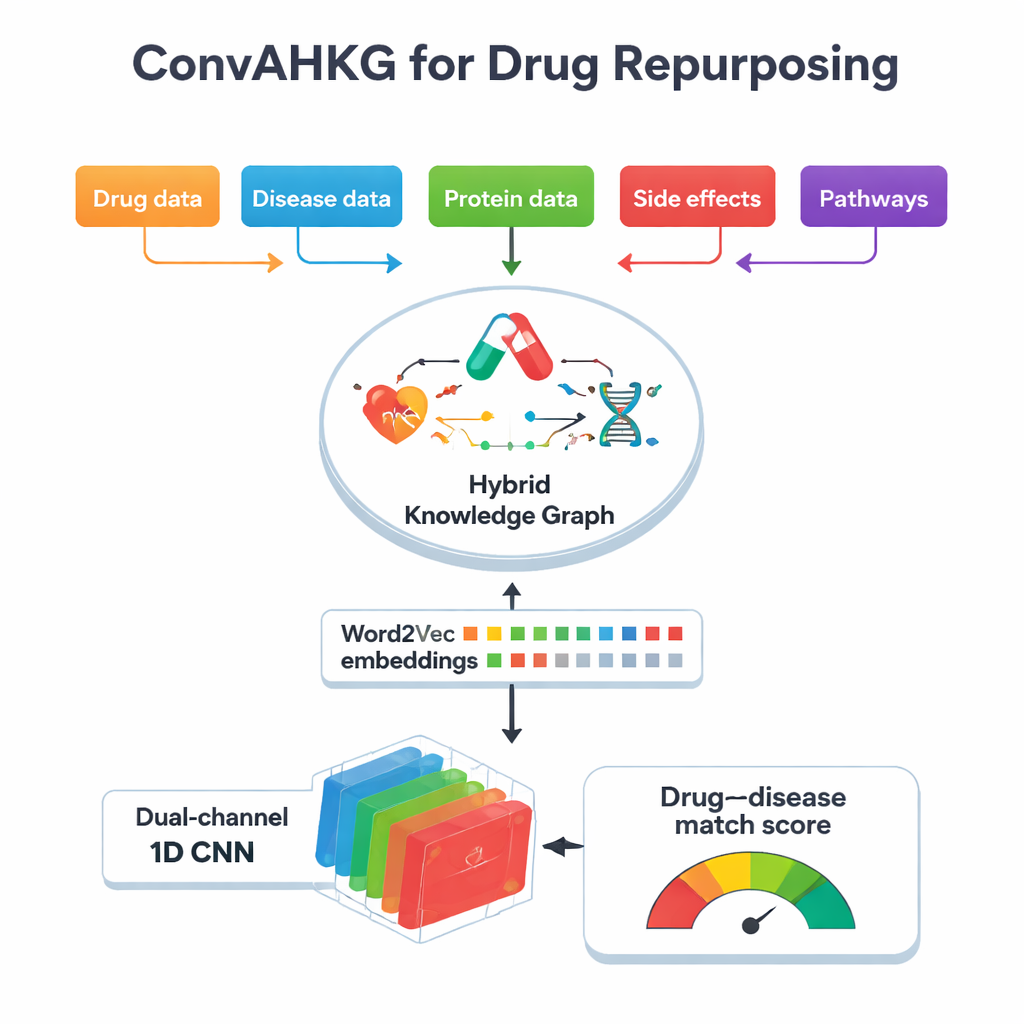
Teaching the Computer the Language of Biology
To make this complex map usable for machine learning, the team converts each connection in the graph into a simple three-part “sentence”: a head (such as a disease), a relation (such as biomarker), and a tail (such as a protein). They then apply Word2Vec, a method originally developed for natural language processing, to learn numeric “embeddings” for every drug, disease, and protein. Items that often appear together in these sentences end up close to each other in this mathematical space, much like words that share similar meanings in text. This approach is far simpler and faster than many graph-specific embedding techniques, yet still captures subtle patterns. In tests against several popular knowledge-graph embedding methods, Word2Vec matched or exceeded their predictive power while using much less computation time.
