Clear Sky Science · en
Identification and validation of a novel ferroptosis-related gene signature associated with inherited retinal degeneration in Rd10 mice
Why dying eye cells matter for all of us
Retinitis pigmentosa is a group of inherited eye diseases that gradually steal people’s sight, often beginning with night blindness and ending in complete vision loss. There is no cure, and existing gene therapies can only help a small fraction of patients. This study explores a different angle: a special form of cell death called ferroptosis, driven by iron and oxidative damage, to uncover new gene targets that might one day protect light-sensing cells in the retina—no matter which mutation a person was born with.
A closer look at a blinding eye disease
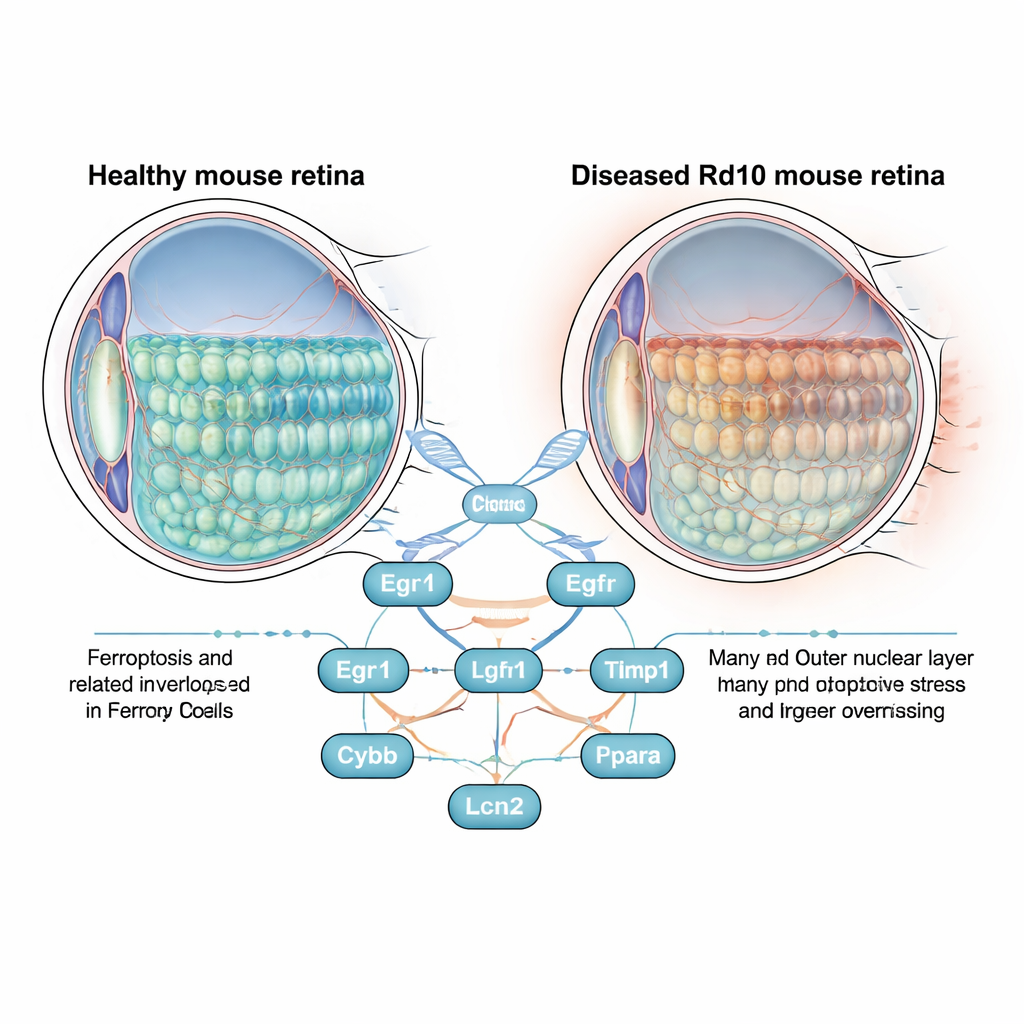
The retina is a thin layer of tissue at the back of the eye that houses rod and cone photoreceptors, the cells that convert light into electrical signals. In retinitis pigmentosa, these cells slowly die, first robbing people of night and side vision and later central sight. Scientists often study this process in Rd10 mice, which carry a mutation that closely mimics the human disease. Instead of focusing solely on the faulty gene, the authors asked a broader question: as the retina degenerates, which genes linked to ferroptosis turn on or off, and could they be driving cell loss?
Finding the key genes in a sea of data
The team mined large public gene-expression datasets from healthy and Rd10 mouse retinas, essentially taking a global “snapshot” of which genes were more or less active. They found over 2,000 genes that changed, then cross-checked these with a curated list of ferroptosis-related genes. This narrowed the field to 37 candidates tied to iron-driven, oxidative cell death. Computer analyses showed these genes clustered in processes like response to oxidative stress, regulation of neuron death, and fat metabolism pathways—hallmarks of ferroptosis and known stress responses in degenerating nerves.
Eight genes that stand out
To pinpoint the most influential players, the researchers built a protein interaction network and applied several ranking algorithms. Eight “hub” genes emerged: Egr1, Cd44, Egfr, Tlr4, Timp1, Cybb, Lcn2, and Ppara. Seven of them were more active in diseased retinas, while Ppara was less active. The group confirmed these patterns in an independent dataset and then looked more closely at where the top genes appeared in the eye. Using fluorescent staining and single-cell RNA sequencing, they showed that Egr1 is strongly increased in rod and cone photoreceptors, while Cd44 is sharply elevated in Müller cells, the supportive glial cells that span the retina. This suggests that stressed photoreceptors and their support cells may cooperate—perhaps harmfully—through ferroptosis-linked pathways.
