Clear Sky Science · en
No-rank tensor decomposition via metric learning
Finding Patterns in a Sea of Data
Modern science is drowning in complex data: stacks of medical scans, brain activity maps, astronomical images, and simulations of materials. Making sense of this data often means squeezing it into simpler forms without losing what really matters. This paper introduces a new way to do that. Instead of trying to faithfully rebuild every pixel, it focuses on capturing the true relationships between samples – which brain looks more like which, which galaxy shape resembles which – so that the resulting map of the data reflects meaning rather than raw detail.
From Rebuilding Images to Measuring Similarity
Traditional tools for simplifying multi-dimensional data, known as tensor decompositions, work a bit like breaking a chord into notes. They factor a data “block” into a small number of basic patterns plus weights. To do this, they must be told in advance how many patterns – the “rank” – they are allowed to use, and they judge success by how well the original data can be reconstructed. That is ideal for compression or denoising, but not necessarily for tasks like “are these two faces the same person?” or “does this brain scan belong to an autistic or a typical subject?” where correct grouping matters more than perfect reconstruction.
In parallel, deep learning has popularized another idea: instead of decomposing a tensor algebraically, learn a compact numerical code, or embedding, through a neural network. Classic autoencoders still focus on reconstruction. This work flips the objective. It proposes a “no-rank” framework that does not fix a rank ahead of time and does not care about pixel-perfect recovery. Instead, it learns a distance measure so that points that ought to be close (same person, same diagnosis, same physical class) end up neighbors in the embedding space, and points that ought to be different are pushed apart.
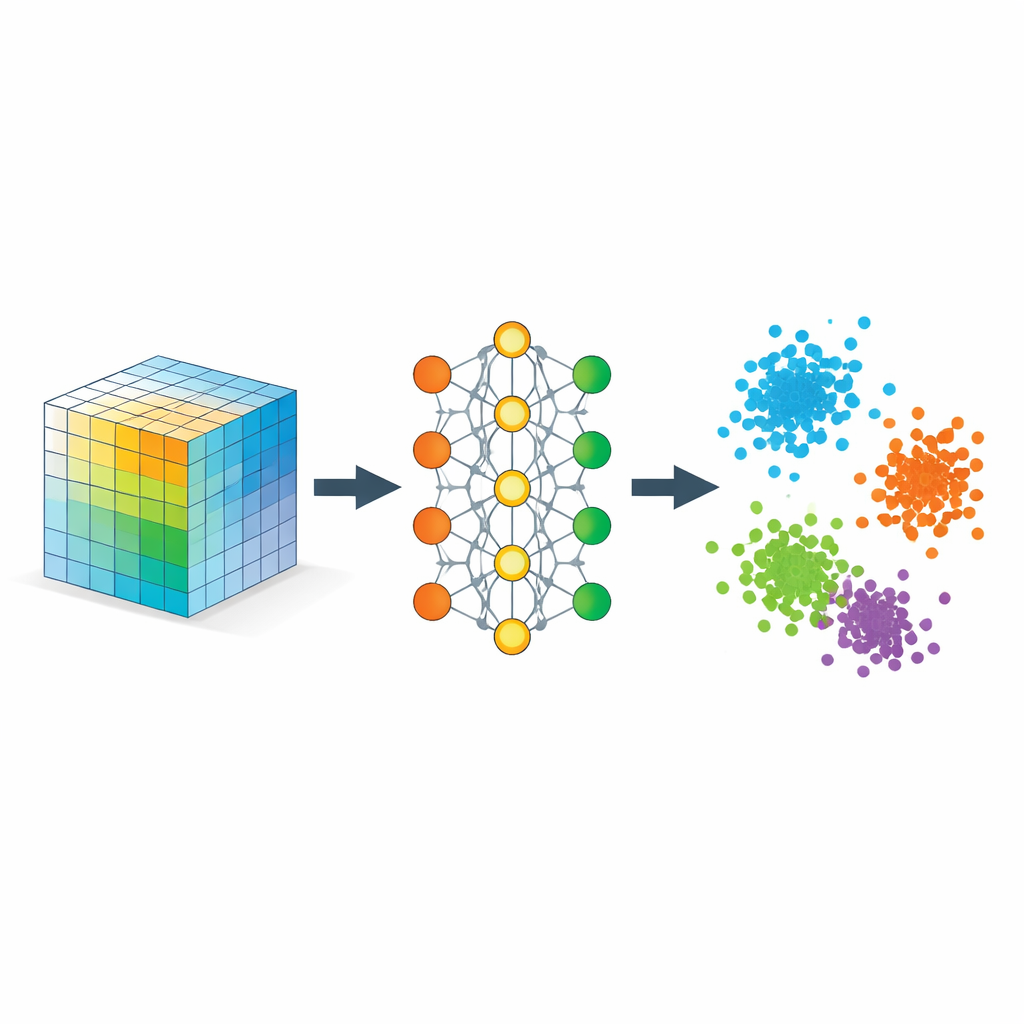
Teaching the Network What “Close” Should Mean
The key ingredient is a strategy called metric learning, implemented here through triplets of examples: an anchor sample, a positive sample of the same type, and a negative sample of a different type. During training, the network is rewarded when the anchor is closer to the positive than to the negative by a safety margin. Over many such triplets, this simple rule sculpts the embedding space so that distances mirror semantic similarity rather than raw pixel similarity. Additional regularizers encourage the network to spread information evenly across dimensions, avoid collapsing everything into a line, and keep local neighborhoods roughly intact, so that nearby points in the original data remain nearby when embedded.
