Clear Sky Science · en
Integrative analysis of transcriptome and single-cell sequencing combined with experimental validation identifies biomarkers associated with T cell and senescence in sepsis
Why this research matters for patients with severe infections
Sepsis is a life-threatening reaction to infection that can turn a routine illness or injury into an emergency, often within hours. Doctors still lack a simple, reliable way to spot which patients are sliding toward organ failure, and there are no drugs designed specifically to correct the immune chaos that sepsis causes. This study explores how certain immune cells, called T cells, and the aging of these cells might drive sepsis—and identifies a small set of genes that could help doctors diagnose the condition earlier and point the way to new treatments.
Sepsis: when the body’s defenses go off the rails
In sepsis, the immune system first explodes into an intense inflammatory response and then often swings into a dangerous shutdown. T cells, which normally help clear infections and remember past ones, become fewer in number and less able to fight. At the same time, many immune cells show signs of “senescence” – a kind of cellular aging in which cells stop dividing but keep releasing inflammatory signals. The authors reasoned that if they could pinpoint genes linked both to T cells and to senescence, they might uncover new biomarkers that reveal who is developing sepsis and why some patients deteriorate more quickly than others.
Digging into blood at the level of single cells
The team combined several layers of data from blood samples of people with and without sepsis. They first used bulk transcriptome data (a readout of which genes are switched on) to find nearly 10,000 genes that changed their activity in sepsis. Then, using single-cell RNA sequencing, they examined tens of thousands of individual blood cells to identify more than 1,300 genes specifically altered in T cells. By overlapping these sets with a curated list of 866 genes known to be involved in cellular aging, they narrowed the field down to 45 candidates that sat at the crossroads of sepsis, T cells, and senescence. 
Finding four key genetic signposts
To filter this list further, the researchers applied two machine-learning techniques that look for the most informative features in complex data. These approaches converged on six promising genes, and when the team checked two independent patient datasets, four of them consistently stood out: PATZ1, SIN3B, BLK, and MTHFD2. In public data, PATZ1, SIN3B, and BLK were reduced in sepsis, while MTHFD2 was increased. In a small group of clinical samples tested by a sensitive method called RT-qPCR, the first three genes were again clearly lower in sepsis patients’ blood, while MTHFD2 showed no clear difference—hinting that its role may be more about how the protein functions than how much of it is made.
