Clear Sky Science · en
A frequency-spatial dual perception network for efficient and accurate medical image segmentation
Sharper Computer Eyes for Medical Scans
When doctors look at a skin spot, a breast ultrasound, or a CT scan, they are really asking one hard question: where exactly is the disease, and where is healthy tissue? The answer often comes from software that outlines suspicious regions in each image, a process called segmentation. This paper introduces a new artificial intelligence system, FDE-Net, that draws those outlines more accurately while using reasonable computing power, making it more suitable for real-world hospital use.
Why Standard Tools Miss the Small Stuff
Most current medical imaging tools rely on "U-shaped" neural networks, such as the well-known U-Net, which compress an image to extract meaning and then expand it back to draw a mask of the target region. These networks are good at capturing sharp edges and textures, but they tend to treat every part of the image the same way when shrinking it down. As a result, faint or tiny lesions can disappear in the process, especially when they blend into complex backgrounds like surrounding organs or tissue. Existing methods also work mostly in the raw pixel space of the image, ignoring a complementary view: how image content is distributed across different frequencies, from broad smooth shapes to fine details.
Listening to Images in Different “Tones”
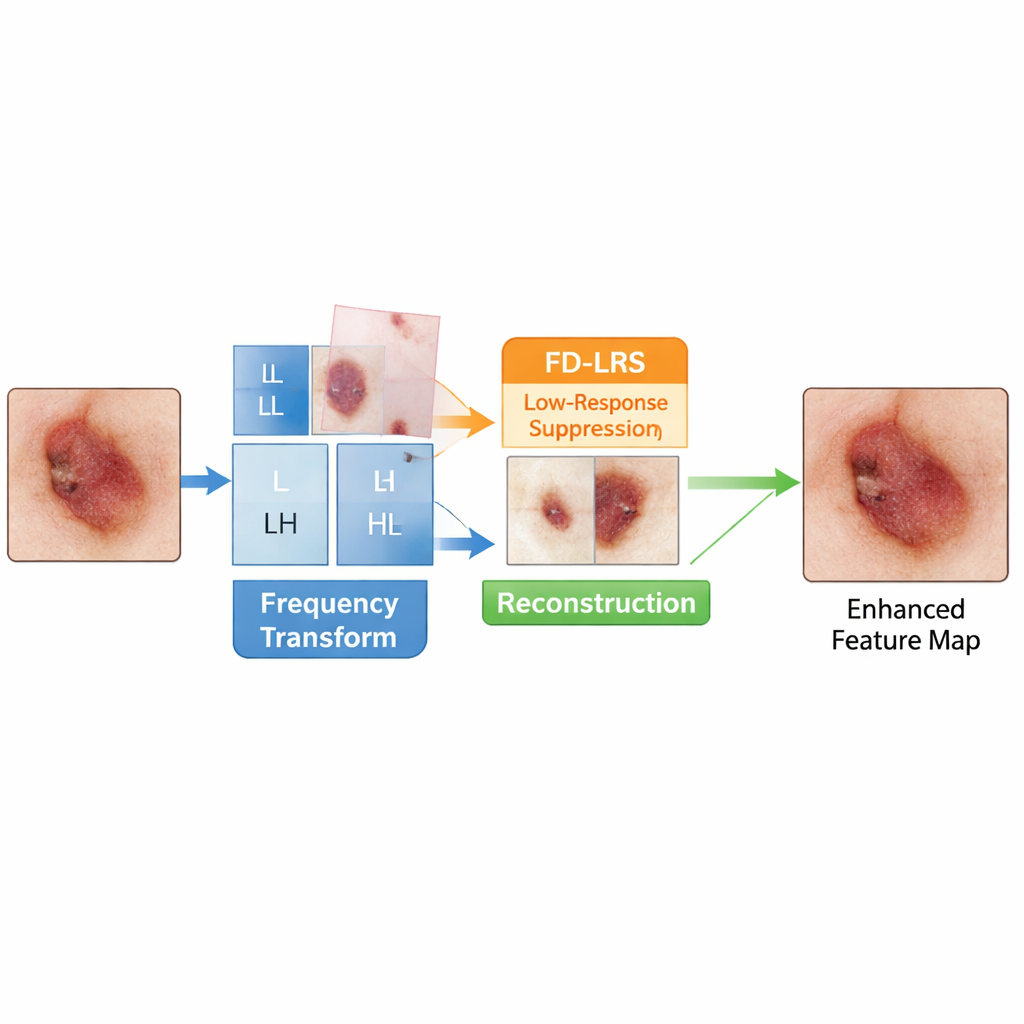
FDE-Net starts by treating a medical image a bit like an audio signal: it separates the picture into low-frequency parts that describe overall structure and high-frequency parts that capture edges and fine detail. Its Low-Frequency Information Extraction Block focuses on the low-frequency piece, which carries crucial cues about the shape and location of organs and lesions but is often polluted by background tissue. A dedicated module, called Frequency Domain Low-Response Area Suppression, learns to tone down low-frequency regions that look like uninformative background while amplifying regions more likely to contain disease. The network then recombines these cleaned-up low- and high-frequency components, giving later layers a clearer, more focused view of what matters.