A Dual Neural Pathway for Yes/No Treatment Decisions
Once each drug and disease is translated into a numeric vector, ConvAHKG feeds them into a dual-channel one-dimensional convolutional neural network. One channel processes the drug vector and the other processes the disease vector using a sequence of convolutional filters that detect local patterns and broader motifs, inspired by designs from image recognition such as InceptionNet and AlexNet. After this separate processing, the two streams are merged and passed through several fully connected layers that output a single probability: does this drug–disease pair represent a likely effective treatment, or more of a harmful side effect? To deal with the fact that known positive pairs are much rarer than negative ones, the authors introduce a weighted loss function that penalizes missed true treatments more heavily, improving performance on these hard-to-find cases. 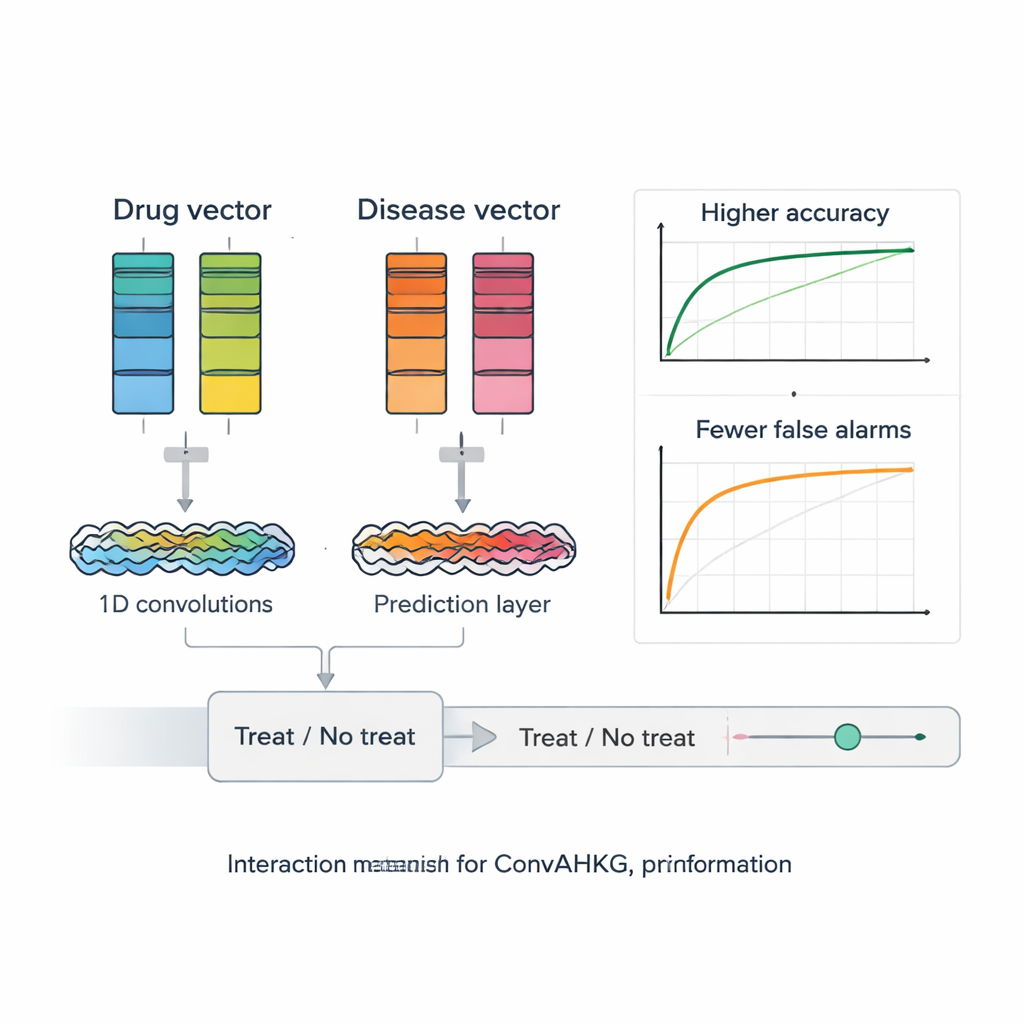
Beating Existing Methods and Spotting Cancer Candidates
The researchers rigorously test their framework against several state-of-the-art drug repurposing tools that use matrix factorization, graph neural networks, and deep autoencoders. ConvAHKG achieves an area under the ROC curve of 0.9836 and an area under the precision–recall curve of 0.9686, outperforming all competing approaches on the same benchmark dataset. They then apply the model to non-small cell lung cancer, the most common and deadly form of lung cancer worldwide. ConvAHKG highlights several drugs not previously labeled as treatments for this disease, including the antibody trastuzumab, which targets the HER2 protein and already has supporting clinical evidence in lung cancer, as well as other biologic agents and even the classic antibiotic benzylpenicillin. Docking simulations suggest that benzylpenicillin may bind strongly to DNA and to topoisomerase II alpha, an enzyme often elevated in these tumors, hinting at a possible anticancer mechanism that now warrants laboratory testing.
Why This Matters for Patients
In plain terms, ConvAHKG acts like a highly informed matchmaker between existing drugs and diseases, using a detailed map of biological actions and a powerful pattern-recognition engine to predict which old medicines might work in new settings. By treating safety-tested drugs as reusable building blocks rather than single-purpose tools, this framework could accelerate the discovery of therapies for conditions such as lung cancer, especially where conventional drug development is too slow or costly. While the predictions still need careful experimental and clinical validation, the study shows that combining rich biological knowledge with modern AI can sharply narrow the search space, bringing potentially life-saving treatments within reach more quickly.
Citation: Khodadadi AghGhaleh, M., Abedian, R., Zarghami, R. et al. ConvAHKG: Action-based hybrid knowledge graph with a dual-channel convolutional approach for drug repurposing. Sci Rep 16, 7592 (2026). https://doi.org/10.1038/s41598-026-38656-8
Keywords: drug repurposing, knowledge graph, deep learning, lung cancer, drug discovery