Clear Sky Science · en
Integrated multi-dataset screening to predict prognosis and identify immunotherapy gene targets in hepatocellular carcinoma patients
Why this matters for people with liver cancer
Hepatocellular carcinoma, the most common form of primary liver cancer, kills hundreds of thousands of people every year. Many patients with the same stage of disease respond very differently to treatment, especially to modern immunotherapies that aim to unleash the body’s own immune system. This study asks a simple but crucial question: can we read a tumor’s gene activity like a fingerprint to predict who will do poorly, who will respond to immune-based drugs, and which genes might be the best new treatment targets?
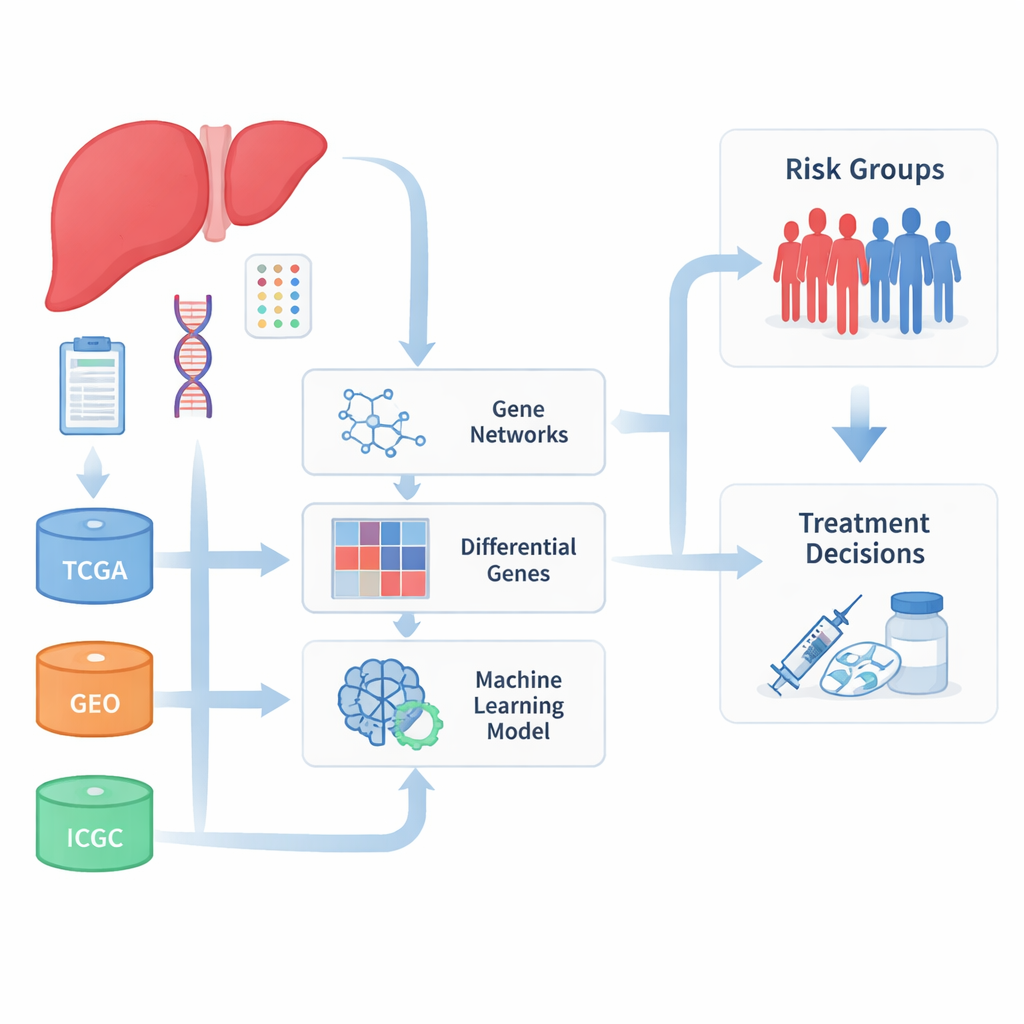
Bringing many datasets together
The researchers began by pooling large collections of liver cancer samples from several international databases that store tumor gene activity and clinical outcomes. By combining data from The Cancer Genome Atlas, the International Cancer Genome Consortium, and multiple Gene Expression Omnibus studies, they assembled a much larger and more diverse patient set than any single hospital or project could provide. Because these datasets were produced in different labs and with different methods, the team first spent substantial effort correcting for technical differences so that real biological signals, rather than laboratory noise, would drive their results.
Finding gene patterns linked to outcome
With the cleaned data in hand, the team searched for groups of genes that tended to turn on and off together and that also tracked with how patients fared. Using a network-style approach, they clustered thousands of genes into modules and then focused on those modules most strongly tied to tumor behavior and patient survival. They also compared tumors to non-tumor tissue to find genes that were clearly more or less active in cancer. The overlap between these two views yielded a set of 93 genes that were both altered in liver cancer and tightly connected to key disease features, many involved in how the liver processes drugs and handles toxic chemicals.
Building a ten-gene risk score
To turn these gene lists into something doctors might eventually use, the authors turned to machine learning. They tested more than one hundred combinations of feature-selection and survival-prediction algorithms, judging them by how accurately they could separate patients into better- and worse-outcome groups across multiple independent cohorts. From this large search, they distilled a compact signature of ten genes that together formed a risk score. Patients with high scores consistently had shorter overall, disease-free, and progression-free survival, both in the main datasets and in outside validation groups. Among these genes, TYMS stood out as a strong indicator of poor prognosis, while APOL3 and FBXO2 were linked with more favorable outcomes.
