Clear Sky Science · en
Reference gene variability across age and sex in 5XFAD mice highlights normalization challenges in Alzheimer’s models
Why this mouse study matters for Alzheimer’s research
When scientists look for early warning signs or treatment effects in Alzheimer’s disease, they often measure how strongly different genes are turned on or off in the brain. But to tell whether a gene has truly changed, they first need a steady “ruler” for comparison: reference genes that are assumed to stay constant. This paper shows that, in a fast-progressing mouse model of Alzheimer’s disease, those supposedly steady genes are not nearly as stable as many researchers assume—especially across different ages, brain regions, and between males and females.
How scientists read the brain’s genetic activity
Modern brain research often relies on a technique called RT-qPCR, which counts how much of a given gene’s message is present in a tissue sample. Because samples never contain exactly the same total amount of material, scientists normalize their measurements to one or more reference genes, often called “housekeeping” genes, that are expected to be constant across conditions. In practice, however, diseases like Alzheimer’s dramatically alter cell types, metabolism, and inflammation in the brain, which can quietly shift the levels of these reference genes. If the reference itself drifts, any measurement built on it can be misleading, exaggerating or hiding real biological changes.
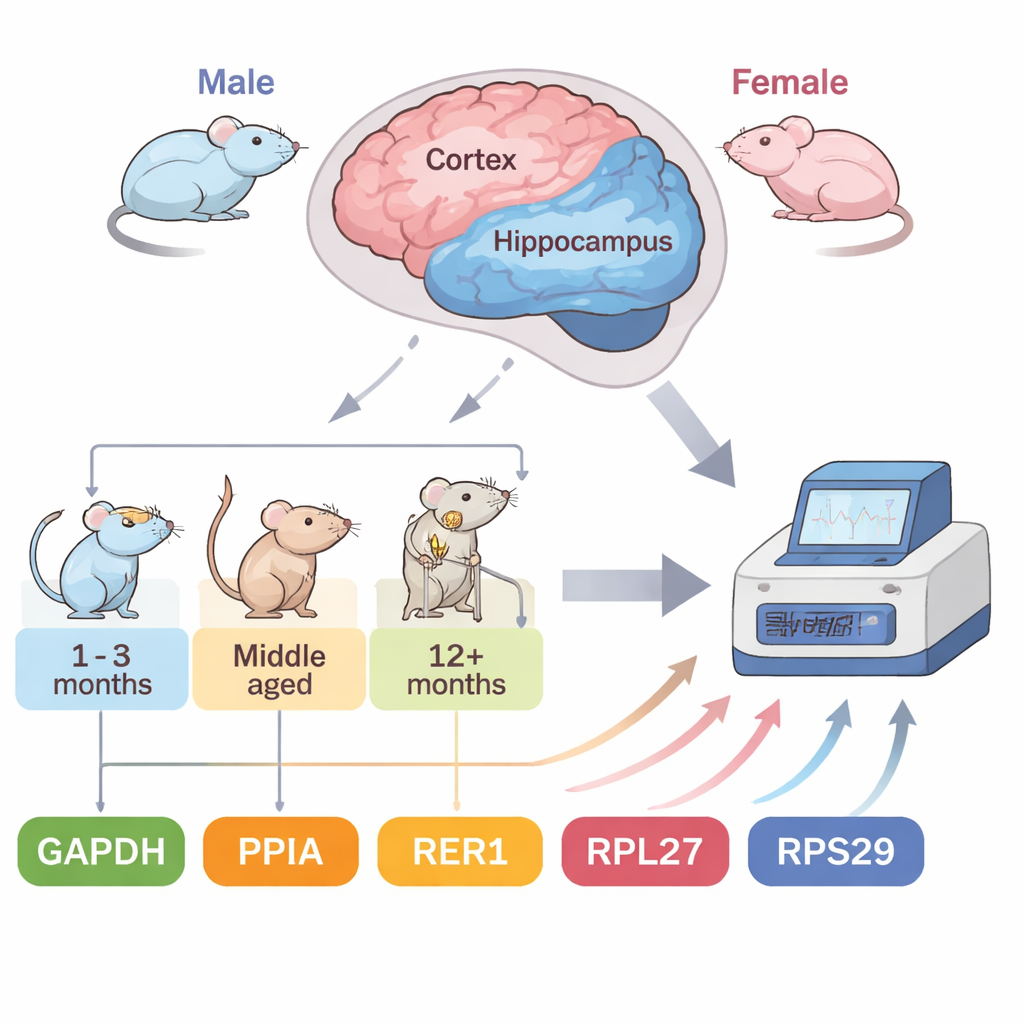
A fast-track mouse model of Alzheimer’s
The team focused on 5XFAD mice, a widely used model that carries five human mutations linked to familial Alzheimer’s disease. These animals develop amyloid plaques—protein clumps that are a hallmark of the disorder—remarkably early: at about two months of age, with heavy plaque buildup by four to five months. This accelerated timeline lets researchers track disease-related changes over months instead of years. The authors examined two vulnerable brain regions, the cortex and the hippocampus, in both male and female mice at four stages: early (2 months), developing (4 months), and more advanced (7 and 10 months). They tested five common reference genes and used four independent statistical tools to rank how stably each one behaved under these shifting conditions.
