Clear Sky Science · en
Machine learning-based prognostic model of stemness and angiogenesis-related genes for predicting prognosis and immune infiltration in patients with HCC
Why this research matters for liver cancer patients
Hepatocellular carcinoma, the most common form of liver cancer, often comes back or spreads even after treatment. This study explores why that happens and how doctors might better predict which patients are at highest risk. By combining big genetic datasets with modern machine learning, the researchers built a tool that links two powerful cancer traits—"stemness" (cells that behave like stubborn seeds) and new blood vessel growth—to patients’ chances of survival and their likely response to immune‑based therapies.
Cancer “seed cells” and new blood vessels
Many tumors contain a small group of cells that act like seeds. These cancer stem cells can renew themselves, resist treatment, and restart the tumor after surgery or chemotherapy. At the same time, tumors need to build new blood vessels to bring in oxygen and nutrients so they can grow and spread. Recent work shows these two processes are intertwined: stem‑like cancer cells release signals that encourage blood vessel growth, while the vessel‑rich environment helps those seed cells survive. Understanding and jointly targeting this partnership is especially important in liver cancer, which is both highly vascular and prone to recurrence.
Using big data to sort patients by risk
The researchers began by searching a large human gene database for genes linked both to stem‑like behavior and to blood vessel formation, ending up with more than 2,600 overlapping candidates. They then examined liver cancer samples from hundreds of patients whose tumor gene activity and clinical outcomes were recorded in public cancer databases. With statistical tests and clustering methods, they grouped patients into two genetic subtypes that showed clear differences in stage, tumor size, and survival, suggesting that combined stemness and vessel‑related signals capture meaningful biology in this disease.
Building a nine‑gene risk score
From this large gene pool, the team used a machine‑learning approach to narrow down to just nine key genes whose combined activity best predicted how long patients lived. These nine genes were used to calculate a risk score for each person. Patients whose tumors had a high score had significantly shorter overall survival, both in the original dataset and in an independent group of liver cancer patients used for validation. The model’s accuracy was competitive with or better than many existing tools, and when the score was combined with simple clinical information such as tumor stage in a chart called a nomogram, prediction of 1‑, 3‑, and 5‑year survival improved further.
Links to the immune system and treatment response
The team also asked what the nine‑gene score revealed about the tumor’s immune environment. Low‑risk tumors tended to be more “inflamed,” with higher levels of various immune cells and stronger signals of attack pathways. High‑risk tumors, in contrast, showed patterns consistent with immune evasion and higher mutation rates in important genes such as TP53. Using established computer tools that estimate how tumors might respond to modern immunotherapy drugs, the study suggested that low‑risk patients could respond better to immune checkpoint blockers than high‑risk patients. While these predictions still need to be tested in real treatment settings, they point to a practical way of identifying patients who might benefit most from such drugs.
