Clear Sky Science · en
RNA sequencing reveals differential expression of circular RNAs in human small cell lung cancer
Why tiny RNA loops could change lung cancer care
Small cell lung cancer is one of the deadliest cancers, spreading quickly and often resisting chemotherapy. Doctors still lack good tests to catch it early or predict who will respond to treatment. This study looks at an unusual class of genetic molecules called circular RNAs—tiny RNA loops that are surprisingly stable in our cells—to see whether they help drive this aggressive cancer and whether they might serve as new, more reliable biomarkers.
Understanding the most aggressive lung cancer
Small cell lung cancer (SCLC) makes up only about one in five lung cancer cases, but it causes a disproportionate share of deaths because it grows fast, spreads early, and usually returns after chemotherapy. Most patients are diagnosed at an advanced stage, when surgery is not an option and tissue samples are limited. While researchers have mapped many of the DNA mutations in SCLC, they know far less about how non‑coding RNAs—molecules that do not make proteins but still strongly influence cell behavior—shape the disease. Circular RNAs (circRNAs) are a particularly intriguing group, because their closed loop structure makes them more resistant to breakdown than ordinary RNAs, raising hopes that they could be measured reliably in tissue or even blood.
Hunting for RNA loops in tumor tissue
The research team in India collected tumor biopsies from patients with newly diagnosed SCLC and compared them with normal lung tissue from people undergoing surgery for early‑stage non‑small cell lung cancer. Using high‑throughput RNA sequencing, they scanned more than 26,000 junctions where circular RNAs can form, then filtered and analyzed the data to identify which circRNAs were truly different between cancerous and non‑cancerous samples. They found 23 circular RNAs whose activity changed significantly in SCLC: 13 were more abundant and 10 were less abundant in tumors than in normal lung tissue. These molecules arose from 21 different genes, hinting at a broad rewiring of RNA regulation in SCLC.
Building a three‑layer RNA control network
Circular RNAs often act like molecular sponges, soaking up microRNAs—short RNA snippets that normally damp down specific genes. To map this control network, the researchers used public databases to predict which microRNAs could bind each of the 23 altered circRNAs, and which genes those microRNAs were likely to regulate. The result was a dense three‑layer web linking 23 circular RNAs to 241 microRNAs and about 7,800 protein‑coding genes. When they examined what these genes do, many clustered in cell‑cycle control, protein breakdown, stress responses, and well‑known cancer pathways such as p53, MAPK, Hippo, and cellular aging. In other words, the misregulated RNA loops appear to sit at the crossroads of key growth and survival circuits in SCLC cells.
Two standout RNA loops with diagnostic promise
Among all the altered circRNAs, two stood out. One, called circLIFR, was consistently reduced in SCLC tissue and in several SCLC cell lines compared with normal lung cells. Its partner microRNAs, including miR‑1234‑3p and miR‑375‑3p, were strongly increased, which fits the idea that losing the circRNA sponge leaves more free microRNAs to influence cancer‑linked genes. The other loop, circCAMSAP1, showed the opposite pattern: it was elevated in tumors, while one of its reported microRNA partners, miR‑145‑5p, was sharply decreased. Patients whose cancer worsened despite first‑line chemotherapy tended to have higher circCAMSAP1 levels, suggesting a link to treatment resistance. When the team tested how well these molecules distinguish SCLC tissue from normal lung, circLIFR performed especially well, with an accuracy measure (AUC) above 0.9, while circCAMSAP1 and the associated microRNAs showed moderate but still useful diagnostic power.