Reference genes are not one-size-fits-all
The results show that no single gene stayed perfectly steady everywhere. Instead, stability depended on where in the brain the sample came from, whether the mouse was male or female, and how far the disease had progressed. Some genes, like Gapdh—a classic workhorse in many studies—performed relatively well in several settings, especially in cortex at earlier ages, but showed more variability in later stages or in certain hippocampal samples. Other candidates, such as Rps29, Ppia, Rpl27 and Rer1, emerged as better partners in specific combinations, and these “best pairs” shifted with age and sex. Overall, the cortex tended to show more variability than the hippocampus, and later disease stages, particularly in male mice, disrupted the supposed stability of several reference genes.
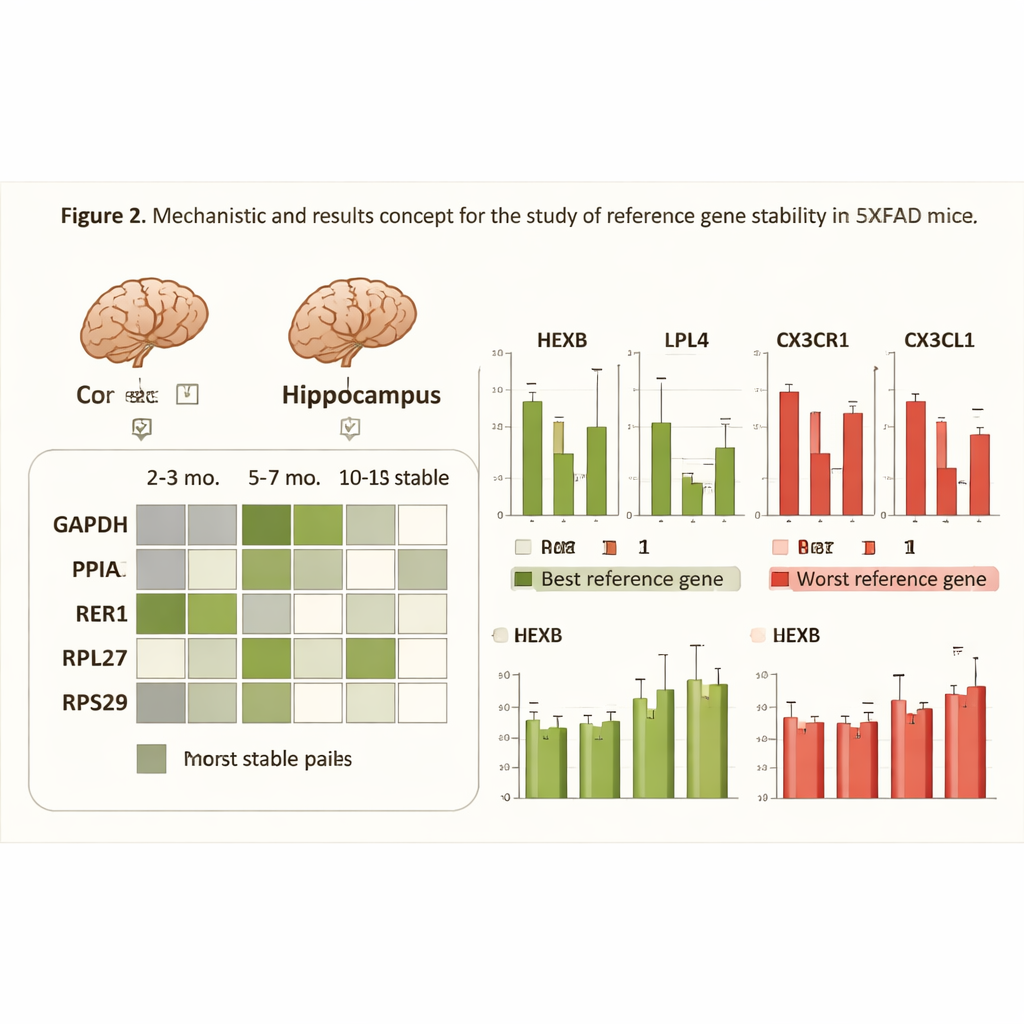
What happens when the ruler bends
To demonstrate why careful choice of reference genes matters, the researchers looked at four brain genes involved in inflammation, metabolism, and immune signaling—processes deeply implicated in Alzheimer’s disease. They compared how the apparent changes in these target genes differed when normalized with either the most stable reference pair, the least stable single gene, or all five genes together. When poor reference genes were used, the data became noisier and some biologically meaningful shifts appeared weaker or even statistically non-significant. In contrast, using the best-matched pair for each condition reduced scatter in the measurements and strengthened the detection of real differences between healthy and diseased mice, especially at later stages of pathology.
Clearer signals for a complex disease
For non-specialists, the takeaway is straightforward: even the basic yardsticks used in gene studies can warp under the pressure of a rapidly changing brain disease like Alzheimer’s. This work offers a practical guide to choosing better reference genes in an important mouse model and underscores that researchers must validate their “rulers” for each brain region, age, and sex they study. By doing so, scientists can obtain clearer, more trustworthy readouts of which genes truly change during Alzheimer’s-like degeneration—an essential step toward understanding the disease’s mechanisms and evaluating potential treatments.
Citation: Daini, E., Antonioni, K., Piemontese, M. et al. Reference gene variability across age and sex in 5XFAD mice highlights normalization challenges in Alzheimer’s models. Sci Rep 16, 7302 (2026). https://doi.org/10.1038/s41598-026-38346-5
Keywords: Alzheimer’s disease, gene expression, 5XFAD mice, reference genes, RT-qPCR