Clear Sky Science · en
Single-cell transcriptomics identifies fibroblast associated immune heterogeneity and prognostic signatures in bladder cancer
Why the cells around a tumor matter
Bladder cancer is common and often comes back or spreads despite surgery and modern drugs. Doctors know that a tumor does not grow alone; it is surrounded by normal-looking cells that can quietly help or hinder the cancer. Among these, a group called fibroblasts help build the tissue scaffold around organs. This study asks a simple but powerful question: can a closer look at individual fibroblasts in and around bladder tumors reveal who is likely to live longer, and point to new ways to treat the disease?
Taking a tumor apart cell by cell
The researchers used single-cell RNA sequencing, a technique that reads out which genes are switched on in thousands of individual cells from tissue next to bladder tumors. Instead of viewing the tumor region as one blurred mass, this method separates it into many distinct cell types. By doing so, the team identified 21 groups of cells, including tumor cells, immune cells, blood vessel cells, and, importantly, over 3,600 fibroblasts. Each group carried its own gene “barcode,” allowing the scientists to map where the cells sit in the tissue and how they differ from one another.
Fibroblasts as signal hubs
Once fibroblasts were singled out, the team explored how these cells talk to their neighbors and what controls their behavior. Using computational tools, they pinpointed key control genes, known as transcription factors, that act like master switches inside fibroblasts. Three such switches—MAF, TWIST1, and TCF21—stood out. They are linked to how cells change shape, remodel the tissue scaffold, and respond to immune signals. Additional analyses of cell-to-cell communication suggested that fibroblasts send and receive many chemical messages, forming a busy crossroads that can shape whether the immune system attacks or tolerates the tumor.
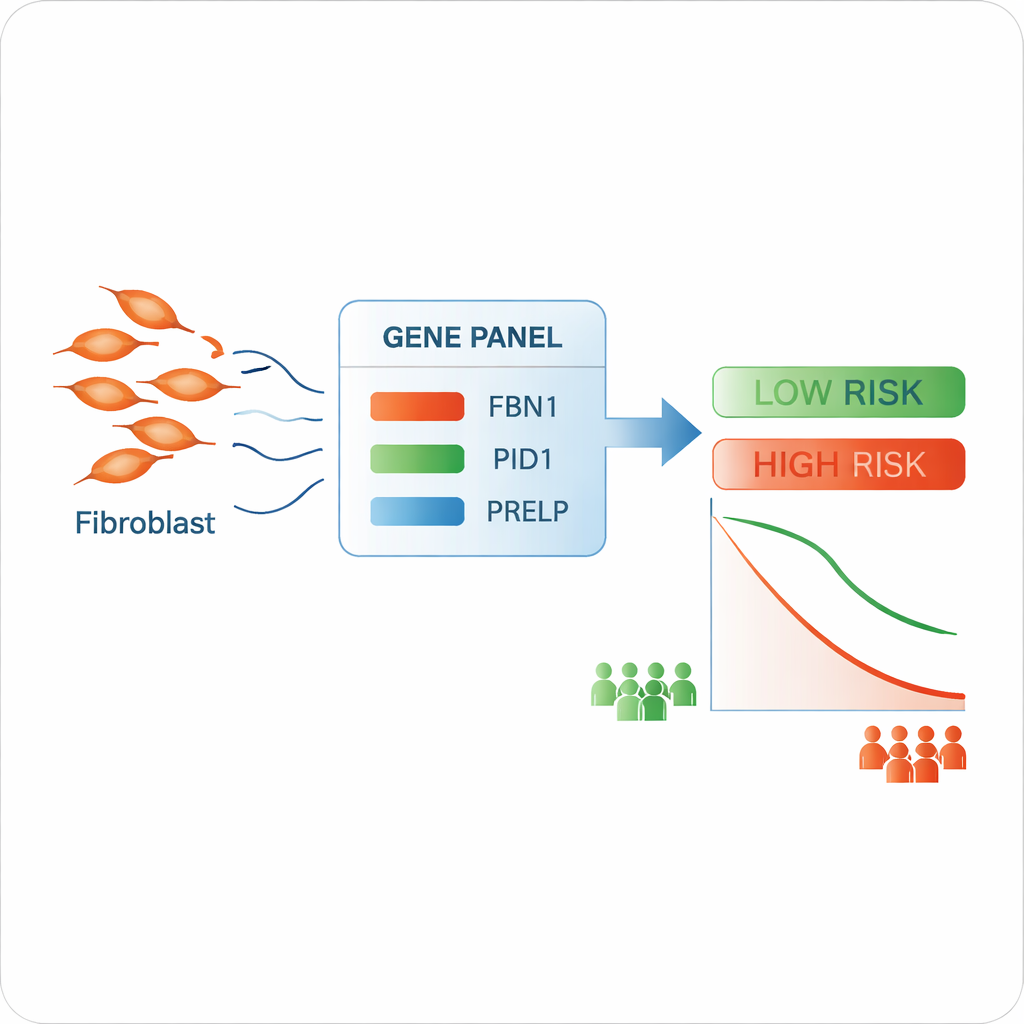
From fibroblast genes to a risk score
The scientists next asked whether fibroblast-linked genes could help predict patient survival. They combined the single-cell findings with large, publicly available gene data from hundreds of bladder cancer patients. Among many candidates, they focused on three genes tied to the tissue scaffold—FBN1, PID1, and PRELP. These genes were used to build a simple risk score: higher combined activity of the three genes in a tumor was associated with poorer overall survival. Patients could be split into high-risk and low-risk groups with clearly different survival curves, and the model showed reasonable accuracy in predicting who would live at least five years after diagnosis.
What metabolism and mutations add to the picture
Beyond gene switches and signals, the study also examined how fibroblasts handle energy and nutrients. A specialized analysis suggested that fibroblasts in the tumor region process fuel in ways that support growth, such as converting sugars into building blocks for fats and complex sugars used in the tissue matrix. These shifts may help create a physical and chemical environment that shelters the tumor. The team also looked at DNA mutations across patients and found that those in the high-risk group tended to have a lower overall number of mutations, a pattern that may relate to how well such tumors respond to modern immunotherapies.
How this could help patients in the future
For non-specialists, the key message is that the “soil” around a bladder tumor—the fibroblasts and the tissue they build—can be just as important as the cancer cells themselves. By reading the activity of a small set of fibroblast-related genes, doctors may one day better estimate a patient’s outlook and choose treatments more wisely. Although the findings need to be tested in larger and more diverse patient groups, this work shows that zooming in on single cells can uncover hidden players in cancer and open new paths for therapies that target not only the tumor, but also the supportive cells that help it thrive.
Citation: Tang, X., Liu, L., Gao, M. et al. Single-cell transcriptomics identifies fibroblast associated immune heterogeneity and prognostic signatures in bladder cancer. Sci Rep 16, 7151 (2026). https://doi.org/10.1038/s41598-026-38219-x
Keywords: bladder cancer, cancer-associated fibroblasts, tumor microenvironment, single-cell sequencing, prognostic biomarkers