How these genes tie into immune balance and potential drugs
Further analyses connected each gene to broader biological pathways. PATZ1 was linked to DNA replication and T cell receptor signaling, processes essential for T cells to multiply and respond to infection. SIN3B was associated with a lipid-based signaling system that influences cell survival and inflammation. BLK, better known in autoimmune diseases, was tied to blood cell development and gut immune defenses, including production of IgA antibodies that protect the intestinal barrier. MTHFD2 was connected to protein synthesis and breakdown machinery, central to how immune cells adjust their metabolism under stress. When the team examined immune cell “infiltration” patterns in blood, higher PATZ1 levels went hand in hand with more CD8 T cells and fewer neutrophils, a balance that may favor better control of infection. Using these four genes, the researchers built an artificial neural network—a simple AI model—that distinguished sepsis from non-sepsis cases in two external datasets with near-perfect accuracy. They also used a drug–gene database and molecular docking simulations to propose several existing compounds, including the antibiotic latamoxef and the molecules emetine and dihydroergotamine, as candidates that bind strongly to these targets. 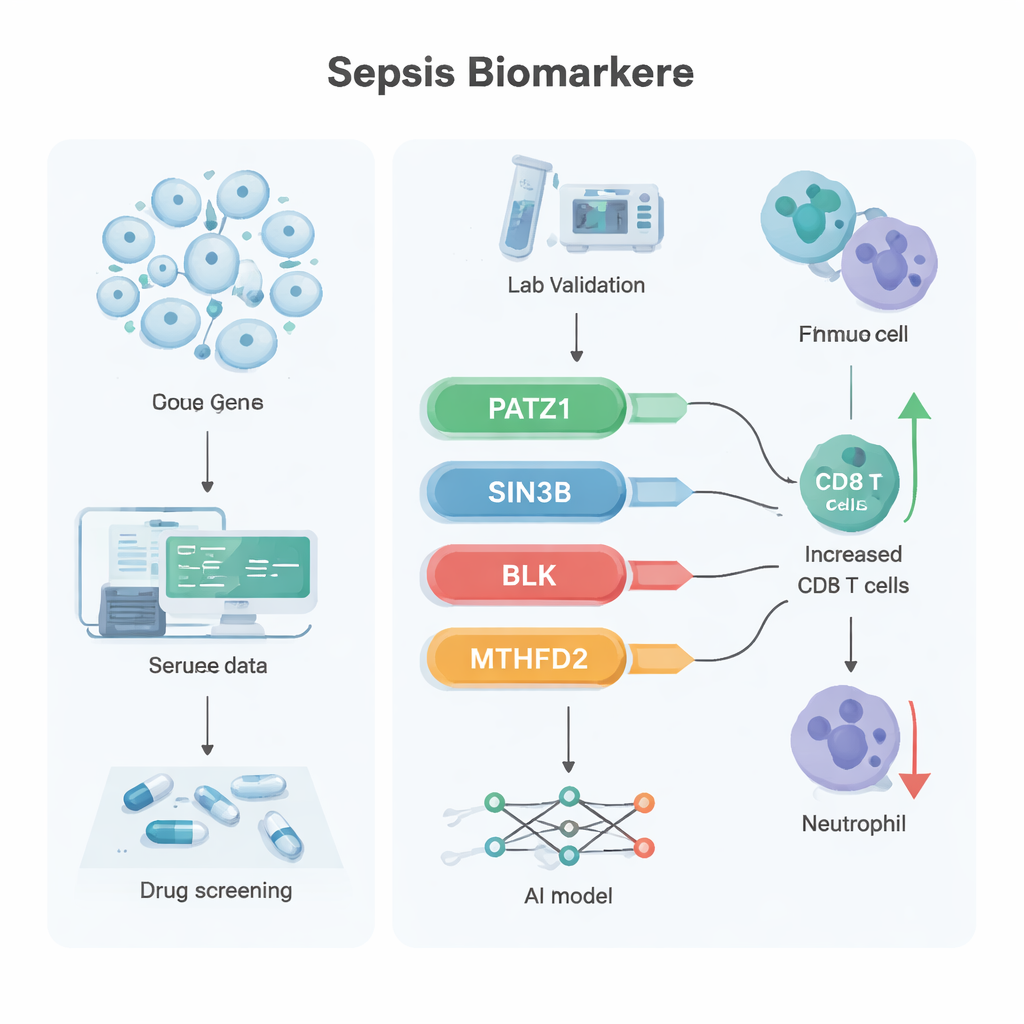
What this could mean for future care
To a layperson, the key takeaway is that this study identifies four genes acting as warning beacons on T cells that misfire and age prematurely during sepsis. Together, these genes form a potential blood test panel that might one day help doctors diagnose sepsis earlier and more accurately than current tools. They also offer clues to how the immune system’s energy use and repair systems break down, suggesting new angles for drugs designed to restore healthy T cell function rather than simply dampening inflammation. While larger studies and laboratory experiments are needed before any of these findings reach the bedside, this work charts a promising path toward more precise, biology-based diagnosis and treatment of sepsis.
Citation: Yang, K., Hu, Y., Ma, C. et al. Integrative analysis of transcriptome and single-cell sequencing combined with experimental validation identifies biomarkers associated with T cell and senescence in sepsis. Sci Rep 16, 7109 (2026). https://doi.org/10.1038/s41598-026-38559-8
Keywords: sepsis biomarkers, T cells, immune senescence, single-cell sequencing, immunometabolism