Clear Sky Science · en
Genome sequencing of Bacillus daqingensis and Alkalicoccus luteus reveals taxonomic insights and adaptive mechanisms
Life in Salty Places
From salt lakes to alkaline soils, some microbes thrive in conditions that would dry out and kill most other forms of life. This study dives into the DNA of two such salt-loving bacteria to understand how they survive extreme environments and to clarify where they truly belong on the tree of life. By comparing their genomes in detail, the researchers show that what were thought to be separate bacterial species are actually the same kind of organism, and they uncover the molecular tricks these microbes use to cope with intense salt stress.
Why These Salt Lovers Matter
The bacteria at the center of this work were originally named Bacillus daqingensis and Alkalicoccus luteus. Both were isolated from salty, alkaline settings such as soda lakes and saline-soil, where most life struggles. Scientists had suspected that Bacillus daqingensis might in fact belong to the genus Alkalicoccus, but its full genome sequence was missing, leaving its official status in limbo. By sequencing and analyzing the complete genomes of these microbes and comparing them with related species, the authors set out to resolve this taxonomic puzzle and, at the same time, learn how members of the genus Alkalicoccus are able to survive under such harsh conditions.
Reading and Comparing Microbial Genomes
The team grew the bacteria in the laboratory, extracted their DNA, and sequenced it using high-throughput methods. They then stitched together the millions of small DNA fragments into near-complete genome maps and checked their quality. Both Bacillus daqingensis and Alkalicoccus luteus turned out to have similarly sized genomes of about 3.4 to 3.5 million base pairs, with almost identical genetic content and very high completeness. A key genetic marker known as the 16S rRNA gene was essentially the same in both organisms: one comparison found 99.8 percent identity to previous lab measurements, and, crucially, the 16S sequences from the two strains were a perfect 100 percent match to each other.
Decoding Survival in Salt and Stress
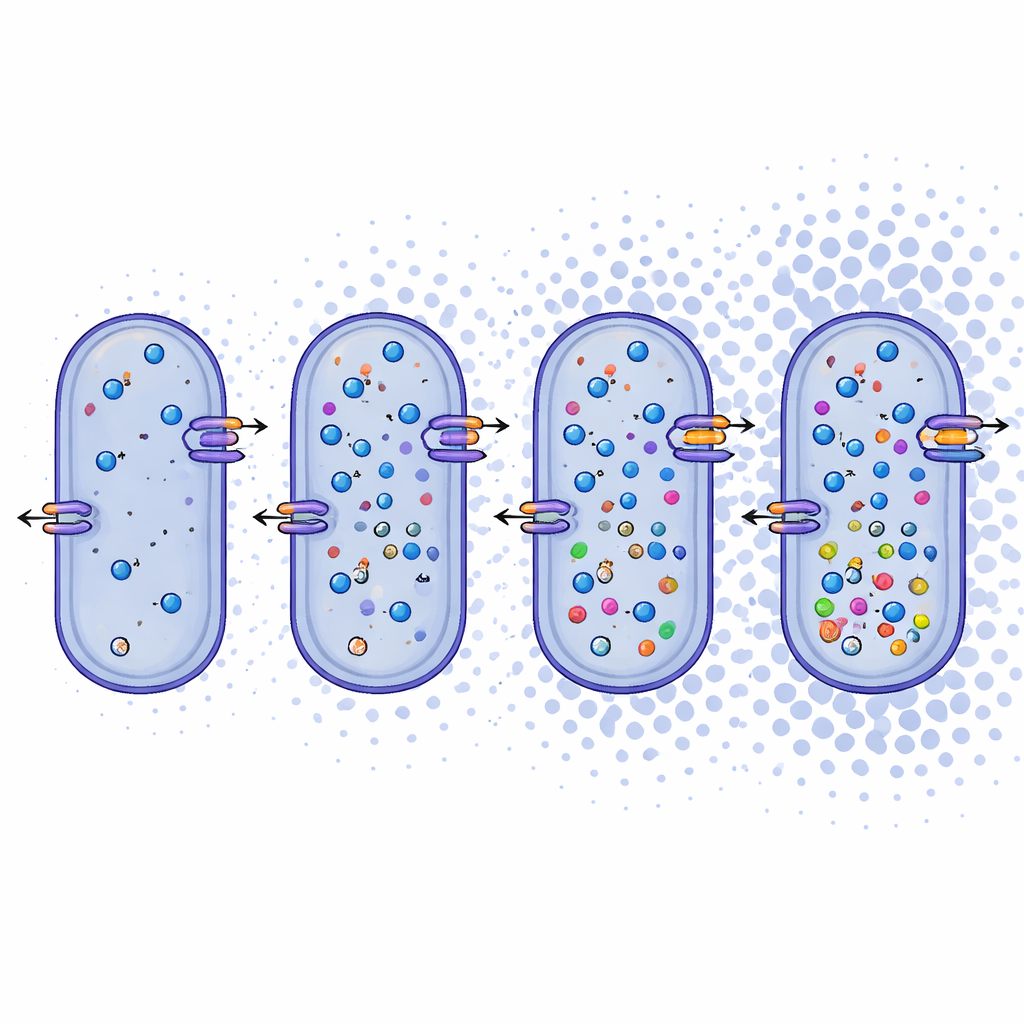
Looking beyond names, the researchers examined what the genomes reveal about how these microbes make a living. They found that Bacillus daqingensis and all surveyed Alkalicoccus species share core metabolic pathways, including standard routes for breaking down sugars and a modified cycle called the glyoxylate shunt that helps conserve carbon under stress. Most strikingly, the genomes carry a rich toolkit for coping