Clear Sky Science · en
Evaluation of commercial kits and purification approaches for DNA extraction from atmospheric samples for 3rd generation sequencing without amplification
Why the Air We Breathe Holds Hidden Clues
The air around us is full of invisible life: grains of pollen, fungal spores, bacteria, even traces of viruses. These airborne passengers can trigger allergies, spread disease, and quietly shape ecosystems. To understand them, scientists want to read their DNA directly from air samples—but there is usually very little biological material to work with. This study asks a practical question with big implications: how can we best pull fragile DNA out of everyday air filters so it can be read by modern long-read sequencing machines without extra amplification steps?
Catching Life on Invisible Dust
Air-quality networks around the world already collect particles on large glass-fiber filters to monitor pollution. The authors saw an opportunity: use those same filters to study biological material in the atmosphere. The catch is that these filters typically hold only tiny amounts of DNA compared with soil or water, and many organisms in the air—like pollen or dried bacteria—are hard to break open. The team had previously built a powerful but cumbersome method that used harsh chemicals to clean up DNA from these filters. It worked well, even when DNA was scarce, but it was slow, labor-intensive, and relied on hazardous reagents that are not ideal for routine monitoring.

Building a Safer and Faster Lab Routine
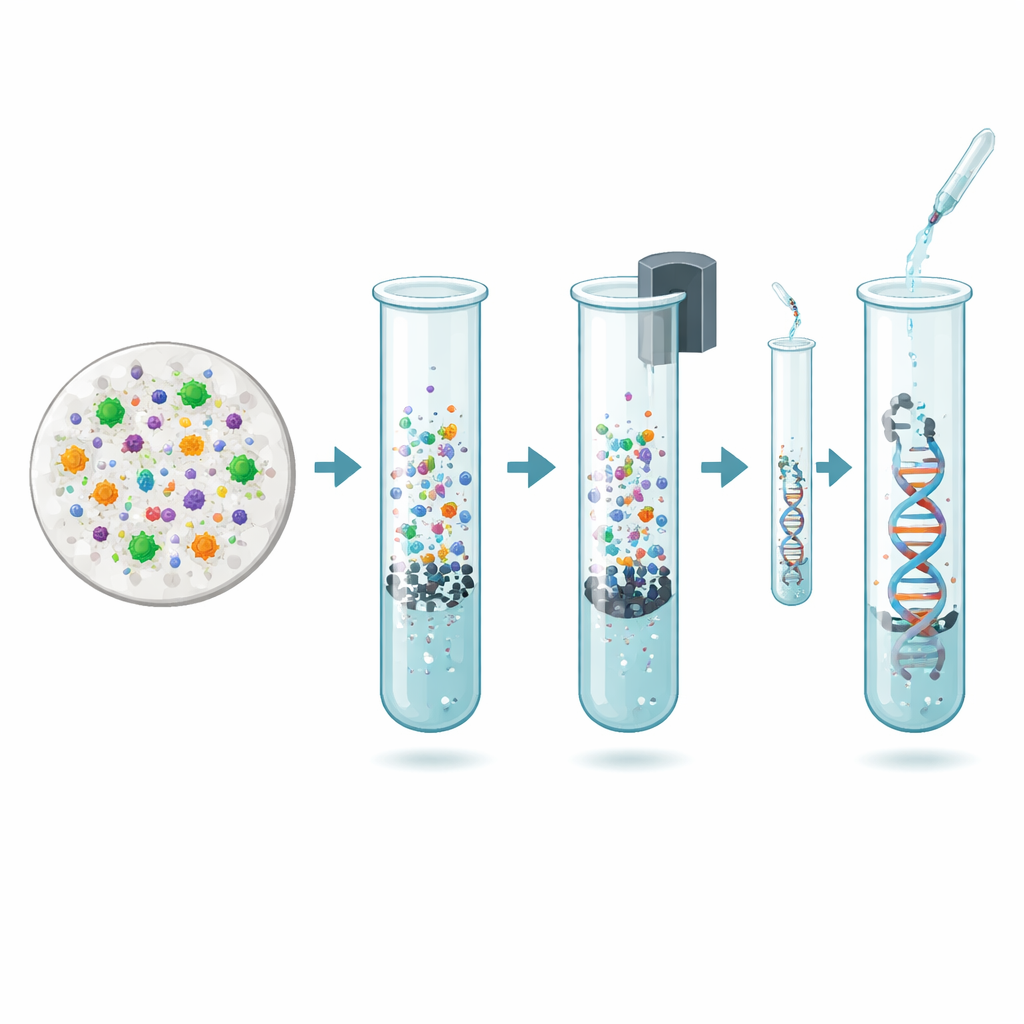
The new work focuses on updating that earlier protocol to make it safer, simpler, and better suited for large studies. The authors designed an improved in-house method that keeps the careful multi-step breakdown of cells—using enzymes, detergents, and gentle heating—but replaces the dangerous solvent purification step with magnetic beads. These tiny beads are coated to grab DNA molecules and can be pulled out of solution with a magnet, leaving many contaminants behind. The researchers then compared this updated method with their original protocol and with five popular commercial kits that use column-based purification approaches originally optimized for plant or soil samples.
Putting Extraction Methods to the Test
To make the comparison fair, the team used archived filters collected on the roof of the Finnish Meteorological Institute in Helsinki during a bioaerosol campaign. They chose one filter known to contain relatively high amounts of DNA and another with much lower levels, and then cut them into identical pieces so every method handled the same starting material. They evaluated each approach by measuring how much DNA it produced, how clean that DNA was, and how consistent the results were between repeated tests. Two of the in-house approaches—the original solvent-based protocol and the new magnetic-bead method—as well as one soil kit stood out for higher yields on the DNA-rich filter. However, when the initial DNA level was low, only the original, harsher protocol reliably recovered enough genetic material.
Does the Extracted DNA Truly Tell the Story?
Quantity is not the only concern: the DNA must also be intact and representative of the community in the air. The team checked how clean the samples were using light-absorption measurements and then fed selected extracts directly into an Oxford Nanopore long-read sequencer without any DNA multiplication step. This is important, because amplification can distort the apparent mix of species. The sequencing results showed that both the magnetic-bead protocol and the original solvent-based method could generate long, diverse reads with a broad range of base compositions, which Nanopore technology can handle well. At the same time, the two methods did not recover the same balance of organisms: the magnetic-bead method tended to favor tougher particles like pollen, while the solvent-based method captured more bacterial DNA, likely because of differences in how easily various cells are broken and preserved on the filters.
What This Means for Watching the Air We Share
For public-health monitoring and environmental surveys, the study delivers a clear message in plain terms. If the air sampler collects enough biological material, the new magnetic-bead protocol offers a safer, quicker way to prepare air filters for state-of-the-art long-read sequencing. Where DNA is scarce, the more demanding solvent-based method is still needed to avoid losing almost everything. Off-the-shelf commercial kits, although convenient, did not perform well enough for these challenging atmospheric samples. Together, the two in-house methods provide a practical toolbox: one tuned for high-yield, routine work, and another reserved for the faintest biological signals in the air we breathe.
Citation: Salokas, J., Sofieva-Rios, S., Paatero, J. et al. Evaluation of commercial kits and purification approaches for DNA extraction from atmospheric samples for 3rd generation sequencing without amplification. Sci Rep 16, 8402 (2026). https://doi.org/10.1038/s41598-026-38534-3
Keywords: airborne DNA, bioaerosols, metagenomics, long-read sequencing, environmental monitoring