Clear Sky Science · en
Genome editing across Dictyostelia species enables comparative functional genetics of social amoebas
How Single Cells Team Up
Most of the time, amoebas are tiny loners crawling around in the soil, eating bacteria. Yet some of them, called social amoebas, can suddenly band together into a multicellular body with a primitive "head" and "stem." This shape-shifting lifestyle offers scientists a rare window into how simple cells learn to cooperate and specialize—key steps in the evolution of complex life. The new study describes a genome-editing toolkit that finally lets researchers probe these behaviors not just in one lab favorite, but across several distant amoeba species.
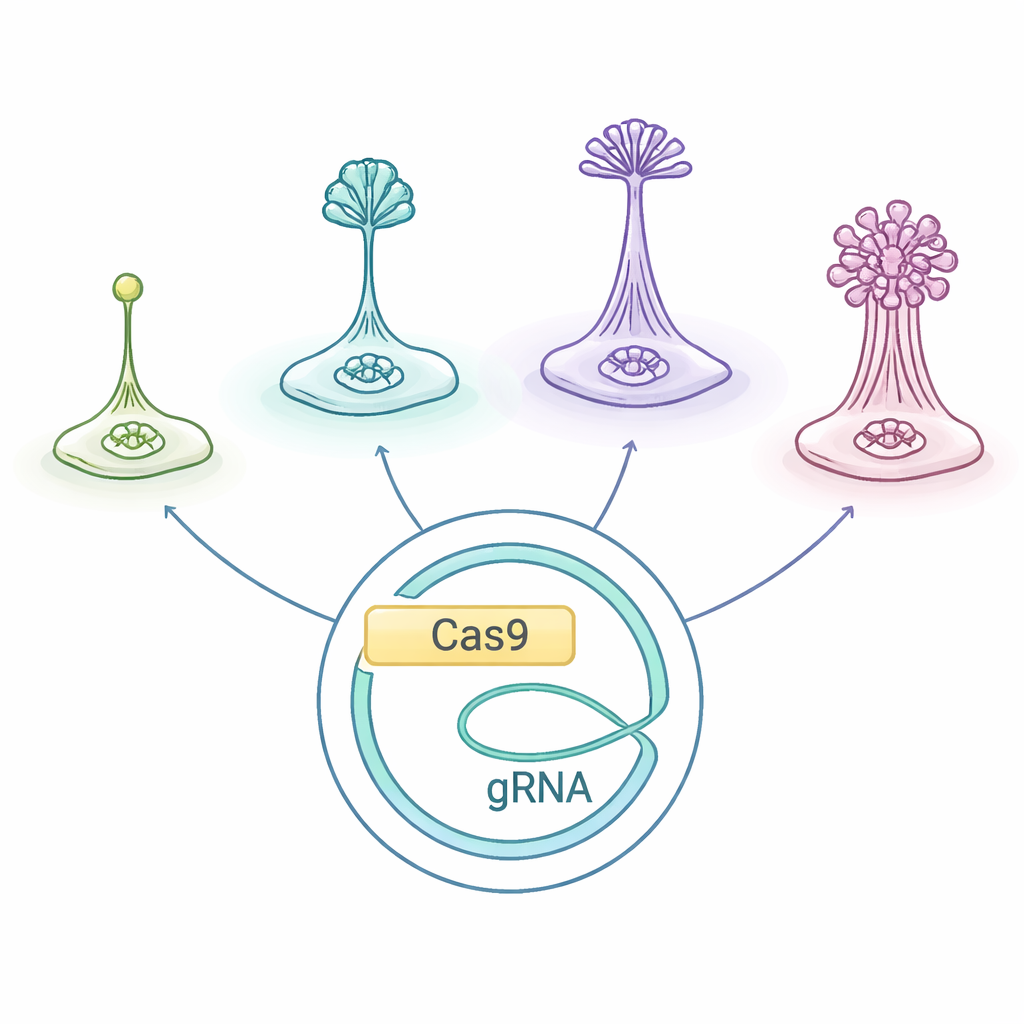
Tiny Creatures with Big Social Lives
Social amoebas, grouped together as Dictyostelia, live as single cells when food is plentiful. When they starve, they start sending out chemical distress calls and crawl toward one another, gathering into a mound that builds a miniature “fruiting body”: a column of sacrificial stalk cells that hoist hardy spores into the air. Different Dictyostelia lineages build these structures in different ways and with different sets of cell types, making them a natural laboratory for asking how multicellular bodies and division of labor evolve.
One Model Species Was Not Enough
Until now, nearly all molecular work focused on a single species, Dictyostelium discoideum, because its genes are relatively easy to tweak. That left more than a hundred other known species—some representing earlier branches of the Dictyostelia family tree—largely unexplored. These overlooked species have distinct life histories, signaling systems, and body plans. To understand how multicellularity changed over time, scientists need to manipulate the same genes in many of these species and compare the results side by side.
Bringing CRISPR to a Diverse Amoeba Family
The researchers adapted a CRISPR/Cas9 gene-editing system originally optimized for D. discoideum and tested whether it works across the Dictyostelia tree, including both early-branching and more derived species. They used a plasmid—an extra circular piece of DNA—that carries the instructions for the Cas9 “molecular scissors,” the guide RNA that points Cas9 to a chosen gene, and a drug-resistance marker. Introducing this plasmid into different amoebas allowed the team to cut two well-studied genes, called stlA and pkaC, which are important for the signals that coordinate aggregation and proper fruiting body formation. In the soil amoeba Polysphondylium violaceum, this strategy reliably produced mutants with clear developmental defects, showing that the borrowed system can work outside the usual model.