Seeing Both the Big Picture and Tiny Lesions
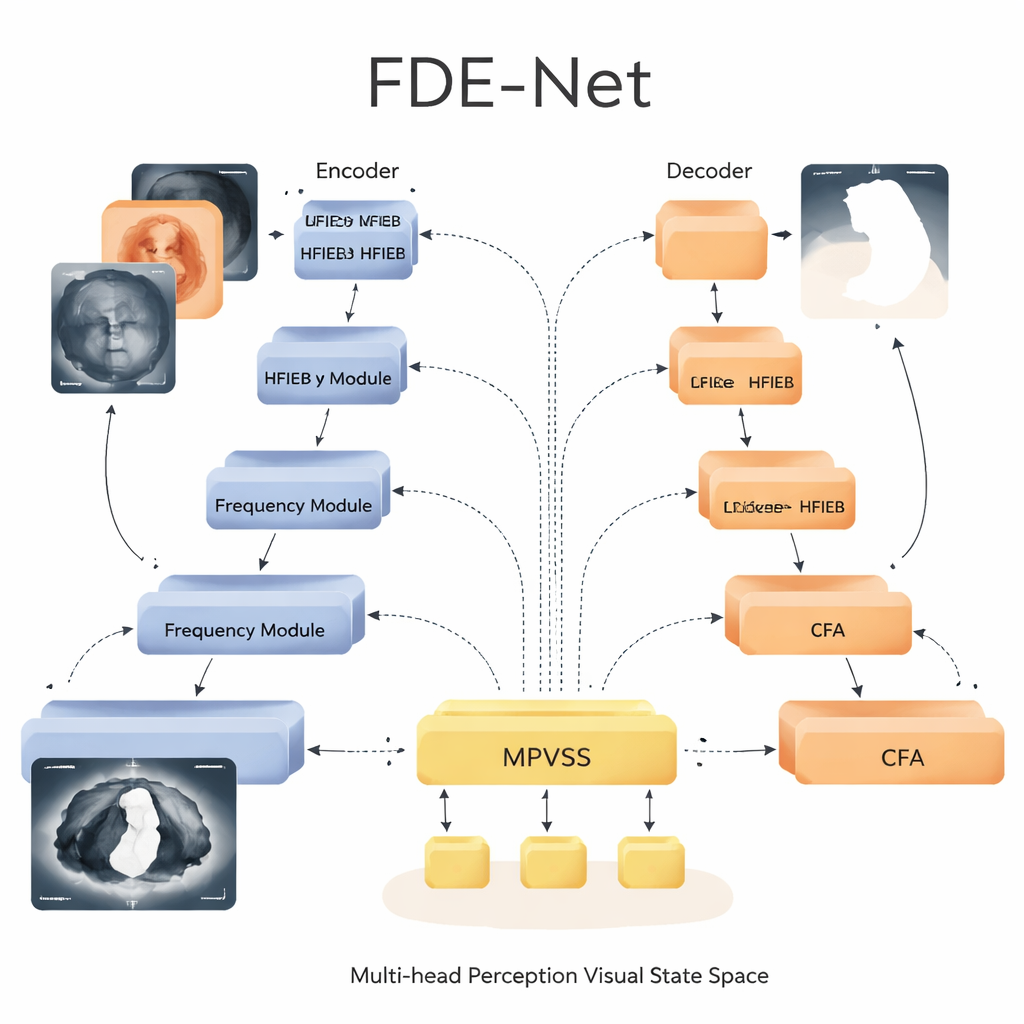
In the center “bottleneck” of the U-shaped architecture, FDE-Net uses a Multi-head Perception Visual State Space module. Instead of relying on heavy Transformer-style attention, which can be very costly for large medical images, this module belongs to a newer family of models known as state space models. It processes information efficiently while still capturing long-range relationships across the image. FDE-Net sends the features through several parallel branches that each look at the image at different scales, from small patches suited to pinpoint tiny spots to broad views capturing large organs. These multi-scale signals are then fused and passed through the state space block, which learns how different regions and sizes relate to each other, all with computational cost that grows only linearly with image size.
Guided Shortcuts that Respect Context
Another key component of FDE-Net lies in how it moves information from early layers to later ones. Traditional U-shaped networks simply copy early details straight to the decoder. FDE-Net instead passes them through a Context Focus Attention mechanism. This module uses very large, but efficient, convolution kernels to let each pixel “see” a wide neighborhood, learning which surrounding regions help clarify whether a boundary is real or just noise. The decoder therefore receives not just crisp edges, but edges informed by the larger anatomy, which leads to smoother, more realistic contours when drawing lesion borders.
What the Tests Show for Real Patients
The researchers tested FDE-Net on three publicly available datasets: two for skin lesions, one for breast tumors in ultrasound, and one for multiple organs in 3D abdominal CT scans. Across all of them, FDE-Net either matched or surpassed strong modern competitors, including classic convolutional networks, Transformer-based models, and recent state space approaches. On a widely used skin-lesion benchmark, it improved a common overlap score (IoU) by more than six percentage points over the original U-Net while using a similar or lower amount of computation than many newer methods. It also showed better detection of small or faint lesions and produced cleaner, more consistent organ outlines in 3D scans.
What This Means for Future Clinical Tools
In simple terms, this work shows that paying attention to both the “frequency view” of images and the multi-scale structure of disease can make computer vision systems more accurate without demanding supercomputers. By carefully suppressing background noise in the frequency domain, efficiently modeling relationships across scales, and enriching the shortcuts between network layers, FDE-Net offers sharper, more reliable segmentation of tumors and organs. With further refinement and validation, such designs could help create faster, more dependable tools to assist doctors in early diagnosis, treatment planning, and tracking how diseases respond to therapy.
Citation: Chen, D., Wu, J., Zhang, XY. et al. A frequency-spatial dual perception network for efficient and accurate medical image segmentation. Sci Rep 16, 7259 (2026). https://doi.org/10.1038/s41598-026-38093-7
Keywords: medical image segmentation, deep learning, frequency domain, state space models, skin and organ lesions