Clear Sky Science · en
Federated learning for heterogeneous electronic health record systems with cost effective participant selection
Why sharing hospital data is so hard
Modern hospitals collect enormous amounts of digital information about their patients, from lab tests and vital signs to medications and procedures. In theory, combining these records across many institutions should let doctors build smarter computer models that predict who is at risk and which treatments might help most. In practice, however, hospitals use different software systems, store data in incompatible formats, and must strictly protect patient privacy and budgets. This study explores how to let hospitals learn from each other’s data without copying it or overspending.
Training together without sharing raw records
The authors build on an approach called federated learning, where each hospital trains a local model on its own patient records and then shares only model updates, not raw data. A central “host” hospital coordinates this process and aims to improve a prediction model for its own needs, such as forecasting complications in intensive care. Other hospitals, called subjects, participate in exchange for compensation. This setup avoids moving sensitive records across institutions, but it raises two tough questions: how to work with many different record systems, and how to avoid paying for partners that do not actually help the model.
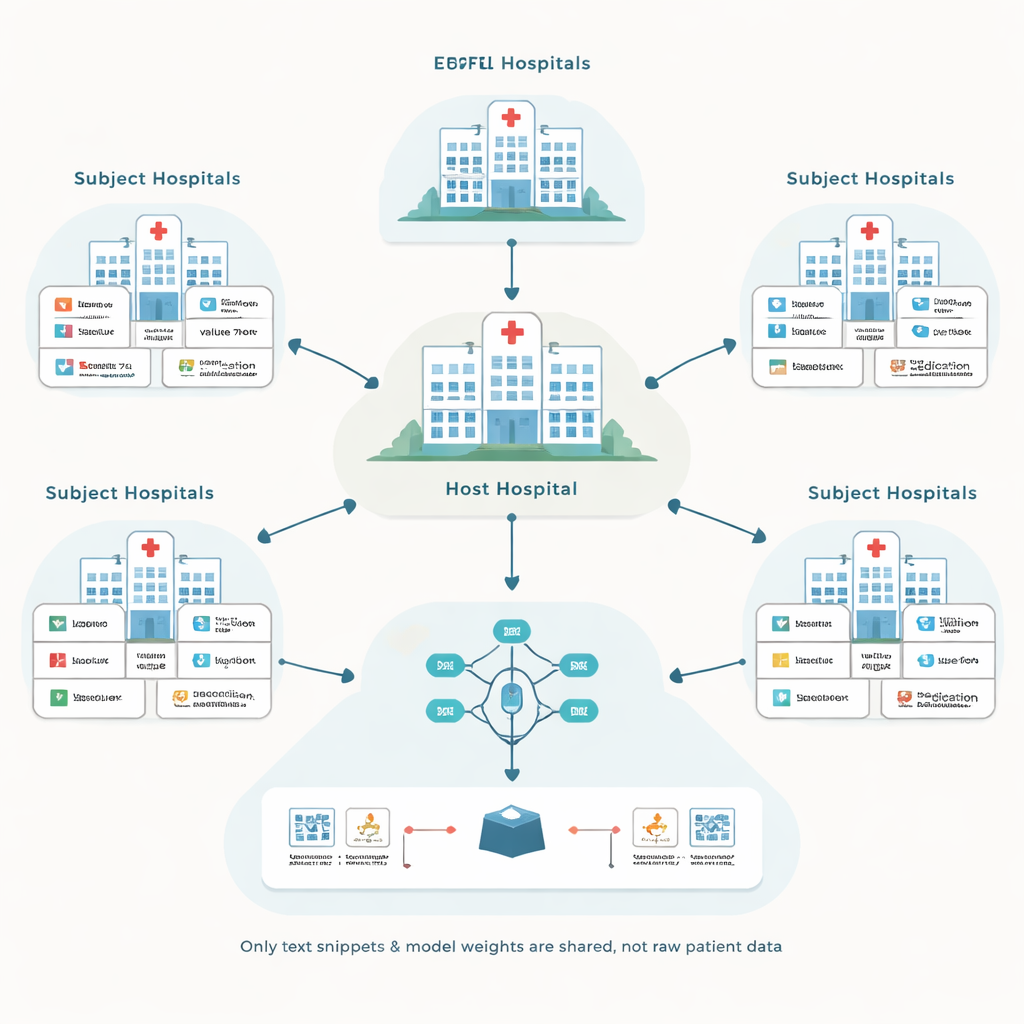
Turning messy records into shared language
Electronic health record systems differ widely in how they label and code information. One hospital might store a blood sugar test under a numeric code, while another uses a different code for the same test. Traditional solutions try to convert everything into a single, carefully designed standard database, which is expensive and requires many expert hours. Instead, the proposed framework, called EHRFL, converts each medical event into a short piece of text. For example, a lab entry like a glucose measurement becomes a phrase such as “lab event glucose value 70 mg/dL.” Because each hospital already keeps dictionaries that map local codes to human-readable names, this conversion can be automated without custom hand-tuning.
Building patient profiles from text
Once events are written as text, EHRFL uses modern language-processing models to turn each event into a numeric vector, and then combines many events into a single “patient embedding” – a compact summary of that person’s medical history over a time window. These embeddings feed into a prediction layer that tackles several clinical tasks at once, such as predicting death in the hospital or kidney injury after an intensive care admission. The authors run federated training on five large, real-world critical-care datasets that span different hospitals, time periods, and record systems. Across a range of algorithms, including commonly used federated methods, models trained with this text-based approach consistently outperform models trained on a single hospital alone, even though the underlying data formats differ.
