Clear Sky Science · en
Automated and robust nonrigid registration of serial section microscopic images using PiCNoR
Seeing Tissues in 3D Without Losing the Details
Modern biology often relies on turning thin slices of tissue into full 3D views of organs and brains. But when each microscopic slice is cut, stained, and imaged, it can stretch, tear, or shift. If these slices are not lined up properly, the resulting 3D picture can be misleading. This paper introduces a new computer method called PiCNoR that helps scientists align such images more accurately and automatically, so that fine structures in embryos and brains are preserved in 3D reconstructions.
Why Lining Up Slices Is So Difficult
To build a 3D view, researchers image long series of ultra-thin sections taken from the same piece of tissue. In theory, each slice should fit perfectly above the previous one, like cards in a neatly stacked deck. In reality, each slice can warp in a slightly different way during cutting and staining. Colors may vary, pieces may stretch, and features can shift. Traditional “rigid” alignment methods assume each entire slice only shifts or rotates, which is often not enough. More flexible, “elastic” methods exist, but they can be slow, require careful tuning by experts, or depend strongly on image brightness, which can change from slice to slice.
A New Way: Local Pieces That Work Together
PiCNoR takes a different, more local view of the problem. Instead of trying to bend an entire image at once, it first divides each slice into many regions based on the pattern of features in the image. Within each region, the method finds matching points between two neighboring slices using robust feature detectors, and estimates how that region must rotate and shift to line up. These local motions are then checked for plausibility and gently combined, so that every pixel in the slice gets a motion that smoothly blends information from surrounding regions. The result is a flexible, “non-rigid” alignment that still behaves in a controlled and realistic way.
Letting the Data Choose the Right Complexity
A key challenge in any region-based method is deciding how many regions to use: too few, and the method cannot correct fine deformations; too many, and it becomes unstable and slow. Previous approaches often relied on trial and error, repeatedly checking the visual quality of the result. PiCNoR avoids this manual tuning by using a statistical tool called the Bayesian information criterion, which automatically balances the amount of detail against the risk of overfitting. In practice, this means the algorithm can decide on its own how many regions are needed for a given dataset, without human supervision, which saves time and reduces bias.
Keeping Results Reliable and Efficient
Not every estimated local motion is trustworthy—some can be distorted by noise or poor matches. PiCNoR addresses this by representing each region as a node in a graph, where neighboring regions influence one another. Motions that look unrealistic in terms of rotation or shift are replaced by a weighted average of the more reliable motions next door. A compact mathematical representation helps combine these motions efficiently. Finally, each pixel’s movement is computed as a probability-weighted blend of the regional motions, ensuring that transitions between regions stay smooth, without sudden kinks or folds in the tissue.
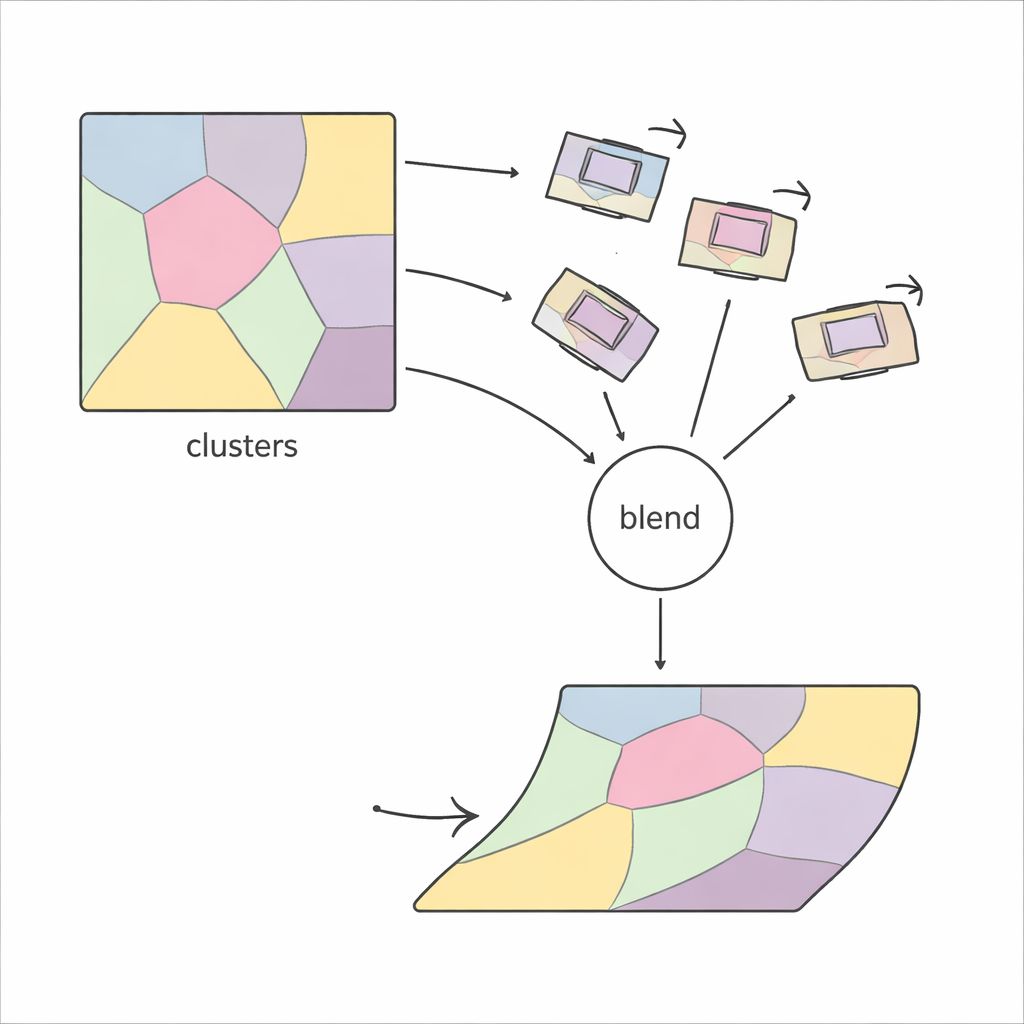
Proving It Works on Real Biological Data
The researchers tested PiCNoR on three very different datasets: human embryo sections from the Kyoto collection, an electron microscopy stack of a fruit fly nerve cord, and a new light-microscope stack from the hippocampus of a rat brain. Across these examples, PiCNoR consistently produced better overlap between slices than standard rigid and widely used non-rigid methods. It maintained continuity of delicate structures in 3D views and avoided the exaggerated distortions sometimes seen with other tools. Importantly, it did this while using fewer local regions than some competitors, and with computational costs that remain practical for large stacks.
What This Means for Future 3D Microscopy
For non-specialists, the takeaway is that PiCNoR offers a more reliable way to turn stacks of 2D microscope images into faithful 3D reconstructions. By automatically choosing how detailed the alignment should be and by safeguarding against bad local corrections, it preserves the true shapes of tissues while keeping processing time manageable. This makes it easier for biologists and pathologists to trust what they see in 3D, whether they are studying how an embryo develops or how brain cells are arranged, and it lays groundwork for more accurate, automated analysis of complex microscopic datasets.
Citation: Adi, P.M., Shabani, H. & Mansouri, M. Automated and robust nonrigid registration of serial section microscopic images using PiCNoR. Sci Rep 16, 7559 (2026). https://doi.org/10.1038/s41598-026-38548-x
Keywords: 3D microscopy, image registration, brain imaging, histology, non-rigid alignment