Mathematically, the authors show that this embedding behaves like a flexible tensor decomposition, but without a pre-set rank. The learned coordinates can be interpreted as factors in a classic decomposition of a similarity tensor whose entries measure how strongly different parts of the data align. Because the model penalizes redundant directions, it tends to use all embedding dimensions effectively, letting the data itself determine how many meaningful components are needed. At the same time, they provide theoretical guarantees that standard training procedures converge and that the resulting geometry faithfully separates classes while not wildly distorting meaningful local relationships.
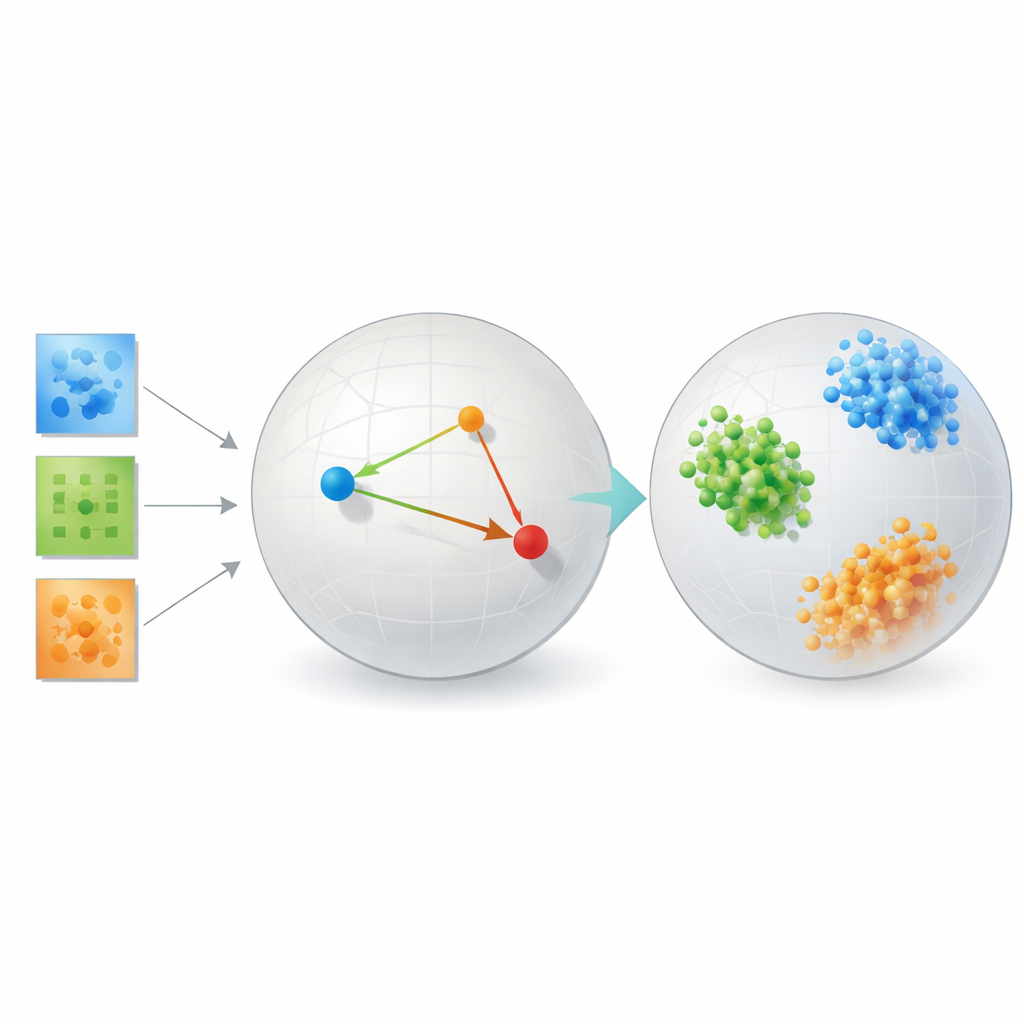
Putting the Method to the Test
To show that their approach is not just elegant theory, the author tests it on several very different problems. In face recognition benchmarks, the learned embeddings group images of the same person into tight, well-separated clusters, dramatically outperforming classic methods such as principal components, popular visualization tools like t-SNE and UMAP, and traditional tensor decompositions that rely on fixed ranks. On brain connectivity data from people with and without autism, the method discovers a space where the two groups are more cleanly separated than with reconstruction-focused tensor tools or autoencoding neural networks, hinting that it is honing in on clinically relevant patterns in how brain regions interact.
The study also includes controlled simulations of galaxy shapes and crystal structures, where the “true” categories are known exactly. Here, the metric learning framework nearly perfectly clusters the synthetic galaxies and crystals by their underlying physical types. Across all these settings, the method consistently trades some faithfulness to the original pixel layout for a representation in which similarity and difference line up with scientific meaning. Importantly, it does this without the vast data and computing resources often needed to train transformer-based deep models, which struggled in these relatively small scientific datasets.
Why This Matters for Future Scientific Data
For scientists seeking patterns in limited, high-dimensional data, this work offers an appealing shift in perspective. Instead of guessing a rank and optimizing for reconstruction, researchers can directly ask for an embedding that reflects the relationships they care about: same diagnosis, same material phase, same astrophysical class. The proposed no-rank metric learning framework shows that such embeddings can be both interpretable and powerful, especially when data are scarce. As the author notes, challenges remain – including dealing with class imbalance and scaling to many categories – but the message is clear: in many scientific problems, learning a good notion of similarity may be more valuable than rebuilding every detail of the original signal.
Citation: Bagherian, M. No-rank tensor decomposition via metric learning. Sci Rep 16, 8326 (2026). https://doi.org/10.1038/s41598-026-38221-3
Keywords: metric learning, tensor decomposition, representation learning, dimensionality reduction, scientific data analysis