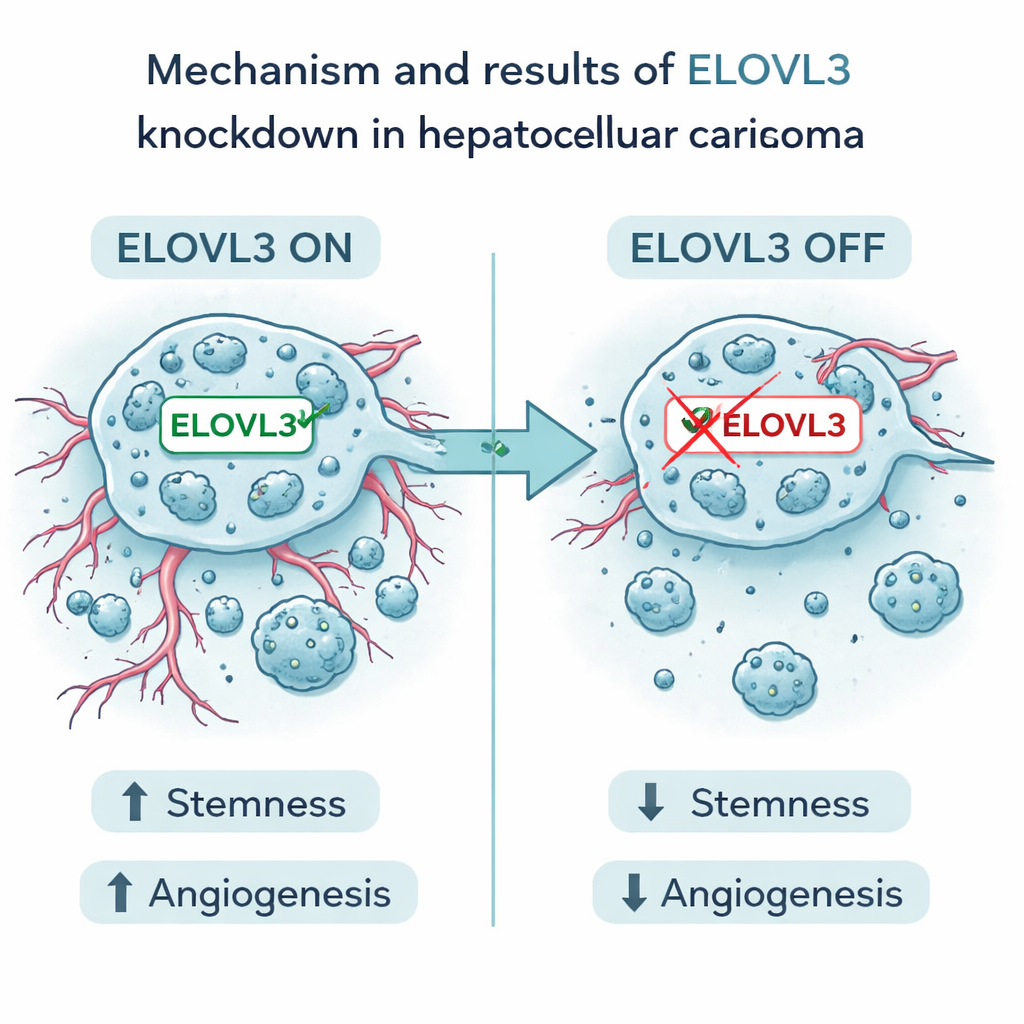
Zooming in on one promising target
Among the nine genes, one called ELOVL3 stood out. High levels of this gene were tied to worse survival. In laboratory experiments, the authors reduced ELOVL3 in human liver cancer cells. This made the cells less able to form stem‑like clusters and lowered the levels of well‑known stemness markers. It also reduced production and release of two major vessel‑promoting factors, VEGFA and FGF2. The cancer cells grew, migrated, and invaded less, and in mouse models, tumors with reduced ELOVL3 were smaller and showed fewer features of aggressive disease. These findings support the idea that ELOVL3 helps fuel both the seed‑like behavior of liver cancer cells and their ability to build supporting blood vessels.
What this could mean for future care
In plain terms, this work delivers two main messages. First, a simple nine‑gene score derived from tumor samples can sort liver cancer patients into higher‑ and lower‑risk groups and hint at how they might respond to immune‑based treatments. Second, one of the genes in that score, ELOVL3, appears to sit at a crossroads between the cancer’s ability to regenerate and its ability to grow new blood vessels, making it an attractive future drug target. Although the model and target need confirmation in prospective clinical trials, they open the door to more personalized liver cancer care and to combination strategies that block both cancer “seed cells” and their blood supply.
Citation: Cheng, F., Shi, Y., Gao, X. et al. Machine learning-based prognostic model of stemness and angiogenesis-related genes for predicting prognosis and immune infiltration in patients with HCC. Sci Rep 16, 7271 (2026). https://doi.org/10.1038/s41598-026-38379-w
Keywords: hepatocellular carcinoma, cancer stem cells, angiogenesis, prognostic gene signature, ELOVL3