Clues from the tumor’s immune neighborhood
The study went beyond prediction to ask why these genes matter. By using several computational tools, the team estimated which types of immune cells were present in each tumor and how strongly the ten-gene score related to that immune landscape. High-risk tumors tended to show immune patterns and gene changes associated with more mutations and with signs of immune escape, including links to well-known checkpoint molecules such as PD-1 and CTLA-4. They also examined actual mutation profiles and found that high-risk tumors harbored more frequent alterations in classic cancer drivers like TP53. Finally, blood tests in patients and healthy volunteers confirmed that TYMS was elevated and FBXO2 reduced in people with liver cancer, supporting the idea that these genes are biologically active in the disease, not just statistical artifacts.
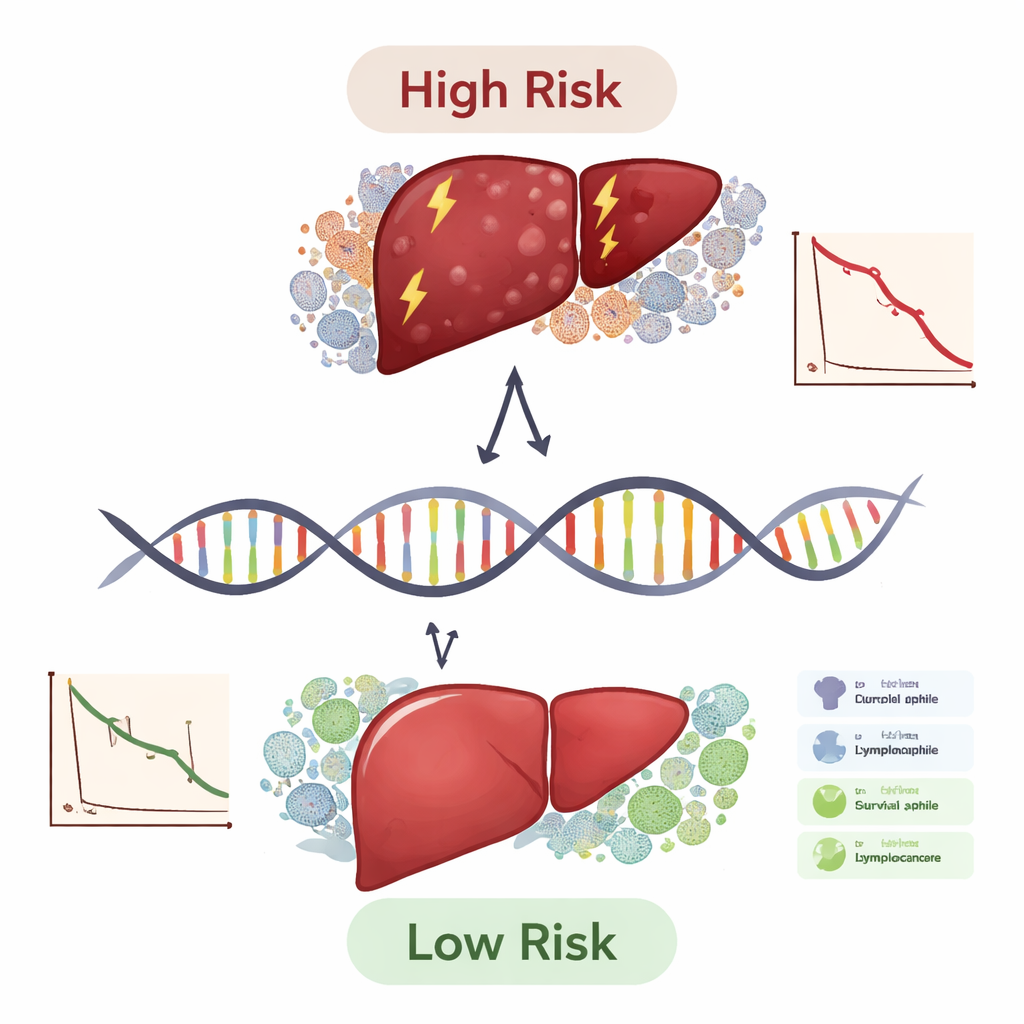
What this means for patients and doctors
In practical terms, this work offers a blueprint for using a small panel of genes to sort liver cancer patients into risk groups and to hint at who might benefit most from immune-based treatments. The ten-gene score is not yet a clinic-ready test, but it performed better than standard staging systems alone and remained useful across different patient subgroups. Just as cholesterol panels guide heart disease prevention, a gene panel like this could one day help oncologists choose more aggressive therapy for high-risk patients, spare low-risk patients from unnecessary side effects, and point drug developers toward new targets such as TYMS, APOL3, and FBXO2. Larger prospective studies and laboratory experiments will be needed, but this integrated analysis marks a significant step toward more personalized, biology-driven care for liver cancer.
Citation: Zhou, L., Zhang, W., Liu, Z. et al. Integrated multi-dataset screening to predict prognosis and identify immunotherapy gene targets in hepatocellular carcinoma patients. Sci Rep 16, 7014 (2026). https://doi.org/10.1038/s41598-026-38424-8
Keywords: hepatocellular carcinoma, gene signature, immunotherapy, prognosis, tumor microenvironment