Testing Genes Across Species
Next, the team moved to more distantly related species, including Heterostelium pallidum from Group 2 and Cavenderia fasciculata from Group 1—lineages that diverged earlier in evolution. Using the same plasmid design, they disrupted the pkaC gene in both species and found that mutants failed to aggregate and build normal fruiting bodies, mirroring the severe defects seen in D. discoideum. Because CRISPR mutations created by this method do not leave behind bulky selection cassettes, the resulting strains were “marker-free.” That made it straightforward to add back a version of the gene from D. discoideum and test whether it could stand in for the local version. In all cases, the foreign pkaC was able to rescue the early step of aggregation but not the later sculpting of a mature fruiting body—evidence that the basic signaling toolkit is shared, while fine-tuned control of timing and position has evolved in species-specific ways.
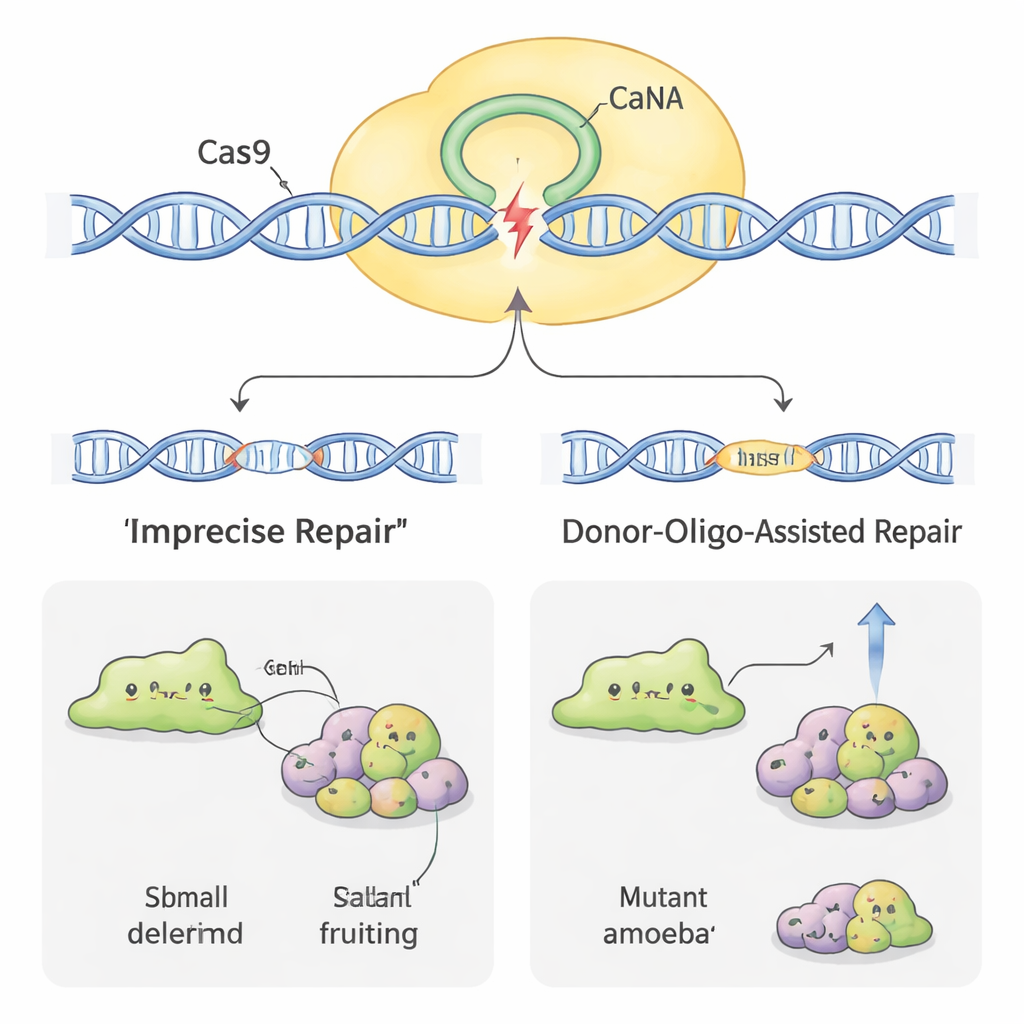
Making Editing More Efficient
Not all amoebas yielded to genome editing equally easily. In some species, such as H. pallidum, the initial success rate was very low. To boost efficiency, the scientists borrowed another trick: adding short “donor” DNA snippets along with the CRISPR plasmid. These donors carry a small tag and a stop signal flanked by short regions matching the target gene. When the cell repairs the Cas9 cut using the donor as a template, it reliably breaks the gene in a recognizable way. In D. discoideum, this raised the share of defective clones and even allowed the team to obtain mutants without using continuous drug selection. In the more stubborn H. pallidum, combining donor DNA with a few days of drug treatment increased mutation rates by nearly an order of magnitude, turning previously rare events into something close to routine.
Opening a Window on the Origins of Complexity
To a non-specialist, the details of plasmid designs and repair templates may sound technical, but the payoff is broad: a flexible CRISPR toolkit that works across several branches of the social amoeba family tree. With it, scientists can now compare how the same gene shapes development in different species, and how subtle changes in gene regulation or protein structure help generate new cell types and body plans. In other words, this work arms researchers with the genetic tools needed to watch, in living detail, how evolution tinkerered with single-celled organisms to build cooperative multicellular societies.
Citation: Oishi, S., Doi, S., Sekida, T. et al. Genome editing across Dictyostelia species enables comparative functional genetics of social amoebas. Sci Rep 16, 7457 (2026). https://doi.org/10.1038/s41598-026-38605-5
Keywords: social amoebas, CRISPR genome editing, multicellularity evolution, Dictyostelium, cell differentiation