What this means for future treatments
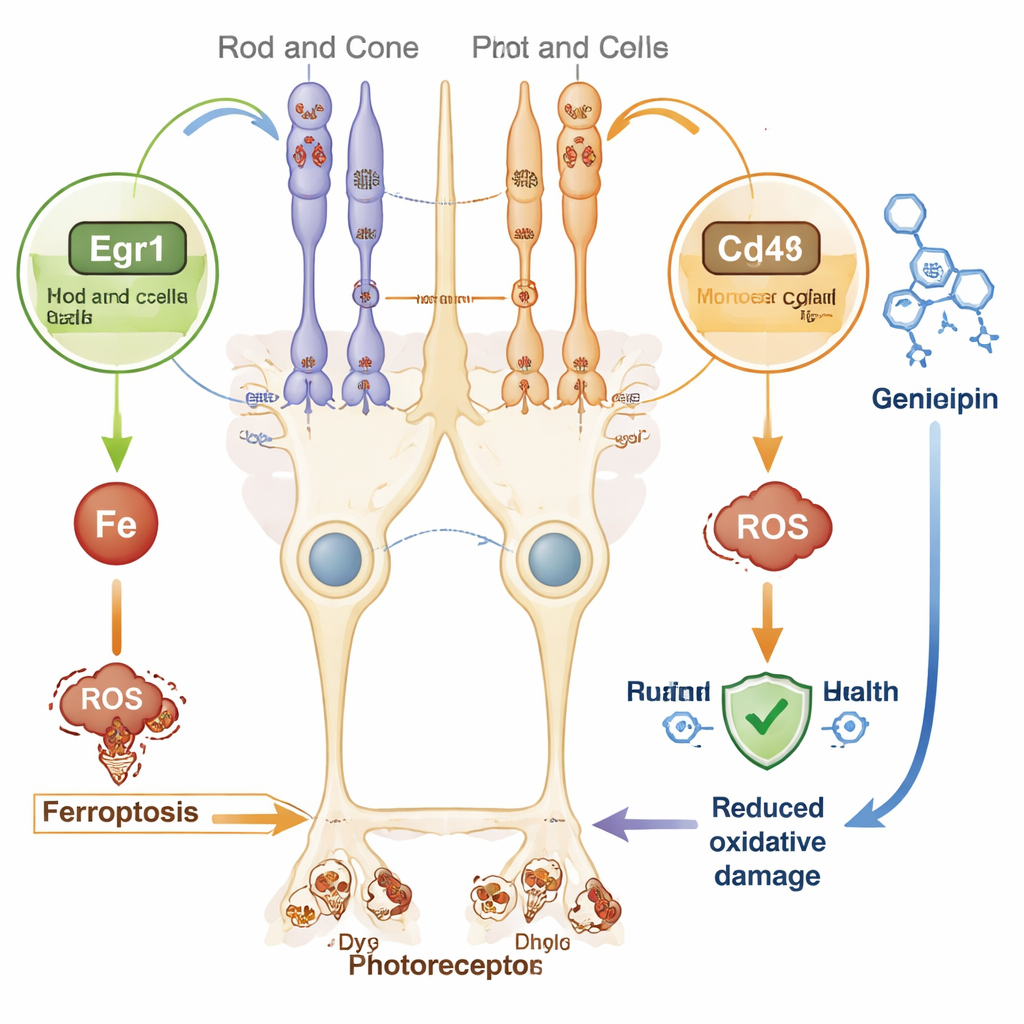
Because gene therapy tailored to each faulty gene is complex and expensive, broad-acting strategies are attractive. The newly identified gene signature hints at several such avenues. For instance, Egr1 is a fast-acting switch that responds to oxidative stress and can promote ferroptosis in other organs. Cd44, a surface molecule on Müller cells, has been shown in different tissues to worsen iron overload and cell death. At the same time, decreased Ppara, a regulator of fat metabolism and energy use, may leave retinal cells more vulnerable to damage. By targeting these shared stress pathways, it may be possible to slow retinal degeneration across many genetic forms of retinitis pigmentosa.
Drugs that might help protect sight
Using a drug–gene interaction database, the authors searched for compounds that could influence Egr1 or Cd44. Several existing molecules surfaced, including the natural compound Genipin, which has shown protective effects in models of Parkinson’s disease, nerve degeneration, and diabetic retinopathy by dampening oxidative stress and inflammation. While none of these candidate drugs has yet been tested in this specific mouse model of retinitis pigmentosa, they provide promising starting points for future laboratory and animal studies aimed at shielding photoreceptors from ferroptosis.
Take‑home message for non‑specialists
This work shows that iron-driven, oxidative cell death is closely intertwined with the gene activity changes seen in a classic model of inherited blindness. By singling out eight key genes, and especially the stress-responsive Egr1 in photoreceptors and Cd44 in Müller cells, the study maps new territory for potential therapies that do not depend on fixing a single mutated gene. If future experiments confirm that calming ferroptosis through these targets can keep retinal cells alive, it could open the door to treatments that benefit a wide range of patients facing progressive vision loss.
Citation: Qiu, X., Fu, XW., Lei, XL. et al. Identification and validation of a novel ferroptosis-related gene signature associated with inherited retinal degeneration in Rd10 mice. Sci Rep 16, 6992 (2026). https://doi.org/10.1038/s41598-026-38193-4
Keywords: retinitis pigmentosa, ferroptosis, photoreceptors, retinal degeneration, Egr1 Cd44