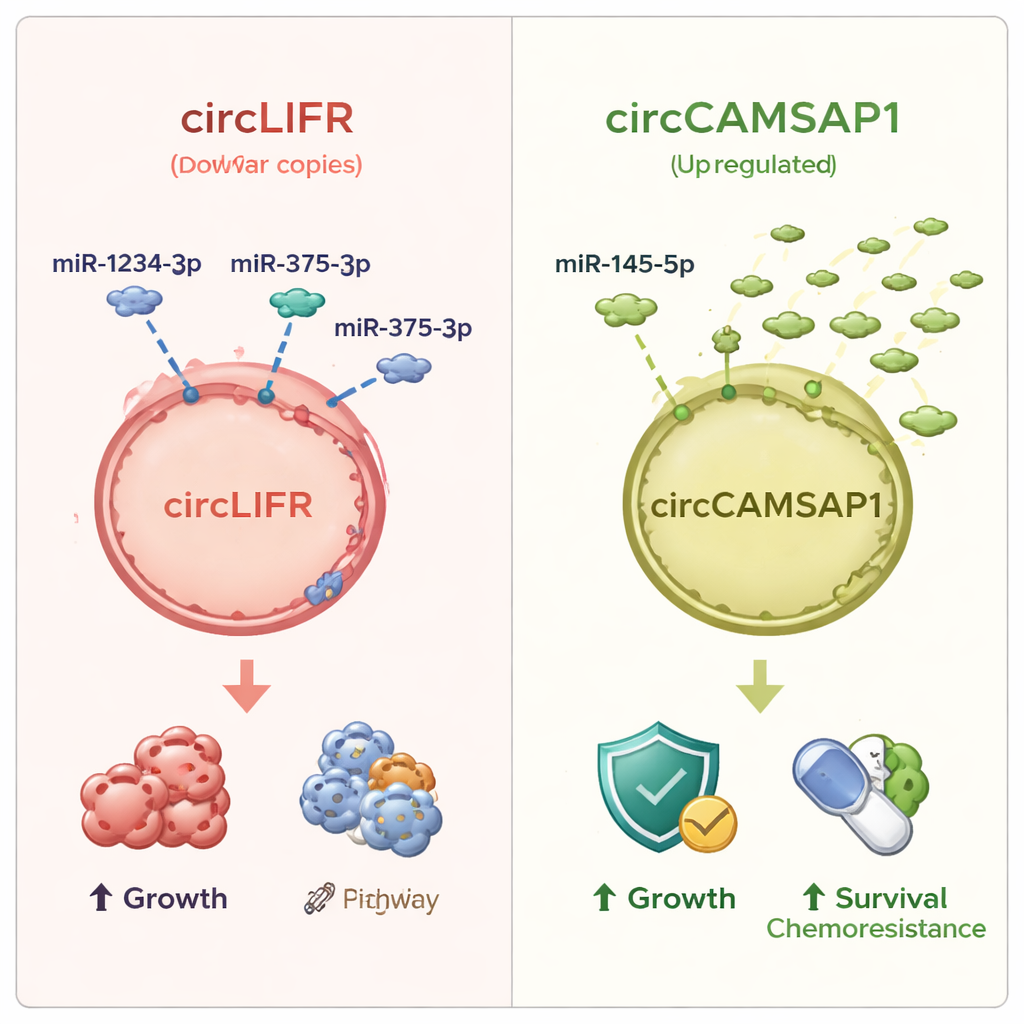
What this could mean for patients
For a layperson, the takeaway is that this study uncovers a new layer of control in small cell lung cancer—built from tiny RNA loops and their partners—that helps regulate how quickly tumor cells grow, how they respond to stress, and potentially how they withstand chemotherapy. Two specific circular RNAs, circLIFR and circCAMSAP1, emerge as especially promising markers: one tends to be lost in tumors, the other gained, and together with their associated microRNAs they can help tell cancerous from normal tissue and may hint at how a patient’s disease will respond to treatment. While these findings still need laboratory experiments to prove cause‑and‑effect and larger clinical studies to test real‑world usefulness, they point toward future blood or tissue tests based on circular RNAs that could improve the diagnosis and management of one of the most challenging forms of lung cancer.
Citation: Saxena, V., Abhilash, D., Budhraja, A. et al. RNA sequencing reveals differential expression of circular RNAs in human small cell lung cancer. Sci Rep 16, 7134 (2026). https://doi.org/10.1038/s41598-026-38145-y
Keywords: small cell lung cancer, circular RNA, microRNA, cancer biomarkers, RNA sequencing