Clear Sky Science · en
Robust imputation-based method for eye, hair, and skin colour prediction from low-coverage ancient DNA
Seeing the Faces Behind Ancient DNA
When archaeologists uncover ancient bones, they rarely know what the people looked like in life—what color their eyes were, how dark their skin was, or whether their hair was black, blond, or red. This study presents a new way to read those visible traits from extremely damaged DNA, allowing scientists to sketch a more human picture of individuals from the distant past and to apply the same ideas to degraded forensic samples today.
Why Old DNA Is So Hard to Read
DNA from long-buried remains is a biological wreck. Time breaks it into tiny fragments and sprinkles in chemical damage that changes one genetic letter into another. Most ancient genomes are sequenced only at “low coverage,” meaning that many positions are read once—or not at all. Existing forensic tools such as HIrisPlex-S, which can predict eye, hair, and skin color from 41 key DNA markers, were built for modern, high-quality DNA and expect reliable information at both copies of each marker. With ancient material, that information is often missing or uncertain, so traditional methods either fail outright or return very shaky predictions.
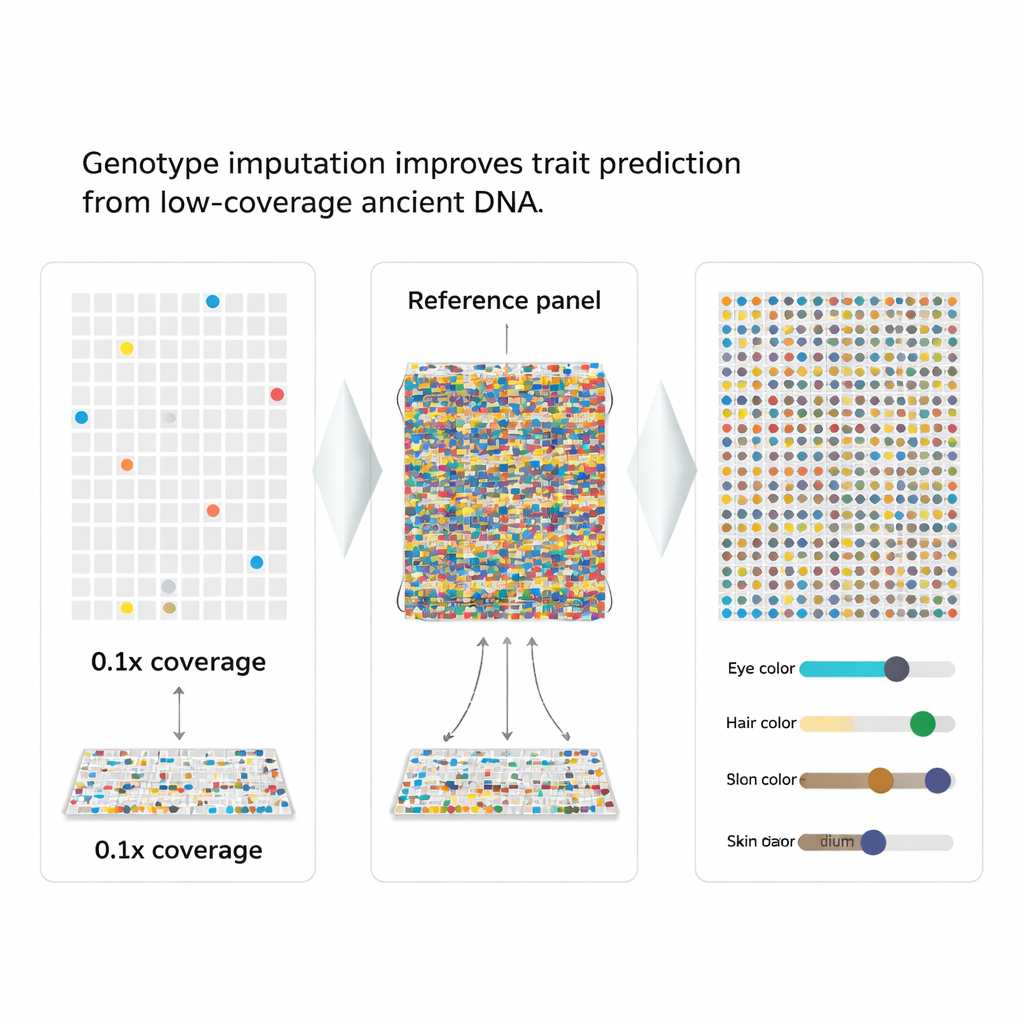
Filling the Gaps with Smart Guesswork
The authors turned to a strategy called genotype imputation, which uses patterns in a large reference panel of modern genomes to “fill in” missing pieces in a damaged one. Because nearby genetic markers are inherited together in chunks, a partially observed pattern can point strongly to the most likely missing letters. The team wrapped this idea into a new workflow, aHISplex, which starts from aligned DNA reads, runs state-of-the-art imputation software, converts the results into the exact format required by HIrisPlex-S, and then automatically turns the predicted probabilities into clear trait categories for eye, hair, and skin color.
Testing Accuracy on Modern and Ancient People
To see how well the method works, the researchers took 93 modern genomes with known traits and artificially “downsampled” them to mimic very low coverage, as poor as one-tenth of a full read-through. They then imputed the missing markers and compared the results to the true data. Even at just 0.5× coverage, the overall error rate for the 41 markers stayed below about 2%, and the most serious mistakes—flipping a marker completely from one homozygous state to the opposite—were extremely rare. Most eye and hair color predictions matched the known traits, and skin color predictions shifted only slightly among neighboring shade categories.
Challenges with Rare Traits and Ancient Diversity
Not all traits are equally easy to recover. Red hair, driven largely by rare variants in the MC1R gene, proved harder: the method almost never invented red hair where it did not exist, but it sometimes missed true red-haired individuals, calling them blond or dark blond instead. The team also applied their workflow to 31 genuinely ancient individuals with high-quality genomes, then pretended those genomes were low coverage and imputed them back. Again, overall accuracy was high, with about 95% of key markers correctly imputed at 0.5× coverage. However, a small set of marker–sample combinations, especially in individuals whose genetic backgrounds are poorly represented in modern reference panels, showed systematically higher error. This “reference bias” could, for example, turn a blue-eyed genotype into a brown-eyed prediction in a fraction of runs.
Bringing Ancient People Into Sharper Focus
Despite these caveats, the study shows that it is now practical to predict eye, hair, and skin color for many ancient individuals even when their DNA is extremely sparse, at only 0.1–0.5× coverage. The new aHISplex pipeline automates the complex steps between raw sequencing files and user-friendly trait calls, making it accessible to archaeogenetics labs and potentially to forensic teams handling degraded samples. As reference panels grow to include more diverse and ancient genomes, and as scientists discover additional trait-linked markers, this approach should become even more precise—helping us move from anonymous bones and DNA sequences to vivid, ethically considered portraits of people who lived long ago.
Citation: Maróti, Z., Nyerki, E., Török, T. et al. Robust imputation-based method for eye, hair, and skin colour prediction from low-coverage ancient DNA. Sci Rep 16, 7371 (2026). https://doi.org/10.1038/s41598-026-38372-3
Keywords: ancient DNA, forensic phenotyping, eye hair skin colour, genotype imputation, archaeogenetics