Picking the right partners while protecting privacy
More partner hospitals do not always mean better results. Some institutions have patient populations or record patterns so different from the host that including them can slow training or slightly hurt performance, while still adding cost. To address this, the authors propose a selection step based on similarity between hospitals’ patient embeddings. The host first trains a model on its own data, shares the model weights, and each candidate hospital uses them to compute patient embeddings. To protect privacy, each subject clips extreme values from its embeddings, averages them into a single vector, and then adds carefully calibrated random noise before sending only this noisy average to the host. The host compares its own average with each subject’s using simple similarity measures and chooses only the most similar hospitals to join the full federated run.
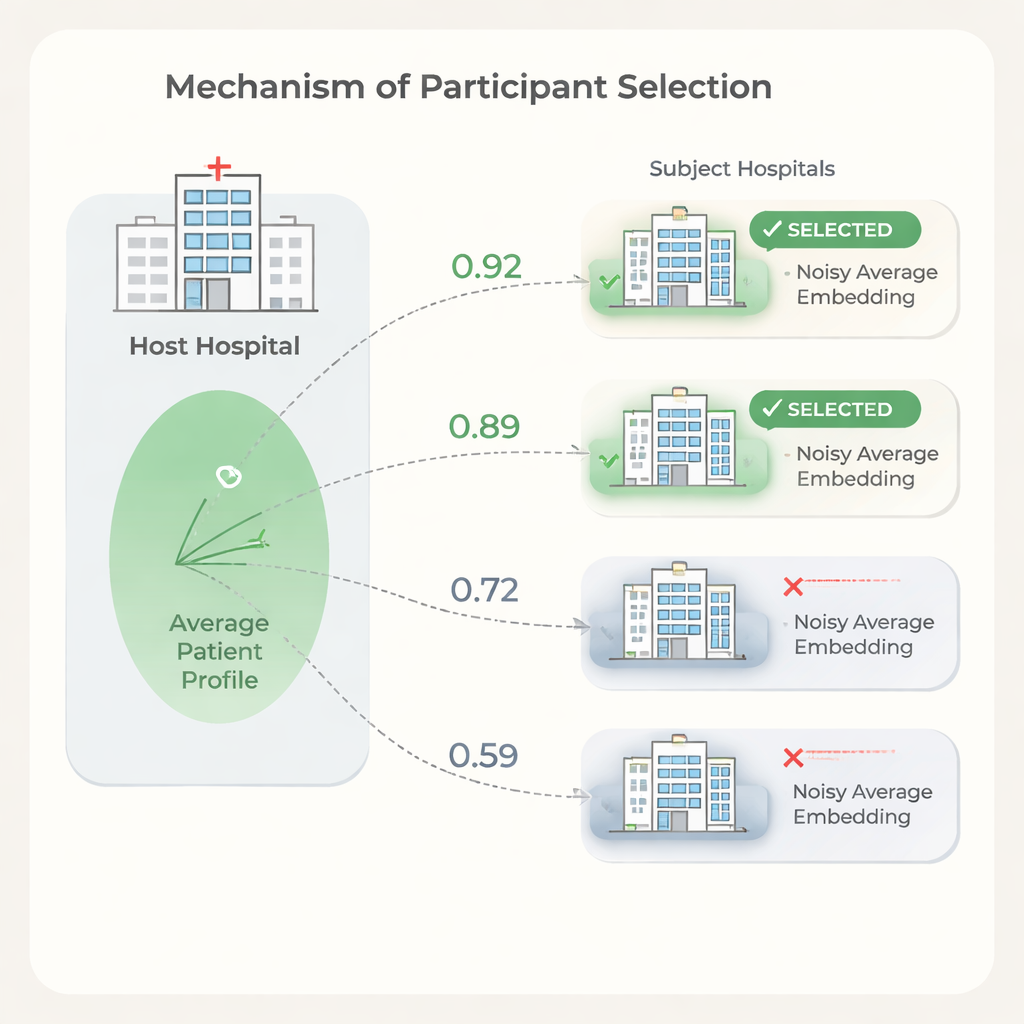
Saving money without losing accuracy
Experiments show that similarity between hospitals’ average patient embeddings aligns with how much each hospital helps or harms the host’s prediction performance. Using this signal to select partners, the host can drop low-similarity hospitals while keeping or even improving prediction quality compared with using all available sites. The authors also outline a cost model showing that, because data usage fees and training time scale with the number of participating hospitals, even modest reductions in partners can lead to substantial savings. At the same time, the selection step is lightweight: the model is trained once, and each hospital only performs simple calculations on a single averaged vector.
What this means for future healthcare AI
For readers outside the field, the key message is that it may be possible for hospitals to “learn together” without pooling raw patient records, and to do so in a way that respects both privacy and financial limits. By translating diverse records into a shared text form and then using privacy-preserving summaries of patient populations to choose compatible partners, EHRFL offers a practical recipe for building hospital-specific prediction tools. While the study focuses on intensive care data, the same ideas could extend to outpatient clinics, emergency rooms, and even non-medical domains where organizations want to collaborate on better models without giving up control of their data.
Citation: Kim, J., Kim, J., Hur, K. et al. Federated learning for heterogeneous electronic health record systems with cost effective participant selection. Sci Rep 16, 6876 (2026). https://doi.org/10.1038/s41598-026-38299-9
Keywords: federated learning, electronic health records, patient privacy, clinical prediction, healthcare AI