with high salt. The bacteria appear to use two complementary tactics: “salt-in” systems that move inorganic ions like sodium and potassium across the cell membrane, and “salt-out” systems that build or import small organic molecules, such as betaine, ectoine, and certain amino acids, which act as internal cushions against dehydration. Genes for ion transporters, antiporters, and the biosynthesis of these compatible solutes are present across the genus, pointing to a robust, flexible response to osmotic stress.
Proving Who Is Who
To settle the taxonomic question, the authors compared whole genomes using several numerical yardsticks that are now standard in microbial systematics. Average nucleotide identity (ANI) measures how similar the DNA sequences are across the genome, while average amino acid identity (AAI) does the same at the protein level. Between Bacillus daqingensis and Alkalicoccus luteus, ANI reached 98.2 percent and AAI reached 98.5 percent—well above the usual cutoffs used to define the same species and even the same genus. Phylogenetic trees built from many genes, along with comparisons of shared protein clusters, showed that these two strains group tightly together and share more gene clusters with each other than with any other Alkalicoccus species. Traditional traits such as cell shape, fatty acid profiles, and key biochemical reactions were also essentially identical between them.
What This Means for Microbial Names
Putting all lines of evidence together, the study concludes that Bacillus daqingensis does not represent a distinct kind of bacterium. Instead, it belongs in the genus Alkalicoccus and is the same species as Alkalicoccus luteus, previously known as Bacillus luteus. The authors formally propose the new combination name Alkalicoccus daqingensis and treat it, along with the older Bacillus name, as later synonyms of Bacillus luteus/Alkalicoccus luteus. For non-specialists, the takeaway is that careful genome sequencing can reveal when different labels are being used for essentially the same microbe, helping clean up the naming system. At the same time, the work highlights how these salt-loving bacteria rely on a blend of ion pumps and protective molecules to stay alive in environments that would otherwise be too salty for life.
Citation: Narsing Rao, M.P., Wang, Kk., Zhu, Ky. et al. Genome sequencing of Bacillus daqingensis and Alkalicoccus luteus reveals taxonomic insights and adaptive mechanisms. Sci Rep 16, 9720 (2026). https://doi.org/10.1038/s41598-026-38640-2
Keywords: halophilic bacteria, genome sequencing, microbial taxonomy, salt stress adaptation, Alkalicoccus luteus